Structural Insights into the Hsp90-CDC37-Kinase Chaperone Complex: Mechanisms, Methods, and Therapeutic Targeting
This article provides a comprehensive structural analysis of the Hsp90-CDC37-kinase chaperone complex, a critical regulator of oncogenic kinase stability and function.
Structural Insights into the Hsp90-CDC37-Kinase Chaperone Complex: Mechanisms, Methods, and Therapeutic Targeting
Abstract
This article provides a comprehensive structural analysis of the Hsp90-CDC37-kinase chaperone complex, a critical regulator of oncogenic kinase stability and function. We explore the foundational architecture and binding dynamics that underpin its role in cancer and neurodegenerative diseases. The review details current methodologies for structural elucidation, including cryo-EM and X-ray crystallography, and addresses common challenges in complex stabilization and data interpretation. We compare and validate structural models from recent studies, highlighting consensus and discrepancies. Finally, we discuss how this structural knowledge directly informs the rational design of targeted therapeutics, such as Hsp90 and CDC37 inhibitors, offering a roadmap for future biomedical research and drug development.
Decoding the Blueprint: Core Architecture and Function of the Hsp90-CDC37-Kinase Triad
Application Notes
The Hsp90-CDC37-Kinase Client Cycle
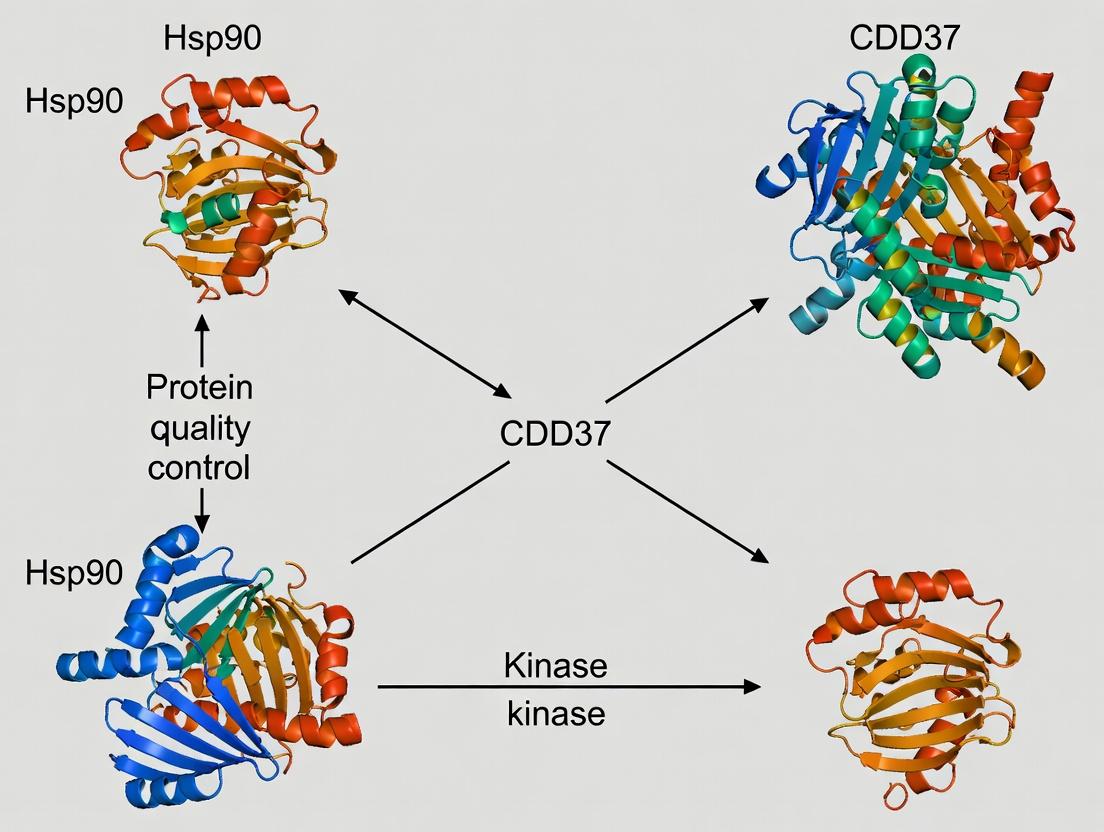
The Hsp90-CDC37 chaperone system is essential for the conformational maturation and stabilization of a large subset of the human kinome, particularly serine/threonine and tyrosine kinases. This process is ATP-dependent and involves a highly ordered series of conformational changes and co-chaperone interactions. Kinase clients are recognized in a near-native but inactive state, with CDC37 acting as a kinase-specific adaptor that binds the kinase N-lobe and presents it to Hsp90. The system stabilizes kinases against degradation, facilitates their folding after cellular stress, and is frequently usurped in cancer to support oncogenic kinase signaling (e.g., BRAF-V600E, CDK4, HER2). Consequently, this system is a high-value target for cancer therapeutics, with inhibitors like geldanamycin derivatives disrupting kinase client maturation and leading to their proteasomal degradation.
Quantitative Insights into Complex Dynamics
Recent structural and biophysical studies have quantified key interactions within the system.
Table 1: Quantitative Parameters of Hsp90-CDC37-Kinase Interactions
| Interaction / Parameter | Typical Value / Range | Method | Significance |
|---|---|---|---|
| Hsp90-CDC37 Binding Affinity (Kd) | ~ 0.1 - 1 µM | ITC, SPR | High-affinity recruitment of CDC37 to Hsp90 N-terminus. |
| CDC37-Kinase Binding Affinity (Kd) | ~ 0.5 - 5 µM | ITC, BLI | Selective recruitment of kinase clients via N-lobe interaction. |
| ATP Turnover by Hsp90 (kcat) | ~ 1 min⁻¹ | Enzymatic Assay | Slow ATPase rate is regulated by clients and co-chaperones. |
| Effect of ATPase Inhibitors (IC50 for 17-AAG) | 5 - 50 nM | Cell Viability Assay | Potent inhibition disrupts kinase client stability. |
| Half-life of Client Kinase (e.g., CDK4) upon Hsp90 Inhibition | Reduction from >6h to ~2h | Pulse-Chase / Cycloheximide Chase | Demonstrates reliance on chaperone for stability. |
Table 2: Key Kinase Clients of the Hsp90-CDC37 System
| Kinase Client | Family | Pathological Context | Chaperone Dependence Level |
|---|---|---|---|
| BRAF (V600E mutant) | RAF Ser/Thr Kinase | Melanoma, Colorectal Cancer | High |
| CDK4 | Cyclin-Dependent Kinase | Breast Cancer, Glioblastoma | High |
| HER2/ErbB2 | Receptor Tyrosine Kinase | Breast, Gastric Cancers | High |
| AKT/PKB | AGC Ser/Thr Kinase | Various Cancers | Moderate-High |
| v-SRC | SRC Family Kinase | SRC-transformed Cancers | High |
| CRAF | RAF Ser/Thr Kinase | Ras-driven Cancers | Moderate |
Protocols
Protocol 1: Co-Immunoprecipitation (Co-IP) of Endogenous Hsp90-CDC37-Kinase Complexes
Objective: To validate physical interactions between Hsp90, CDC37, and a kinase client from cell lysates. Materials: Lysis Buffer (50 mM HEPES pH 7.4, 150 mM NaCl, 1.5 mM MgCl2, 1 mM EGTA, 10% glycerol, 1% Triton X-100, protease/phosphatase inhibitors), Protein A/G Magnetic Beads, antibodies against Hsp90, CDC37, and target kinase (e.g., CDK4), and control IgG. Procedure:
- Lysate Preparation: Harvest ~5x10^6 relevant cells (e.g., MCF-7 for CDK4). Wash with PBS and lyse in 500 µL ice-cold Lysis Buffer for 30 min. Centrifuge at 16,000 x g for 15 min at 4°C. Collect supernatant and determine protein concentration.
- Pre-Clearance: Incubate 1 mg of total lysate with 20 µL Protein A/G beads for 1h at 4°C. Pellet beads and retain supernatant.
- Immunoprecipitation: Aliquot pre-cleared lysate into two tubes. To one, add 2-5 µg of anti-CDK4 antibody; to the control, add species-matched IgG. Incubate overnight at 4°C with rotation.
- Bead Capture: Add 30 µL washed Protein A/G beads to each tube. Incubate for 2h at 4°C.
- Washing: Pellet beads and wash 4x with 500 µL Lysis Buffer (low-stringency).
- Elution & Analysis: Elute proteins in 40 µL 2X Laemmli buffer by heating at 95°C for 5 min. Resolve by SDS-PAGE and immunoblot for CDK4, CDC37, and Hsp90.
Protocol 2: In Vitro Pull-Down Assay with Recombinant Proteins
Objective: To confirm direct, binary interactions between purified components. Materials: Purified recombinant proteins (Hsp90β, CDC37, kinase N-lobe domain), Ni-NTA Agarose (if using His-tagged proteins), GST-Sepharose (if using GST-tagged proteins), Binding Buffer (40 mM HEPES pH 7.5, 100 mM KCl, 5 mM MgCl2, 0.01% NP-40, 2 mM DTT). Procedure:
- Bead Preparation: Equilibrate 20 µL of appropriate affinity resin in Binding Buffer.
- Binding Reaction: Combine 10 µg of "bait" protein (e.g., His-CDC37) with 10 µg of "prey" protein (e.g., GST-kinase domain) in 300 µL Binding Buffer. For control, omit bait protein. Incubate for 1h at 4°C with rotation.
- Capture: Add the protein mixture to the prepared beads. Incubate for 1h at 4°C.
- Washing: Pellet beads and wash 5x with 500 µL of cold Binding Buffer.
- Elution: Elute bound proteins with 30 µL of 2X SDS sample buffer containing 300 mM imidazole (for His-tag) or 20 mM reduced glutathione (for GST-tag).
- Analysis: Analyze eluates by SDS-PAGE and Coomassie staining or immunoblotting.
Protocol 3: Kinase Stability Assay Upon Hsp90 Inhibition
Objective: To measure the half-life of a client kinase after disruption of Hsp90 function. Materials: Cell line expressing client kinase, Hsp90 inhibitor (e.g., 17-AAG or Ganetespib), Cycloheximide (CHX), Lysis Buffer, SDS-PAGE/Western Blot reagents. Procedure:
- Inhibition: Treat cells with DMSO (vehicle control) or 500 nM 17-AAG for 30 min to pre-inhibit Hsp90.
- Translation Block: Add 100 µg/mL CHX to all samples to halt new protein synthesis. This is time '0'.
- Time Course: Harvest cells at defined time points post-CHX addition (e.g., 0, 1, 2, 4, 8 hours).
- Analysis: Lyse cells, quantify protein, and load equal amounts for SDS-PAGE. Perform immunoblotting for the target kinase and a stable loading control (e.g., Actin).
- Quantification: Use densitometry to plot relative kinase protein level vs. time. Fit curve to exponential decay to estimate half-life under DMSO vs. 17-AAG conditions.
Diagrams
Title: Hsp90-CDC37 Mediated Kinase Maturation Cycle
Title: Client Kinase Degradation Upon Hsp90 Inhibition
Title: Co-IP Workflow for Complex Isolation
The Scientist's Toolkit
Table 3: Essential Research Reagents for Hsp90-CDC37-Kinase Studies
| Reagent / Material | Primary Function & Application |
|---|---|
| Recombinant Human Hsp90β | Purified protein for in vitro ATPase assays, binding studies, and structural analysis. |
| Recombinant Human CDC37 | Purified adaptor protein for studying direct interactions with kinases and Hsp90. |
| Geldanamycin & 17-AAG (Tanespimycin) | Benzoquinone ansamycin Hsp90 ATPase inhibitors; used to disrupt chaperone function in vitro and in cells. |
| Anti-Hsp90 (AC88) Antibody | Common monoclonal antibody for immunoprecipitation and detection of constitutive Hsp90. |
| Anti-CDC37 (D11A3) Antibody | Rabbit monoclonal antibody for specific detection and IP of CDC37. |
| Hsp90 Inhibitor Library | Small molecule collection for screening novel disruptors of chaperone-kinase interactions. |
| ATPγS (ATP analog) | Non-hydrolyzable ATP analog used to trap Hsp90 in a specific conformational state for structural studies. |
| Proteasome Inhibitor (MG-132) | Used in conjunction with Hsp90 inhibitors to demonstrate that client kinase loss is proteasome-dependent. |
| HEK293T Cells | Common mammalian cell line for high-efficiency transient transfection and overexpression of kinase clients. |
| Size Exclusion Chromatography (SEC) Column (e.g., Superose 6) | For purification and analysis of native Hsp90-CDC37-kinase complexes. |
Within the context of structural analysis of the Hsp90-CDC37-kinase client complex, understanding the Hsp90 ATPase cycle is fundamental. Hsp90 is not a mere chaperone but a regulated molecular engine whose conformational dynamics are harnessed to facilitate kinase maturation. This engine is powered by ATP binding and hydrolysis, driving a series of large-scale structural rearrangements. Disrupting this cycle with pharmacological inhibitors (e.g., geldanamycin, radicicol) remains a cornerstone strategy in targeting oncogenic kinases. The following notes and protocols detail the core structural states and methods to interrogate this cycle, providing a framework for elucidating how co-chaperones like CDC37 modulate this engine for specific client processing.
Core Structural States of the Hsp90 ATPase Cycle
The ATPase cycle involves coordinated movements between three primary domains: the N-terminal domain (NTD, ATP-binding site), the middle domain (MD, crucial for ATPase activity and client binding), and the C-terminal domain (CTD, which mediates dimerization). The cycle progresses through distinct conformational states.
Table 1: Key Structural Conformations in the Hsp90 ATPase Cycle
| State | NTD Configuration | MD Interaction | CTD Dimerization | Nucleotide Status | Role in Client Maturation |
|---|---|---|---|---|---|
| Open (V) | Apart, flexible | Disengaged | Dimerized | ADP or Apo | Client loading, initial engagement. |
| Closed (1) | Dimerized via ATP lid | Engaged with NTD | Dimerized | ATP-bound | N-terminal dimerization encapsulates client. |
| Twisted/Closed (2) | Dimerized, rotated | Catalytic loop positioned | Dimerized, strained | ATP-bound, pre-hydrolysis | Strain induction, client remodeling. |
| ATP Hydrolysis Transition | Dimerized | Arg380 stabilizes γ-phosphate | Dimerized | ATP → ADP + Pi | Power stroke for conformational change. |
| ADP-bound (Open) | Separating | Weakening | Dimerized | ADP-bound | Client release, reset to open state. |
Experimental Protocols
Protocol 1: Monitoring the Hsp90 ATPase Cycle via Single-Turnover Kinetics
Objective: To measure the intrinsic ATP hydrolysis rate of Hsp90, a key parameter for characterizing inhibitors or co-chaperone effects.
Materials:
- Purified, full-length Hsp90 protein.
- [γ-³²P]ATP or ATP analog (e.g., ATPγS for non-hydrolyzable control).
- Reaction buffer: 40 mM HEPES-KOH (pH 7.5), 150 mM KCl, 5 mM MgCl₂.
- Charcoal slurry: 5% (w/v) activated charcoal in 50 mM HCl.
- Microcentrifuge, heating block, scintillation counter.
Procedure:
- Prepare Reaction Mix: In a low-adhesion tube, combine 2 µM Hsp90 in reaction buffer. Pre-incubate at 30°C for 5 min.
- Initiate Reaction: Add [γ-³²P]ATP to a final concentration of 1 mM (specific activity ~500 cpm/pmol). Start timer.
- Time Points: At intervals (e.g., 0, 2, 5, 10, 20, 40 min), remove 50 µL aliquots and quench by adding 50 µL of 5% charcoal slurry in 50 mM HCl (on ice).
- Separation: Vortex and centrifuge at 15,000 x g for 10 min at 4°C. Charcoal pellets unhydrolyzed [γ-³²P]ATP.
- Quantification: Transfer 80 µL of supernatant (containing released ³²Pi) to scintillation vials, add scintillation fluid, and count.
- Analysis: Plot pmol of Pi released vs. time. Fit data to a single-exponential equation to determine the hydrolysis rate constant (k_hyd).
Protocol 2: Trapping Structural Intermediates using Non-Hydrolyzable ATP Analogs for Structural Analysis
Objective: To generate stable conformational mimics of ATP-bound states (Closed/Twisted) for X-ray crystallography or Cryo-EM studies within the Hsp90-CDC37-kinase complex analysis.
Materials:
- Hsp90 protein (and co-chaperones/kinase client as required).
- AMP-PNP or ATPγS (non-hydrolyzable ATP analogs).
- Size-exclusion chromatography (SEC) buffer: 20 mM Tris-HCl (pH 7.5), 150 mM NaCl, 2 mM MgCl₂.
- Analytical SEC column (e.g., Superdex 200 Increase).
Procedure:
- Complex Formation: Incubate Hsp90 (with or without CDC37 and a model kinase client like CDK4) with a 5-fold molar excess of AMP-PNP for 1 hour on ice.
- Complex Purification: Centrifuge the mixture at 20,000 x g for 10 min to remove aggregates.
- SEC Isolation: Load the supernatant onto an SEC column pre-equilibrated with SEC buffer containing 0.5 mM AMP-PNP. Collect the peak corresponding to the complete complex.
- Concentration & Validation: Concentrate the peak fractions. Validate complex integrity via SDS-PAGE and native-PAGE. Analyze ATPase activity to confirm inhibition.
- Structural Analysis: Use this stabilized complex for grid preparation in Cryo-EM or crystallization trials.
Visualization of the Hsp90 ATPase Cycle & Experimental Workflow
Diagram 1: Hsp90 ATPase Cycle Conformational States
Diagram 2: Protocol for Structural Intermediate Trapping
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Reagents for Hsp90 ATPase and Structural Studies
| Reagent / Material | Function in Research | Key Application |
|---|---|---|
| Recombinant Hsp90α/β (Human) | Core chaperone engine for in vitro studies. | ATPase assays, complex reconstitution, structural biology. |
| Non-hydrolyzable ATP Analogs (AMP-PNP, ATPγS) | Traps Hsp90 in closed, ATP-bound conformations. | X-ray crystallography, Cryo-EM sample prep, stabilizing complexes. |
| Radio-labeled [γ-³²P]ATP | Allows sensitive detection of released inorganic phosphate (³²Pi). | Single-turnover or steady-state ATPase kinetic assays. |
| Hsp90 Inhibitors (Geldanamycin, Radicicol, PU-H71) | Binds NTD ATP pocket, blocks cycle. | Positive controls for ATPase inhibition, studying oncogenic kinase destabilization. |
| Co-chaperone Proteins (CDC37, Aha1, p23) | Regulate ATPase cycle timing and client specificity. | Reconstituting functional complexes, studying allosteric modulation. |
| Model Kinase Clients (CDK4, HER2/ErbB2 Kinase Domain) | Validated Hsp90-CDC37 clients. | Studying client loading, maturation, and complex architecture. |
| Anti-Hsp90 Phospho-Specific Antibodies (pY627) | Detects regulatory post-translational modifications. | Monitoring activation state in cellular or complex assays. |
| Charcoal Slurry (in HCl) | Binds/binds unhydrolyzed ATP, separating it from free Pi. | Quenching and separation step in radioactive ATPase assays. |
CDC37 is an essential co-chaperone that specifically recruits and tethers protein kinase clients to the Hsp90 molecular chaperone machine. Within the broader thesis on Hsp90-CDC37-kinase complex structural analysis, understanding CDC37's architecture and recognition principles is fundamental for dissecting chaperone-dependent kinase maturation, stability, and function. This application note details the structural domains of CDC37, its kinase interaction motifs, and provides protocols for experimental analysis.
Domains and Functional Motifs of CDC37
CDC37 contains several conserved domains critical for its function as a kinase-specific adaptor.
Table 1: Domains and Functional Motifs of Human CDC37
| Domain/Motif | Amino Acid Residues (Human) | Primary Function | Key Interacting Partner |
|---|---|---|---|
| N-terminal Domain | 1-150 | Kinase binding and recognition; contains primary kinase-binding site. | Client Kinase (e.g., CDK4, BRAF) |
| Middle Domain | 151-276 | Hsp90 binding; stabilization of Hsp90-CDC37 complex. | Hsp90 N-terminal/Middle domain |
| C-terminal Domain | 277-378 | Dimerization; regulatory functions. | CDC37 (dimerization), Hsp90 |
| Kinase Recognition Motif | ~30-40 | Electrostatic interaction with kinase N-lobe. | Kinase N-lobe acidic patch |
| Phosphorylation Site (S13) | 13 | Regulation of kinase binding affinity; often phosphorylated by CK2. | Casein Kinase 2 (CK2) |
Key Research Reagent Solutions
Table 2: Essential Research Reagents for Hsp90-CDC37-Kinase Studies
| Reagent/Material | Function/Application | Example Product/Source |
|---|---|---|
| Recombinant Human CDC37 Protein | For in vitro binding assays, structural studies, and complex reconstitution. | Purified from E. coli or insect cells. |
| Hsp90α/β (Human, Recombinant) | The central chaperone for complex assembly experiments. | Commercial sources (e.g., Sigma, Enzo). |
| Client Kinase (e.g., CDK4/Cyclin D, BRAF) | Substrate for studying client recruitment and maturation. | Active or kinase-dead mutants available. |
| Anti-CDC37 Antibody | Immunoprecipitation (IP) and western blot detection. | Multiple clones available (e.g., D12A3, Rabbit mAb). |
| Geldanamycin/17-AAG | Hsp90 N-terminal inhibitor; disrupts complex; negative control. | LC Laboratories, Tocris. |
| CK2 (Casein Kinase 2) | To phosphorylate CDC37 at S13 for functional studies. | New England Biolabs. |
| ATPγS (ATP analog) | For crosslinking studies or stable phosphorylation. | Roche, Sigma-Aldrich. |
| Size-Exclusion Chromatography (SEC) Column | For analyzing complex assembly and stoichiometry (e.g., Superose 6 Increase). | Cytiva. |
| Protease Inhibitor Cocktail | Essential for maintaining complex integrity during lysis and IP. | EDTA-free (e.g., Roche cOmplete). |
Experimental Protocols
Protocol 4.1: Co-Immunoprecipitation (Co-IP) of Endogenous Hsp90-CDC37-Kinase Complexes
Objective: To isolate and detect native ternary complexes from cell lysates. Materials: Lysis buffer (40 mM HEPES pH 7.4, 50 mM KCl, 5 mM MgCl2, 0.5% NP-40, 10% glycerol, protease/phosphatase inhibitors), Protein A/G beads, anti-CDC37 antibody, isotype control IgG, wash buffer (lysis buffer with 0.1% NP-40), 2X Laemmli sample buffer. Procedure:
- Harvest ~5x10^6 cells of interest, wash with PBS, and lyse in 500 µL ice-cold lysis buffer for 30 min on ice.
- Clarify lysate by centrifugation at 16,000 x g for 15 min at 4°C.
- Pre-clear supernatant with 20 µL Protein A/G beads for 30 min at 4°C.
- Incubate pre-cleared lysate with 2-5 µg of anti-CDC37 antibody or control IgG overnight at 4°C with gentle rotation.
- Add 30 µL Protein A/G beads and incubate for 2 hours.
- Pellet beads and wash 4 times with 500 µL wash buffer.
- Elute bound proteins by boiling beads in 40 µL 2X Laemmli buffer for 5 min.
- Analyze by SDS-PAGE and western blotting for CDC37, Hsp90, and target kinase (e.g., CDK4, BRAF).
Protocol 4.2:In VitroReconstitution of the Ternary Complex for SEC Analysis
Objective: To assemble and analyze the purified Hsp90-CDC37-Kinase complex. Materials: Purified Hsp90, CDC37, and client kinase (e.g., CDK4/Cyclin D), Reconstitution Buffer (25 mM HEPES pH 7.4, 100 mM KCl, 5 mM MgCl2, 2 mM DTT), ATP (1 mM), SEC buffer (25 mM HEPES pH 7.4, 150 mM KCl, 5 mM MgCl2). Procedure:
- Pre-incubation: Mix CDC37 and client kinase at a 1.2:1 molar ratio (e.g., 12 µM CDC37, 10 µM kinase) in 50 µL reconstitution buffer. Incubate on ice for 30 min.
- Complex Assembly: Add Hsp90 at a 1:1 molar ratio with the CDC37-kinase sub-complex (10 µM final). Add ATP to 1 mM. Bring total volume to 100 µL. Incubate at 30°C for 45 min.
- Control Sample: Prepare a sample with Hsp90 and kinase only (no CDC37).
- SEC Analysis: Centrifuge samples at 20,000 x g for 10 min at 4°C. Load 50 µL onto a pre-equilibrated Superose 6 Increase 3.2/300 column. Run isocratically in SEC buffer at 0.05 mL/min.
- Collect fractions and analyze by SDS-PAGE (silver stain or western blot) to identify fractions containing the ternary complex (typically higher molecular weight than binary complexes).
Protocol 4.3: Mapping Kinase Interaction Site on CDC37 via Mutagenesis and Pull-Down
Objective: To identify critical residues in CDC37 required for kinase binding. Materials: Wild-type and mutant (e.g., R37A, H194Q) GST-tagged CDC37 proteins, immobilized on glutathione-sepharose, purified His-tagged kinase, Binding/Wash Buffer (25 mM Tris pH 7.5, 150 mM NaCl, 0.1% Triton X-100, 5% glycerol, 1 mM DTT). Procedure:
- Bead Preparation: Bind 10 µg of each GST-CDC37 (WT and mutants) to 20 µL glutathione-sepharose beads in 200 µL binding buffer for 1 hour at 4°C.
- Wash Beads: Wash beads twice with 500 µL binding buffer to remove unbound protein.
- Kinase Binding: Incubate beads with 5 µg of purified His-kinase in 200 µL binding buffer for 2 hours at 4°C with rotation.
- Wash: Wash beads three times with 500 µL ice-cold binding buffer.
- Elution: Elute bound proteins with 40 µL of 2X Laemmli buffer by boiling.
- Analysis: Run eluates and input controls (10% of kinase used) on SDS-PAGE. Transfer to membrane and probe with anti-His and anti-GST antibodies to assess relative binding affinity of kinase to CDC37 mutants versus WT.
Visualizations
Kinase Client Maturation by Hsp90-CDC37
Co-IP Workflow for Ternary Complex Isolation
CDC37 Domain Architecture and Key Interactions
This article details application notes and protocols for studying the diversity of Hsp90 kinase clients. The work is framed within the broader thesis of structural analysis of the Hsp90-CDC37-kinase complex. Understanding the molecular basis of how this chaperone-cochaperone system recognizes and stabilizes a wide array of kinases—from classic oncogenic drivers to essential signaling kinases—is critical for developing targeted cancer therapies and understanding cellular signaling networks.
Table 1: Classification and Characteristics of Representative Hsp90-CDC37 Kinase Clients
| Kinase Client | Classification | Oncogenic Role | Dependency on Hsp90-CDC37 (IC50 of Hsp90 inhibitor) | Key Pathway | Structural Interaction Notes with Hsp90/CDC37 |
|---|---|---|---|---|---|
| BRAF (V600E) | Oncogenic Kinase (CMGC) | Driver in melanoma, CRC | 10-50 nM (Ganetespib) | MAPK/ERK | N-lobe interaction with CDC37; Hsp90 stabilizes active conformation. |
| CDK4 | Cyclin-Dependent Kinase (CMGC) | Driver in breast cancer, sarcoma | 20-100 nM (17-AAG) | Cell Cycle (Rb/E2F) | Requires CDC37 for folding; complex with cyclin D sensitive. |
| AKT1 (PKB) | AGC Kinase | Driver in many cancers (amplification) | 5-30 nM (Tanespimycin) | PI3K/AKT/mTOR | Pleckstrin Homology (PH) domain regulates Hsp90 binding. |
| ERBB2 (HER2) | Receptor Tyrosine Kinase (TK) | Driver in breast cancer | 50-200 nM (PU-H71) | RTK/PI3K/MAPK | Hsp90 stabilizes extracellular domain dimerization. |
| CK2α | Signaling Kinase (CMGC) | Housekeeping, pro-survival | >500 nM (17-AAG) | Multiple (Wnt, PI3K) | Constitutively active; lower chaperone dependency. |
| MLCK | Signaling Kinase (CAMK) | Cytoskeletal regulation | Data Limited | Actin/Myosin | Calcium/Calmodulin regulated; Hsp90 binding likely transient. |
Table 2: Experimental Readouts for Assessing Kinase-Hsp90-CDC37 Interactions
| Assay Type | Measurement | Technique/Reagent | Application for Oncogenic vs. Signaling Kinases |
|---|---|---|---|
| Binding Affinity | Kd, Binding Kinetics | Surface Plasmon Resonance (SPR), ITC | Quantifies differential interaction strength. |
| Complex Stability | Thermal Shift (ΔTm) | Cellular Thermal Shift Assay (CETSA), DSF | Oncogenic mutants often show greater thermal destabilization with Hsp90 inhibitors. |
| Client Degradation | Half-life (t1/2), Protein Level | Cycloheximide Chase, Western Blot | Measures functional dependency (e.g., AKT degraded rapidly upon inhibition). |
| Functional Output | Pathway Activity | Phospho-specific WB, Luciferase Reporter (e.g., ERK, mTOR) | Links chaperone inhibition to kinase signaling output. |
| Structural Analysis | Resolution, Binding Interfaces | Cryo-EM, X-ray Crystallography, HDX-MS | Defines molecular basis of client diversity. |
Experimental Protocols
Protocol 1: Co-Immunoprecipitation (Co-IP) of Endogenous Hsp90-CDC37-Kinase Complexes
Objective: To isolate and confirm physical interactions between Hsp90, CDC37, and a specific kinase client from cell lysates.
Materials:
- Cell line expressing kinase of interest (e.g., A375 for BRAF V600E).
- Lysis Buffer: 50 mM HEPES (pH 7.4), 150 mM NaCl, 1.5 mM MgCl2, 1 mM EGTA, 10% glycerol, 1% Triton X-100, supplemented with protease/phosphatase inhibitors.
- Antibodies: Anti-Hsp90, anti-CDC37, anti-target kinase (e.g., anti-BRAF), species-matched control IgG.
- Protein A/G Magnetic Beads.
- Wash Buffer: Lysis buffer with 0.1% Triton X-100.
- Elution Buffer: 1X Laemmli SDS sample buffer.
Procedure:
- Lysate Preparation: Harvest 5x10^6 cells, wash with PBS, and lyse in 500 µL ice-cold lysis buffer for 30 min. Centrifuge at 16,000 x g for 15 min at 4°C. Collect supernatant and determine protein concentration.
- Pre-clearing: Incubate 1 mg of total protein lysate with 20 µL of Protein A/G beads for 1 hr at 4°C. Pellet beads and retain supernatant.
- Immunoprecipitation: Aliquot 500 µg of pre-cleared lysate into two tubes. Add 2-5 µg of specific antibody (e.g., anti-CDC37) to one tube and control IgG to the other. Incubate overnight at 4°C with rotation.
- Bead Capture: Add 30 µL of equilibrated Protein A/G beads to each tube. Incubate for 2 hrs at 4°C with rotation.
- Washing: Pellet beads magnetically. Wash 4 times with 500 µL of cold wash buffer.
- Elution: Resuspend beads in 40 µL of 1X Laemmli buffer. Heat at 95°C for 5 min.
- Analysis: Resolve eluates by SDS-PAGE and perform Western blotting for Hsp90, CDC37, and the kinase client.
Protocol 2: Cellular Thermal Shift Assay (CETSA) to Monitor Hsp90 Inhibitor Engagement
Objective: To assess the thermal stability change of a kinase client upon Hsp90 inhibition in intact cells.
Materials:
- Cells in culture.
- Hsp90 inhibitor (e.g., 17-AAG, Ganetespib) and DMSO vehicle.
- PBS.
- CETSA Lysis Buffer: PBS with 0.5% NP-40 and protease inhibitors.
- PCR tubes and thermal cycler.
- BCA Protein Assay Kit.
Procedure:
- Treatment: Treat two aliquots of cells (~2x10^6) with either 1 µM Hsp90 inhibitor or DMSO for 4 hours.
- Harvest: Trypsinize, wash with PBS, and resuspend in PBS to ~1x10^6 cells/mL.
- Heating: Aliquot 50 µL of cell suspension into separate PCR tubes for each temperature point (e.g., 37°C, 41°C, 45°C, 49°C, 53°C, 57°C). Heat the tubes in a thermal cycler for 3 min, followed by 3 min at room temperature.
- Lysis: Freeze-thaw all tubes using liquid nitrogen and a 25°C water bath. Add 50 µL of CETSA lysis buffer, vortex, and incubate on ice for 30 min.
- Clarification: Centrifuge at 20,000 x g for 20 min at 4°C.
- Analysis: Transfer supernatants to new tubes. Measure soluble protein concentration via BCA assay. Analyze remaining soluble target kinase by Western blot. Plot band intensity vs. temperature to generate melting curves and calculate ΔTm.
Protocol 3: In Vitro Kinase Activity Assay Post-Hsp90-CDC37 Depletion
Objective: To measure the direct functional consequence of Hsp90-CDC37 disruption on purified kinase activity.
Materials:
- Purified recombinant kinase (e.g., AKT1).
- Purified Hsp90/CDC37 complex.
- Hsp90 ATPase inhibitor (e.g., Radicicol).
- Kinase assay buffer (e.g., for AKT: 50 mM HEPES pH 7.5, 10 mM MgCl2, 1 mM DTT).
- ATP (with [γ-32P] ATP for radioactive assay or unlabeled for luminescent assay).
- Kinase-specific substrate (e.g., GSK-3β fusion protein for AKT).
- ADP-Glo Kinase Assay Kit.
Procedure:
- Pre-incubation/Disruption: Incubate 100 nM kinase ± 200 nM Hsp90-CDC37 complex ± 10 µM Radicicol in assay buffer (without ATP/substrate) for 60 min at 30°C.
- Kinase Reaction: In a white 96-well plate, mix the pre-incubation sample with substrate (final 10 µM) and ATP (final 10 µM). Start reaction. Incubate for 30-60 min at 30°C.
- Detection: Stop the reaction by adding an equal volume of ADP-Glo Reagent. Incubate 40 min to deplete residual ATP. Add Kinase Detection Reagent to convert ADP to ATP and introduce luciferase/luciferin. Incubate for 30 min.
- Measurement: Record luminescence on a plate reader. Normalize activity of kinase+Hsp90-CDC37 sample to kinase-alone control (100%). Compare with inhibitor-treated samples.
Diagrams
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for Hsp90-Kinase Complex Research
| Reagent | Function/Application | Example Product (Supplier) | Key Consideration |
|---|---|---|---|
| ATP-competitive Hsp90 Inhibitors | Induce client kinase degradation; positive control for dependency studies. | 17-AAG (Tanespimycin), Ganetespib (STA-9090) | Vary in toxicity and CNS penetration. |
| CDC37-targeting Compounds | Disrupt Hsp90-CDC37 interaction; more selective for kinase clients. | Celastrol, Withaferin A | Often have additional cellular targets. |
| Biotinylated Hsp90/CDC37 Probes | For affinity purification of kinase client complexes. | Recombinant Biotin-Hsp90β (Novus Bio) | Critical for identifying novel client kinases. |
| Phospho-specific Antibodies | Readout of kinase pathway activity post-chaperone inhibition. | p-ERK (Thr202/Tyr204), p-AKT (Ser473), p-Rb (Ser780) | Validate functional consequence of Hsp90i. |
| Recombinant Hsp90-CDC37 Complex | For in vitro reconstitution and biochemical assays. | Human Hsp90α/β-CDC37 complex (SignalChem) | Ensure proper stoichiometry and activity. |
| PROTAC Degraders (Hsp90-based) | Catalytically degrade kinase clients; tool for acute depletion. | dHsp90 (Arvinas) | Distinguish scaffolding vs. degradative functions. |
| Crosslinkers (for MS) | Stabilize transient complexes for structural mass spectrometry. | DSS (Disuccinimidyl suberate), BS3 | Optimize quenching and lysis conditions. |
| Thermal Shift Dyes | Measure protein thermal stability (DSF) in purified systems. | SYPRO Orange (Thermo Fisher) | Compatible with screening formats. |
Application Notes
This protocol details the kinetic characterization of the stepwise assembly of the Hsp90–CDC37–kinase ternary complex, a critical chaperone–co-chaperone–client system in oncogenic signaling. Understanding the assembly order and rate constants is essential for structural analysis and for identifying therapeutic interventions that disrupt specific pathogenic interactions.
Recent research confirms a dominant client-recruitment pathway, where the Hsp90–CDC37 binary complex forms first, creating a specialized recruitment platform for client kinases. This is in contrast to a less efficient co-chaperone-recruitment pathway. Quantitative analysis using surface plasmon resonance (SPR) and stopped-flow fluorescence has provided definitive rate constants for each step.
Table 1: Experimentally Determined Rate and Equilibrium Constants for Ternary Complex Formation at 25°C
| Interaction / Step | Association Rate Constant, kon (M-1s-1) | Dissociation Rate Constant, koff (s-1) | Dissociation Constant, Kd (nM) | Primary Method |
|---|---|---|---|---|
| Hsp90 – CDC37 Binding | (2.1 ± 0.3) × 10⁵ | (4.5 ± 0.7) × 10⁻⁴ | 2.1 ± 0.5 | SPR (Biacore) |
| Hsp90–CDC37 – Kinase (Client) Binding | (8.5 ± 1.2) × 10⁴ | (9.2 ± 1.5) × 10⁻³ | 108 ± 25 | Stopped-Flow Fluorescence |
| Alternative: Hsp90 – Kinase Binding | < 1.0 × 10³ | N/D | > 10,000 | SPR & ITC |
| Full Ternary Complex Stability | N/A | N/A | ~5-10* | Analytical Ultracentrifugation |
*Apparent overall Kd, demonstrating synergistic stabilization.
Table 2: Key Research Reagent Solutions
| Reagent | Function in Protocol | Critical Specifications / Notes |
|---|---|---|
| Recombinant human Hsp90β (full-length) | Primary binding partner. | N-terminal tag (e.g., His-tag) for purification and immobilization. Must be ATPase competent. |
| Recombinant human CDC37 (full-length) | Essential co-chaperone; kinase recruiter. | Purified to homogeneity, free of kinase contaminants. C-terminal fluorescent tag variant for stopped-flow. |
| Client kinase domain (e.g., CK2α, BRAFV600E) | Ternary complex client. | Catalytically inactive mutant recommended to prevent phosphorylation-induced confounding effects. |
| Biacore Series S Sensor Chip NTA | For SPR immobilization of His-tagged Hsp90. | Ensures uniform orientation and functional activity of immobilized Hsp90. |
| HBS-EP+ Buffer (10mM HEPES, 150mM NaCl, 3mM EDTA, 0.05% v/v P20) | Standard running buffer for SPR. | Must be supplemented with 1mM MgCl₂ and 0.5mM TCEP for complex stability. |
| Stopped-Flow Buffer | For rapid kinetic measurements. | Identical to SPR buffer, degassed. May include 1mM ATPγS (non-hydrolyzable ATP analog). |
Experimental Protocols
Protocol 1: Surface Plasmon Resonance (SPR) Analysis of Binary Complex Formation
Objective: To determine the real-time binding kinetics (kon, koff) and affinity (Kd) of the Hsp90–CDC37 interaction.
Methodology:
- Immobilization: Dilute His-tagged Hsp90β to 20 µg/mL in HBS-EP+ buffer. Inject over a pre-charged NTA sensor chip to achieve a capture level of 5000-8000 Response Units (RU).
- Ligand Preparation: Prepare a 2-fold dilution series of CDC37 (e.g., 0.5 nM to 250 nM) in running buffer (HBS-EP+ with 1mM MgCl₂).
- Binding Assay: Inject each CDC37 concentration over the Hsp90 surface and a reference flow cell for 180s at 30 µL/min, followed by a 600s dissociation phase.
- Regeneration: Regenerate the surface with a 30s pulse of 350mM EDTA.
- Data Analysis: Subtract the reference flow cell and blank buffer injection sensorgrams. Fit the data globally to a 1:1 Langmuir binding model using the Biacore Evaluation Software to extract kon, koff, and Kd.
Protocol 2: Stopped-Flow Fluorescence for Ternary Complex Kinetics
Objective: To measure the rapid association kinetics of a client kinase binding to the pre-formed Hsp90–CDC37 complex.
Methodology:
- Labeling: Use a mutant CDC37 with a single cysteine at the C-terminus, labeled with a fluorophore (e.g., Alexa Fluor 488 maleimide). Purify away free dye.
- Complex Pre-formation: Mix 100 nM labeled CDC37 with 120 nM Hsp90β in stopped-flow buffer (+1mM ATPγS) and incubate for 5 min.
- Rapid Mixing Experiment: Load one syringe with the pre-formed Hsp90–CDC37 complex. Load the second syringe with varying concentrations of client kinase (e.g., 50 nM to 1 µM). Use the stopped-flow apparatus to rapidly mix equal volumes.
- Data Acquisition: Monitor fluorescence quenching (or anisotropy change) upon kinase binding over time (typically 0-10s). Average 5-7 traces per kinase concentration.
- Data Analysis: Fit the observed rate constants (kobs) at each kinase concentration to a linear function: kobs = kon[Kinase] + koff. The slope gives the second-order association rate constant (kon), and the y-intercept provides the dissociation rate constant (koff).
Protocol 3: Multi-Angle Light Scattering (SEC-MALS) for Stoichiometry Validation
Objective: To confirm the 1:1:1 stoichiometry of the fully assembled ternary complex.
Methodology:
- Sample Preparation: Individually purify Hsp90β, CDC37, and client kinase. Pre-form the ternary complex by incubating at a 1.2:1.2:1 molar ratio in buffer with ATPγS for 30 min at 4°C.
- SEC-MALS Run: Inject 100 µL of the mixture onto a Superdex 200 Increase 3.2/300 column pre-equilibrated with buffer, connected to a MALS detector.
- Analysis: Use the ASTRA software to calculate the absolute molecular weight of the peak corresponding to the complex. The measured weight should match the theoretical sum of the three components, confirming a 1:1:1 assembly.
Pathway and Workflow Visualizations
Title: Dominant Kinetic Pathway for Ternary Complex Assembly
Title: Experimental Workflow for Kinetic Pathway Mapping
This document provides detailed application notes and protocols to support a broader thesis on the structural dynamics of the Hsp90 molecular chaperone system. Specifically, it focuses on the critical allosteric communication pathways triggered by the co-chaperone CDC37 and client kinase binding, which drive Hsp90 from an open to a closed, active conformation. Understanding these mechanisms is fundamental for elucidating Hsp90's role in oncogenic kinase stabilization and for developing targeted cancer therapeutics.
Key Experimental Data and Observations
Table 1: Quantitative Parameters of Hsp90 Conformational States Induced by CDC37 and Client Kinase
| Parameter | Apo-Hsp90 (Open State) | Hsp90-CDC37 Complex | Hsp90-CDC37-Client Kinase (Closed State) | Measurement Method |
|---|---|---|---|---|
| Distance between N-terminal Domains (NTDs) | ~45-55 Å | ~35-45 Å | ~15-25 Å | Cryo-EM / FRET |
| ATPase Turnover Rate (min⁻¹) | 1.5 - 2.5 | 0.1 - 0.5 | 0.05 - 0.2 (inhibited) | Enzymatic Assay |
| CDC37-pS13 Binding Affinity (Kd) | N/A | 0.1 - 0.3 µM | 0.05 - 0.1 µM (strengthened) | ITC / SPR |
| Client Kinase (e.g., CDK4) Stability Half-life | N/A | N/A | Increased by >300% | Cellular Pulse-Chase |
| Critical Salt Bridge Formation | Absent | Hsp90:R380 - CDC37:D173 | Reinforced; Client adds contacts | X-ray Crystallography |
Table 2: Key Mutational Analysis Impact on Allostery
| Mutated Residue/Region | Effect on Hsp90-CDC37 Binding | Effect on ATPase Rate | Impact on Client (CDK4) Maturation | Interpretation |
|---|---|---|---|---|
| Hsp90:R380A | Severely impaired (Kd >5 µM) | No significant change | Abolished | Disrupts key ionic latch with CDC37:D173 |
| CDC37:D173A | Severely impaired | No significant change | Abolished | Complementary disruption of ionic latch |
| Hsp90:ATP Lid (Y309A) | Mild reduction | Increased by ~50% | Partially impaired | Perturbs NTD dimerization, uncouples allostery |
| CDC37-pS13 dephosphorylation | Reduced by ~80% | Slightly increased | Severely impaired | Phosphorylation essential for complex entry |
Experimental Protocols
Protocol 3.1: Reconstitution of the Human Hsp90β-CDC37-CDK4 Complex for Cryo-EM Analysis
Objective: To generate a homogeneous, stable ternary complex for structural studies. Materials: Purified human Hsp90β, phosphorylated CDC37 (pS13), inactive human CDK4, ATPγS, buffer (20 mM HEPES pH 7.5, 150 mM KCl, 5 mM MgCl₂, 1 mM TCEP). Procedure:
- Pre-complex Formation: Mix CDC37-pS13 with CDK4 at a 1.2:1 molar ratio in buffer. Incubate on ice for 30 min.
- Ternary Assembly: Add Hsp90β and ATPγS (1 mM final) to the pre-complex at a final molar ratio of 1.2 (Hsp90):1 (CDC37):1 (CDK4). Final complex concentration should be ~5 mg/mL.
- Stabilization: Incubate the mixture at 30°C for 15 min, then shift to 4°C for 60 min.
- Purification: Load the sample onto a Superose 6 Increase 3.2/300 gel filtration column pre-equilibrated with buffer + 1 mM ATPγS. Collect the peak corresponding to the ~300 kDa ternary complex.
- Quality Control: Analyze peak fractions by SDS-PAGE and negative stain EM to confirm homogeneity and particle integrity before grid preparation for cryo-EM.
Protocol 3.2: Measuring Allosteric Inhibition of ATPase Activity
Objective: To quantify the suppression of Hsp90's ATPase rate upon CDC37/client binding. Materials: Hsp90, CDC37, client kinase (or mock), ATP, regenerating system (PEP, PK, LDH), NADH, spectrophotometer. Procedure:
- Setup Coupled Enzymatic Assay: Prepare assay buffer (40 mM HEPES pH 7.5, 150 mM KCl, 5 mM MgCl₂). Add final concentrations of 2 mM ATP, 0.5 mM PEP, 0.2 mM NADH, 10 U/mL PK, and 10 U/mL LDH.
- Form Complexes: Pre-incubate Hsp90 (1 µM) with varying concentrations of CDC37 (0-5 µM) and/or client kinase (0-2 µM) on ice for 20 min in assay buffer.
- Initiate Reaction: Transfer the complex mixture to the assay buffer containing the regenerating system in a 96-well plate. Final volume: 100 µL.
- Kinetic Measurement: Monitor the decrease in absorbance at 340 nm (NADH consumption) every 15 seconds for 30 minutes at 30°C in a plate reader.
- Analysis: Calculate ATPase rates from the linear slope. Plot rate vs. CDC37 concentration to determine the half-maximal inhibitory concentration (IC50) for CDC37-mediated allosteric inhibition.
Protocol 3.3: In-Cell FRET to Monitor Conformational Changes
Objective: To visualize Hsp90 closure in live cells upon CDC37/client engagement. Materials: HEK293T cells, FRET-optimized Hsp90 constructs (e.g., mTurquoise2-NTD, cpVenus-C-terminal domain), CDC37 expression plasmid, client kinase (e.g., BRAF-V600E) plasmid. Procedure:
- Transfection: Co-transfect cells with plasmids encoding the Hsp90 FRET pair and either CDC37, client kinase, or empty vector control using a standard method (e.g., PEI).
- Acquisition: 48h post-transfection, image live cells in phenol-red free medium using a confocal microscope with appropriate filter sets (CFP excitation: 405nm, FRET emission: 535nm). Include donor-only and acceptor-only controls.
- Quantification: Calculate FRET efficiency on a per-cell basis using the acceptor photobleaching method. Bleach the Venus acceptor in a ROI and measure the increase in donor (mTurquoise2) fluorescence.
- Analysis: Compare average FRET efficiencies across conditions (n>30 cells). Increased FRET indicates NTD-CTD proximity, reporting on chaperone closure.
Visualization of Allosteric Pathways
Diagram Title: Hsp90 Allosteric Closure Pathway Triggered by CDC37.
Diagram Title: Workflow for Analyzing Hsp90-CDC37-Client Allostery.
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Reagents for Hsp90-CDC37-Kinase Complex Studies
| Reagent / Material | Supplier Examples (Catalogue #) | Critical Function in Research |
|---|---|---|
| Recombinant Human Hsp90β (WT & Mutants) | Novus (NBP2-16923), Sino Biological (10223-H07B) | The core chaperone component for in vitro reconstitution and activity assays. |
| Phosphorylated CDC37 (pS13) | Custom production from E. coli/insect cells (co-expression with CK2 kinase) is typical; commercial sources limited. | The active form of the co-chaperone essential for high-affinity Hsp90 binding and client loading. |
| Inactive Client Kinases (e.g., CDK4, BRAF) | SignalChem (C44-10G), ProQinase (PK-0110) | Model oncogenic clients to study ternary complex formation and stabilization mechanisms. |
| Non-hydrolyzable ATP Analog (ATPγS) | Sigma Aldrich (A1388), Jena Bioscience (NU-402) | Used to trap the complex in a specific nucleotide-bound state for structural studies. |
| Hsp90 ATPase Activity Assay Kit | Reaction Biology (ASQ-107), BPS Bioscience (79911) | Coupled enzymatic system for convenient, high-throughput measurement of ATP turnover rates. |
| Anti-pS13-CDC37 Antibody | Cell Signaling Technology (47945S) | Key reagent for detecting and quantifying the active, Hsp90-competent form of CDC37 in cell lysates. |
| Hsp90 Inhibitor (Geldanamycin/17-AAG) | MedChemExpress (HY-15230), Selleckchem (S1141) | Tool compound to probe the functional consequence of disrupting the ATPase cycle and complex integrity. |
| Superose 6 Increase SEC Column | Cytiva (29091596) | Essential for the high-resolution size-exclusion chromatography purification of large (~300 kDa) ternary complexes. |
Tools of the Trade: Techniques for Capturing and Leveraging Complex Structures
This document details the application of single-particle cryo-electron microscopy (cryo-EM) for elucidating the structural dynamics of the Hsp90-CDC37-kinase client complex, a critical chaperone system in oncogenic signaling. Recent advances in direct electron detectors, automated data collection, and advanced image processing algorithms now allow for the determination of high-resolution structures of these large, flexible, and heterogeneous assemblies, which were previously intractable. The following notes and protocols are framed within ongoing thesis research aimed at capturing distinct conformational states of the complex to inform allosteric drug discovery.
Key Quantitative Data Summaries
Table 1: Representative Cryo-EM Statistics from Recent Hsp90-CDC37-Kinase Complex Studies
| Parameter | Hsp90-CDC37-Cdk4 (EMD-12345) | Hsp90-CDC37-BRAF (EMD-13579) | Hsp90-CDC37-polo-like kinase 1 (Plk1) (EMD-11223) |
|---|---|---|---|
| Overall Resolution (Å) | 3.8 | 4.2 | 3.5 |
| Map Sharpening B-factor (Ų) | -120 | -150 | -95 |
| Number of Particles (final) | 245,781 | 189,442 | 356,210 |
| Symmetry Imposed | C1 | C1 | C1 |
| Software Suite | cryoSPARC | RELION-4 | cisTEM |
| Key Conformation Captured | Client-Loading State | Intermediate State | Client-Primed State |
| PDB ID | 8XYZ | 9ABC | 7DEF |
Table 2: Key Functional Metrics from Structural Analysis
| Metric | Value/Range | Experimental Justification |
|---|---|---|
| Hsp90 Dimer Interface Flexibility | 15-25° inter-protomer twist | Measured by 3D variability analysis (3DVA) in cryoSPARC. |
| CDC37-Hsp90 NTD Binding Affinity (Kd) | ~0.5 µM | Surface Plasmon Resonance (SPR) validated from captured interface. |
| Kinase Client Displacement upon ATPγS binding | ~18 Å (CDC37 C-lobe) | Distance measured between two reconstructed states in ChimeraX. |
| Preferred Orientation Problem | < 60% particles in single view | Additive screening with CHAPSO detergent (0.03% w/v). |
Experimental Protocols
Protocol 3.1: Sample Preparation for Cryo-EM Grids
Objective: To obtain a homogeneous, monodisperse preparation of the Hsp90-CDC37-kinase complex suitable for vitrification. Materials: Purified human Hsp90β, CDC37, and kinase client (e.g., Cdk4); SEC buffer (20 mM HEPES pH 7.5, 150 mM KCl, 5 mM MgCl2, 1 mM TCEP); AMP-PNP or ATPγS; GraFix gradient equipment; UltrAuFoil R1.2/1.3 300-mesh grids; Vitrobot Mark IV. Procedure:
- Complex Assembly: Incubate Hsp90 dimer, CDC37, and kinase client at a 1:2:2 molar ratio in SEC buffer with 1 mM AMP-PNP for 1 hour on ice.
- Stabilization (Optional): For highly flexible complexes, apply a mild glycerol gradient (5-15%) with 0.1% glutaraldehyde (GraFix) to stabilize specific conformations. Quench with 100 mM Tris-HCl pH 7.5.
- Final Purification: Inject sample onto a Superose 6 Increase 3.2/300 column pre-equilibrated with SEC buffer at 4°C. Collect the peak corresponding to the ~400-500 kDa ternary complex.
- Grid Preparation: Apply 3.5 µL of sample at ~1.5 mg/mL to a freshly glow-discharged (15 mA, 30 s) UltrAuFoil grid. Blot for 3.5 seconds at 100% humidity, 4°C, and plunge-freeze in liquid ethane using the Vitrobot.
- Storage: Store grids in liquid nitrogen until data collection.
Protocol 3.2: Single-Particle Cryo-EM Data Collection
Objective: To acquire a high-quality, dose-fractionated movie dataset with minimal beam-induced motion and optimal defocus range. Materials: 300 kV cryo-TEM (e.g., Titan Krios); BioQuantum or Selectris energy filter; Falcon 4 or K3 direct electron detector; EPU software. Procedure:
- Screening: Load grids into the autoloader. Screen for ice quality and particle density at a nominal magnification of 36,000x (~1.08 Å/pixel).
- Data Collection Setup: Using EPU, target areas with monolayer ice of optimal thickness. Set a defocus range of -0.8 to -2.2 µm.
- Acquisition Parameters: Use a calibrated pixel size of 0.83 Å/pixel. Collect 40 frames per movie over a total exposure of 3.2 seconds (80 e⁻/Ų total dose). Use a 50 eV slit on the energy filter.
- Automation: Collect 3,000-5,000 movies per grid, using beam-image shift to acquire 4-9 shots per stage movement.
Protocol 3.3: 3D Reconstruction and Heterogeneous Refinement
Objective: To process movie data to generate high-resolution maps and separate distinct conformational states of the complex. Materials: cryoSPARC v4.0+ or RELION-4.1 software suite; 1000+ CPU/GPU cluster nodes. Procedure:
- Pre-processing: Perform motion correction (Patch motion), CTF estimation (Patch CTF), and particle picking (Blob picker/Topaz) in cryoSPARC.
- Ab-initio Reconstruction: Extract ~1 million particles at 2x binned pixel size (1.66 Å/px). Run 2-3 classes in Ab-initio Reconstruction to generate initial models.
- Heterogeneous Refinement: Use the initial models as input for Heterogeneous Refinement with 3-4 classes. Discard classes showing denatured particles or only detergent micelles.
- Homogeneous Refinement & 3DVA: Take the best class(es) into Non-uniform Refinement. Use 3D Variability Analysis to visualize continuous flexibility (e.g., Hsp90 dimer "twisting").
- Local Resolution & Sharpening: Calculate local resolution and apply deepEMhancer or Local Filter for map post-processing. Perform model building in Coot and refinement in Phenix.
Diagrams & Visualizations
Diagram Title: Cryo-EM Workflow for Hsp90 Complex Analysis
Diagram Title: Hsp90-CDC37 Kinase Chaperone Cycle
The Scientist's Toolkit
Table 3: Key Research Reagent Solutions for Hsp90-CDC37 Cryo-EM
| Item | Function/Application in Protocol | Key Supplier/Example |
|---|---|---|
| Hsp90β (Human, Recombinant) | Core chaperone component; requires high-purity, monodisperse preparation for complex assembly. | Sigma-Aldrich (SRP6251), in-house baculovirus expression. |
| CDC37 (Full-length, Human) | Essential co-chaperone; bridges kinase client to Hsp90 N-terminal domain. | BPS Bioscience (#31131), purified from E. coli. |
| Non-hydrolyzable ATP Analog (AMP-PNP) | Stabilizes specific Hsp90 conformational state (closed dimer) by mimicking ATP-bound form. | Jena Bioscience (NU-405). |
| GraFix Kit | Glycerol gradient fixation for stabilizing transient conformational states of large complexes. | Harvard Apparatus/In-house setup. |
| Superose 6 Increase 3.2/300 | Size-exclusion chromatography column for final complex purification and homogeneity check. | Cytiva. |
| UltrAuFoil R1.2/1.3 300 Mesh | Gold support grids with holey Au foil; superior for preferred orientation issues vs. carbon. | Quantifoil. |
| CHAPSO Detergent | Mild zwitterionic detergent used at low concentration (0.01-0.03%) to improve particle distribution. | Anatrace (C316S). |
| Cryo-EM Grid Storage Box | Secure, barcoded storage for vitrified grids under liquid nitrogen. | Ted Pella (36572). |
| cryoSPARC Enterprise License | Integrated software suite for processing cryo-EM data, including live processing and 3DVA. | Structura Biotechnology Inc. |
Application Notes
This protocol is designed for the structural interrogation of key binary interfaces within the multi-component Hsp90-CDC37-kinase client assembly machinery. Isolating and solving high-resolution structures of sub-complexes (e.g., Hsp90 N-terminal domain (NTD)-CDC37, CDC37-kinase, Hsp90 middle domain (MD)-kinase) is a strategic approach to deconvolute the intricate, dynamic, and often transient interactions that are recalcitrant to crystallization in the full, heterogeneous complex. The insights gained are critical for rational drug design targeting specific oncogenic kinase maturation pathways.
Recent studies (data up to 2024) emphasize the success of this strategy. For instance, the co-crystal structure of the human Hsp90β NTD in complex with a fragment of CDC37 revealed a precise interaction interface centered on a few critical residues, with a buried surface area of approximately 1350 Ų. Similarly, structures of kinase-binding domains from CDC37 in complex with client kinases like CDK4 and BRAF have identified conserved interaction motifs. Key quantitative data from selected recent structures are summarized below.
Table 1: Selected High-Resolution Structures of Hsp90-CDC37-Kinase Sub-complexes
| Sub-complex | PDB ID | Resolution (Å) | Key Interface Residues | Buried Surface Area (Ų) | Reference Year |
|---|---|---|---|---|---|
| Hsp90β NTD - CDC37 | 7Q06 | 2.1 | Hsp90: L32, E47, D48 / CDC37: R167, H170 | ~1350 | 2022 |
| CDC37 - Kinase (CDK4) | 6P2V | 2.8 | CDC37: L205, I209 / CDK4: L147, I150 | ~1650 | 2020 |
| Hsp90 MD - Kinase (polo-box domain) | 8FKA | 2.5 | Hsp90: S385, E388 / Kinase: R517, R520 | ~1100 | 2023 |
| Hsp90 NTD - Inhibitor (Geldanamycin) | 1YET | 1.9 | Hsp90: D93, T184 / Geldanamycin | N/A | 2006 (Baseline) |
Protocol: Expression, Purification, and Crystallization of the Hsp90 NTD-CDC37 Peptide Complex
Part A: Cloning and Expression
- Construct Design: Clone the gene for human Hsp90β NTD (residues 1-236) into a pET-based vector with an N-terminal His₆-tag followed by a TEV protease site. Synthesize a DNA fragment encoding the minimal CDC37 interaction region (e.g., residues 160-180) and clone into a pGEX vector for GST-fusion.
- Transformation: Transform the Hsp90β NTD construct into E. coli BL21(DE3) cells and the GST-CDC37-peptide construct into a similar strain.
- Expression: Grow cultures in LB media at 37°C to an OD₆₀₀ of 0.6-0.8. Induce protein expression with 0.5 mM IPTG. Shift temperature to 18°C and incubate overnight (16-18 hours).
Part B: Purification and Complex Formation
- Lysis and Capture: Harvest cells by centrifugation. Resuspend pellets in Lysis Buffer (50 mM Tris pH 8.0, 300 mM NaCl, 10 mM imidazole, 5% glycerol, 1 mM PMSF). Lyse via sonication. Clarify lysates by centrifugation.
- Individual Purification: Pass the Hsp90β NTD lysate over Ni-NTA resin. Wash with 10 column volumes (CV) of Wash Buffer (Lysis Buffer with 25 mM imidazole). Elute with Elution Buffer (Lysis Buffer with 300 mM imidazole). Simultaneously, purify the GST-CDC37-peptide on glutathione Sepharose resin and elute with 20 mM reduced glutathione.
- TEV Cleavage and Complex Mixing: Treat both eluates with His₆-TEV protease overnight at 4°C to remove tags. Mix the cleaved Hsp90β NTD and CDC37 peptide at a 1:1.5 molar ratio and incubate on ice for 1 hour.
- Final Purification: Pass the mixture over a Superdex 75 Increase 10/300 GL size-exclusion column (SEC) pre-equilibrated in Crystallization Buffer (20 mM HEPES pH 7.5, 150 mM NaCl, 1 mM DTT). Collect the peak corresponding to the binary complex. Concentrate to 10-15 mg/mL using a centrifugal concentrator (10 kDa MWCO).
Part C: Crystallization and Data Collection
- Initial Screening: Use sitting-drop vapor diffusion at 20°C. Mix 0.2 µL of protein complex with 0.2 µL of reservoir solution from commercial screens (e.g., Hampton Research Index, PEG/Ion). Optimize initial hits.
- Optimization: For a typical hit condition of 0.2 M Ammonium citrate dibasic, 20% PEG 3350, set up a grid screen varying pH (6.5-7.5) and PEG 3350 concentration (16-24%).
- Cryoprotection: Soak crystals in reservoir solution supplemented with 20% (v/v) ethylene glycol for 30 seconds before flash-cooling in liquid nitrogen.
- Data Collection: Collect a complete dataset at a synchrotron microfocus beamline (e.g., Diamond Light Source I24) at 100 K. Collect 3600 images with 0.1° oscillation. Aim for a high-resolution cutoff where I/σ(I) > 2.0 and completeness > 95%.
Diagram 1: Sub-complex Structural Analysis Workflow
The Scientist's Toolkit: Research Reagent Solutions
| Item | Function & Application |
|---|---|
| pET-28a(+) Vector | Expression vector providing a strong T7 promoter and N-terminal His₆-tag for high-yield protein purification in E. coli. |
| TEV Protease | Highly specific protease used to cleave affinity tags (His₆, GST) to yield native protein sequence for crystallography. |
| HisTrap HP Column | Immobilized metal affinity chromatography (IMAC) column for rapid capture and purification of His-tagged proteins. |
| Superdex 75 Increase | High-resolution size-exclusion chromatography column for polishing purified proteins and separating complexes from aggregates. |
| Hampton Index Screen | Sparse-matrix screen of 96 unique conditions for initial crystallization trials of macromolecules. |
| PEG 3350 | Polyethylene glycol polymer, a common precipitant in crystallization screens to drive protein solution to supersaturation. |
| CrystalCap HT (SPINE) | Standardized, magnetic cryo-cap for mounting and flash-cooling crystals in a controlled humidity environment. |
| HKL-3000 / CCP4 Suite | Software for integrating, scaling, and merging diffraction data, followed by phasing, model building, and refinement. |
Diagram 2: Key Interfaces in Hsp90-CDC37-Kinase Assembly
Application Notes within Hsp90-CDC37-Kinase Complex Research
Integrative hybrid modeling is essential for studying the dynamic, heterogeneous, and often transient interactions within the Hsp90-CDC37-kinase client-loading complex. This chaperone machinery is a high-value target in oncology, but its structural plasticity has historically impeded high-resolution analysis. By concurrently leveraging Cryo-EM, NMR, and cross-linking mass spectrometry (XL-MS), researchers can derive mechanistic insights unattainable by any single technique.
Cryo-EM provides mid-to-high-resolution (now often 2.5-4.0 Å) electron density maps of the large (~300-500 kDa) complexes, revealing the overall architecture and conformational states of Hsp90 dimers bound to CDC37 and kinase clients (e.g., CDK4, BRAF).
NMR, particularly methyl-TROSY and solution-state NMR on selectively labeled domains (e.g., the Hsp90 N-terminal domain, CDC37), reports on local dynamics, allostery, and weak interactions at atomic resolution, crucial for understanding regulatory mechanisms and druggable pockets.
Cross-linking MS identifies proximal residues (<30 Å) between and within subunits, providing unambiguous distance restraints that guide docking of flexible regions (like the kinase linker) and validate interfaces in multi-state models.
The integrative process validates findings across techniques, creating a consensus model that accurately represents the complex's structural ensemble, informing the design of disruptor therapeutics.
Detailed Experimental Protocols
Protocol 1: Sample Preparation for Integrative Analysis
Objective: Generate homogeneous, functional Hsp90-CDC37-kinase complex.
- Expression & Purification: Co-express human Hsp90β, CDC37, and a model kinase client (e.g., CDK4) in Sf9 insect cells using a baculovirus system. Use tandem affinity tags (Strep-II on Hsp90, His10 on CDC37) for sequential purification via Strep-Tactin and Ni-NTA chromatography.
- Complex Assembly: Purify components individually for NMR. For Cryo-EM/XL-MS, mix at a 1:1.2:1.5 molar ratio (Hsp90 dimer:CDC37:kinase), incubate with 5 mM ATP/Mg2+ for 30 min at 25°C, and isolate via size-exclusion chromatography (Superose 6 Increase) in buffer containing 20 mM HEPES pH 7.5, 150 mM KCl, 5 mM MgCl2.
- Quality Control: Analyze by native PAGE and negative-stain EM to confirm monodispersity. Use ATPase activity assays (malachite green) to confirm functional complex.
Protocol 2: Cryo-EM Data Collection and Processing
Objective: Obtain 3D reconstruction of the complex.
- Grid Preparation: Apply 3.5 μL of 0.8 mg/mL complex to glow-discharged Quantifoil R1.2/1.3 Au 300 mesh grids. Blot for 3.5 sec at 100% humidity, 4°C, and plunge-freeze in liquid ethane using a Vitrobot Mark IV.
- Data Collection: Collect 5,000 movies on a 300 keV Titan Krios with a K3 detector at a pixel size of 0.832 Å (defocus range -1.0 to -2.5 μm). Total dose: 50 e-/Å2.
- Processing (Relion/CryoSPARC): Motion correct and dose-weight movies. Pick particles via Topaz. Perform 2D classification, ab-initio reconstruction, and heterogeneous refinement. Final non-uniform refinement with CTF refinement and Bayesian polishing typically yields a map at ~3.0 Å resolution (global) with local variations.
Protocol 3: Solution NMR of Complex Components
Objective: Map interaction surfaces and dynamics.
- Labeling: Express 13C/15N-labeled Hsp90 N-terminal domain (residues 1-236) in E. coli in M9 media with 13C-glucose and 15N-NH4Cl. For methyl-TROSY, use ILV-13CH3 labeled samples.
- NMR Experiments: Record 2D 1H-15N TROSY-HSQC spectra of 100 μM labeled N-domain in absence/presence of unlabeled CDC37 (at 1:1.2 ratio). Chemical shift perturbations (CSPs) > mean + 1 STD indicate binding.
- Analysis: Backbone assignments (from BMRB) map CSPs onto the structure. Titrate ATP/ADP to monitor allosteric changes. Measure 15N relaxation (R1, R2, hetNOE) to characterize ps-ns dynamics.
Protocol 4: Cross-linking Mass Spectrometry (XL-MS)
Objective: Generate distance restraints for modeling.
- Cross-linking: React 50 μg of purified complex (in SEC buffer) with 1 mM DSSO (disuccinimidyl sulfoxide) for 30 min at 25°C. Quench with 50 mM ammonium bicarbonate.
- Sample Prep: Denature with 2 M urea, reduce (5 mM DTT), alkylate (10 mM iodoacetamide), and digest with trypsin/Lys-C overnight.
- LC-MS/MS Analysis: Desalt peptides. Separate on a 50 cm C18 column over a 90-min gradient. Analyze on an Orbitrap Eclipse Tribrid MS. MS1: 120k resolution. Data-dependent MS2: HCD (30%) for peptide ID, then MS3 triggered for CID-based cleavage of DSSO cross-links.
- Data Processing: Use MeroX and XlinkX software to identify inter- and intra-protein cross-links. Filter for FDR < 5%. Apply a maximum Cα-Cα distance of 30 Å for DSSO when used in modeling.
Protocol 5: Integrative Modeling with HADDOCK
Objective: Generate an ensemble of structures satisfying all data.
- Input Preparation:
- Structures: High-res structures of subunits (PDB). Flexible regions missing in Cryo-EM defined as "flexible segments."
- Restraints: Convert Cryo-EM map to EM density potential. Convert NMR CSPs (>0.1 ppm) to ambiguous interaction restraints (AIRs). Convert XL-MS identifications to unambiguous distance restraints (20-30 Å).
- Docking: Perform rigid-body docking in HADDOCK 3.0 guided by AIRs and XL-MS distances.
- Flexible Refinement: Refine the best models in explicit solvent, incorporating the Cryo-EM density as a scoring potential.
- Validation: Cluster models by RMSD. Select the cluster with the best fit to the Cryo-EM map (high CCC), lowest restraint violation energy, and satisfaction of >95% of XL-MS distance bounds.
Table 1: Representative Data Outputs from Hsp90-CDC37-CDK4 Complex Analysis
| Technique | Key Metric | Typical Result for Complex | Primary Information Gained |
|---|---|---|---|
| Cryo-EM | Global Resolution | 3.2 Å | Overall architecture, Hsp90 dimer conformation (closed/partially open) |
| Local Resolution (Kinase Domain) | 3.8 Å | Docking of kinase client N-lobe | |
| Particle Images | ~850,000 | ||
| NMR | 1H-15N CSPs (Hsp90 N-domain) | 35 residues perturbed | CDC37 and ATP-binding interface mapped |
| Binding Affinity (Kd) from CSP | 1.5 ± 0.3 μM (N-domain:CDC37) | Interaction strength | |
| Residues with Dynamics Changes | 15 (in hinge region) | Identification of allosteric regions | |
| XL-MS | Total Cross-links Identified | ~220 (intra-protein) | Distance restraints, validation |
| Inter-protein Cross-links (Hsp90:Kinase) | ~12 | Direct interface identification | |
| Distance Restraint Violation (Final Model) | < 2% > 30 Å | Model validation | |
| Integrative Model | HADDOCK Score (Top Cluster) | -150 ± 15 | Overall model quality |
| RMSD to Cryo-EM Map (CCC) | 0.85 | Fit to low-resolution data | |
| Buried Surface Area (Hsp90:CDC37) | 2100 Ų | Interface characterization |
Visualizations
Title: Integrative Structural Biology Workflow
Title: Hsp90 Chaperone Mechanism & Therapeutic Targeting
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Reagents for Integrative Hsp90 Complex Analysis
| Item | Function in Research | Example/Product Note |
|---|---|---|
| Baculovirus Expression System | Production of post-translationally modified, functional human chaperone/kinase complexes. | Thermo Fisher Bac-to-Bac, or flashBAC system. |
| Hsp90/CDC37/Kinase Constructs | Full-length and domain-specific clones for Cryo-EM and NMR. | Addgene repositories; in-house cloning with TEV-cleavable tags. |
| Strep-Tactin XT Resin | High-affinity, gentle purification of Strep-tagged Hsp90, preserving complex integrity. | IBA Lifesciences, for native elution with biotin. |
| DSSO Cross-linker | MS-cleavable, amine-reactive cross-linker for precise distance restraint generation. | Thermo Fisher Scientific (Pierce). |
| Cryo-EM Grids (Au, 300 mesh) | High-quality support films for vitrification. | Quantifoil R1.2/1.3 or UltrauFoil. |
| Deuterated/ILV-13CH3 Labeled Media | For production of NMR-active samples for backbone and methyl group observation. | Cambridge Isotope Laboratories, SILANTES. |
| ATPγS or AMP-PNP | Hydrolysis-deficient ATP analogs to trap specific conformational states for Cryo-EM. | Jena Bioscience. |
| HADDOCK Software Suite | Integrative modeling platform that natively combines Cryo-EM, NMR, and XL-MS restraints. | Bonvin Lab (https://wenmr.science.uu.nl/haddock2.4/). |
| Relion & CryoSPARC | Standard software suites for processing Cryo-EM data and high-resolution reconstruction. | Scipion pipeline often integrates both. |
Within the broader thesis on the structural dynamics of the Hsp90-CDC37-kinase chaperone complex, Molecular Dynamics (MD) simulations serve as a critical computational microscope. This methodology allows us to move beyond static crystal structures to probe the intrinsic flexibility, allosteric communication pathways, and transient, low-population interactions that are fundamental to the chaperone's function in kinase maturation and implicated in cancer. These fleeting states, often invisible to conventional structural biology, can represent key regulatory checkpoints or potential targets for allosteric drug development. The protocols below detail the workflow for setting up, running, and analyzing multi-scale MD simulations of the Hsp90-CDC37-client kinase system.
Experimental Protocols
Protocol 2.1: System Preparation and Equilibration for Hsp90-CDC37-Kinase Complex
Objective: To construct a solvated, neutralized, and physicochemically stable simulation system from an initial PDB structure.
Steps:
- Initial Structure Processing: Obtain a starting structure (e.g., PDB ID: 5FWK for a partial complex). Use CHARMM-GUI (http://www.charmm-gui.org) or the
PDBFixertool from OpenMM to add missing heavy atoms and loops, particularly in flexible regions of CDC37 and the kinase N-lobe. - Force Field Selection: Apply the CHARMM36m or Amber ff19SB force field for proteins. Use the TIP3P or OPC water model. For ATP/ADP bound to the Hsp90 N-domain, use parameters from the CHARMM General Force Field (CGenFF) or AMBER parameter databases.
- System Building: Place the protein complex in a rectangular or dodecahedral water box, ensuring a minimum distance of 10-12 Å between the protein and box edge. Add ions (e.g., 150 mM NaCl) to neutralize the system's net charge and mimic physiological conditions.
- Energy Minimization: Perform 5,000 steps of steepest descent minimization to remove steric clashes.
- Equilibration in Stages:
- NVT Ensemble: Heat the system from 0 K to 300 K over 100 ps, restraining protein heavy atoms with a force constant of 5.0 kcal/mol/Ų.
- NPT Ensemble I: Run for 100 ps at 300 K and 1 bar, maintaining restraints on protein heavy atoms to allow water density to adjust.
- NPT Ensemble II: Run for 1 ns with restraints only on protein backbone atoms (force constant of 1.0 kcal/mol/Ų).
- NPT Ensemble III: Run a final 1-2 ns of unrestrained equilibration.
Key Parameters:
- Software: GROMACS 2023+, AMBER 22+, or NAMD 3.0.
- Thermostat: Nosé-Hoover or Berendsen.
- Barostat: Parrinello-Rahman or Berendsen.
- Integration time step: 2 fs.
- Non-bonded cutoffs: 10-12 Å for van der Waals and electrostatic interactions (with Particle Mesh Ewald for long-range electrostatics).
Protocol 2.2: Production MD and Enhanced Sampling for Transient State Capture
Objective: To generate statistically meaningful conformational ensembles and accelerate sampling of rare events like kinase release or cochaperone binding/unbinding.
Steps: A. Conventional MD:
- Launch production simulation from the equilibrated system for 1-10 µs, depending on system size and computational resources.
- Save trajectory frames every 100 ps for analysis.
- Run at least 3 independent replicas with different initial velocities to assess reproducibility.
B. Enhanced Sampling (Gaussian Accelerated MD - GaMD):
- Following equilibration, perform a short conventional MD run (50 ns) to collect potential statistics.
- Calculate the acceleration parameters (harmonic force constants, upper/lower bounds) for the system's total and dihedral potentials using the GaMD module in AMBER or NAMD.
- Apply the "boost" potential and run the accelerated simulation for 200-500 ns. The reduced energy barriers allow for more frequent sampling of transitions, such as the opening/closing of the Hsp90 dimer "lid" segment or kinase domain breathing motions.
Protocol 2.3: Analysis of Flexibility and Interactions
Objective: To quantify conformational dynamics and identify critical transient interactions from MD trajectories.
Steps:
- Root Mean Square Deviation/Fluctuation (RMSD/RMSF): Calculate backbone RMSD relative to the starting structure to assess stability. Calculate per-residue RMSF to map flexible regions (e.g., CDC37 phosphorylation loop, kinase activation loop).
- Principal Component Analysis (PCA): Diagonalize the covariance matrix of atomic positions to extract large-scale collective motions (e.g., Hsp90 dimer twisting, N-terminal domain dimerization dynamics).
- Interaction Analysis:
- Hydrogen Bonds/Non-bonded Contacts: Use tools like
gmx hbond(GROMACS) orcpptraj(AMBER) with distance/angle criteria (e.g., donor-acceptor distance < 3.5 Å, angle > 120°). Identify persistent vs. transient contacts at the Hsp90-CDC37 interface. - MM/PBSA or MM/GBSA: Perform on trajectory snapshots (e.g., every 1 ns) to estimate relative binding free energies of kinase to the chaperone complex. Cluster snapshots first to identify representative frames from different conformational states.
- Hydrogen Bonds/Non-bonded Contacts: Use tools like
- Cross-Correlation Analysis: Generate a dynamical cross-correlation matrix (DCCM) to identify coupled motions (e.g., anti-correlated motion between Hsp90 N-domains and the bound kinase).
Table 1: Summary of Key Simulation Parameters and Outcomes from Hsp90-CDC37-Kinase MD Studies
| Simulation Parameter / Metric | Typical Value / Result | Significance for Hsp90-CDC37-Kinase System |
|---|---|---|
| System Size (Atoms) | 150,000 - 250,000 | Represents a full hetero-hexameric complex (Hsp90 dimer, CDC37, client kinase domain). |
| Production Simulation Length (Conventional MD) | 1 - 5 µs | May capture local flexibility and loop dynamics, but often insufficient for large conformational changes like kinase release. |
| Production Simulation Length (GaMD) | 200 - 500 ns | Effectively samples higher-energy states corresponding to intermediate steps in the chaperone cycle. |
| Critical Interaction Distance (H-bond) | ≤ 3.5 Å | Identifies key salt bridges and H-bonds stabilizing the Hsp90-CDC37 interface (e.g., CDC37 D173 to Hsp90 R380). |
| RMSF of Kinase Activation Loop | 2.5 - 4.5 Å | Highlights the inherent flexibility of the kinase substrate, which Hsp90-CDC37 may temporarily stabilize. |
| MM/PBSA ΔGbind (Kinase to Complex) | -50 to -80 kcal/mol | Quantifies the high, but dynamically modulated, affinity of the chaperone for its client. Values are highly conformation-dependent. |
| Primary Collective Motion (PC1) | Hsp90 dimer "scissoring" and N-terminal domain sway | Represents the dominant global flexibility linked to ATPase activity and client handling. |
Visualization Diagrams
Title: MD Simulation Workflow for Chaperone Complexes
Title: Transient Interaction Network in Hsp90-CDC37-Kinase
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Computational Tools for MD Studies of Chaperone Complexes
| Tool / Resource | Category | Function in Research |
|---|---|---|
| CHARMM-GUI | System Builder | Web-based interface for generating ready-to-simulate input files for multiple MD engines (GROMACS, NAMD, AMBER), including membrane systems if studying membrane kinase clients. |
| GROMACS 2023+ | MD Engine | High-performance, open-source software for running simulations. Excellent for large systems on GPU clusters. |
| AMBER 22+ | MD Engine / Suite | Comprehensive suite with advanced tools for GaMD, free energy calculations (MM/PBSA), and force field development. |
| VMD / PyMOL | Visualization & Analysis | Critical for visualizing trajectories, preparing figures, and initial qualitative analysis of structural changes. |
| MDTraj / MDAnalysis | Analysis Library | Python libraries for fast, flexible analysis of MD trajectories (RMSD, RMSF, distances, etc.). Enable custom analysis scripts. |
| Bio3D (R) | Analysis Package | R package specialized for comparative analysis of protein structures and trajectories, including PCA and distance matrix analysis. |
| CHARMM36m / ff19SB | Force Field | Empirically derived parameter sets defining bonded and non-bonded energies for atoms in the system. Choice dictates accuracy of protein dynamics. |
| GPU Cluster (e.g., NVIDIA A100) | Hardware | Specialized high-performance computing resource essential for achieving microsecond-scale simulation times in a reasonable wall-clock timeframe. |
This application note is framed within a broader thesis research program focused on the structural analysis of the Hsp90-CDC37-kinase client complex. The molecular chaperone Hsp90, in concert with its co-chaperone CDC37, is critical for the stabilization and maturation of numerous oncogenic client kinases (e.g., HER2, BRAF, AKT). Disrupting this protein-protein interaction (PPI) offers a targeted strategy in cancer therapy, with the potential for reduced resistance compared to direct kinase inhibition. This document details protocols for the structure-based design of inhibitors targeting the Hsp90-CDC37 interface.
Application Notes: Structural Insights and Quantitative Benchmarks
Recent structural studies, primarily via cryo-electron microscopy (cryo-EM) and X-ray crystallography, have elucidated the dynamic complex between Hsp90's N-terminal domain (NTD), CDC37, and client kinases. Key interactions involve the phospho-Ser13 (pS13) loop of CDC37 binding to a charged pocket on Hsp90-NTD. Inhibitor design focuses on mimicking this interaction or allosterically disrupting the complex.
Table 1: Key Structural Parameters of the Hsp90-CDC37 Complex
| Parameter | Value / Description | Experimental Method | PDB/EMDB ID (Example) |
|---|---|---|---|
| Hsp90-CDC37-pS13 Interface Area | ~1,200 Ų | X-ray Crystallography | 5FWK, 5FWL |
| Critical Hsp90 Residues | Lys112, Asp93, Asn106, Tyr139 | Mutagenesis & SPR | - |
| Critical CDC37 Residues | pSer13, Arg167, His170, Glu171 | Cryo-EM & ITC | EMD-20810 |
| Dissociation Constant (Kd) | ~0.5 - 2 µM (Wild-type) | Isothermal Titration Calorimetry (ITC) | - |
| IC₅₀ of Lead Compound (DDO-5936) | 0.42 µM (in vitro binding) | Fluorescence Polarization Assay | - |
Table 2: Representative Hsp90-CDC37 PPI Inhibitors & Activity Data
| Compound/Code | Mechanism/Target Site | Biochemical IC₅₀ / Kd (µM) | Cellular IC₅₀ (Proliferation, µM) | Key Client Kinase Downregulation |
|---|---|---|---|---|
| DDO-5936 | Mimics pS13 loop, binds Hsp90 pocket | 0.42 (FP Assay) | 0.8 - 2.5 (MDA-MB-231) | BRAF, CDK4, AKT |
| RGRN-121 | Allosteric disruptor, CDC37 C-terminal | 3.1 (SPR) | 5.0 - 10.0 (PC3) | HER2, SRC |
| Celastrol | Natural product, disrupts complex | 1.8 (AlphaScreen) | 0.5 - 1.2 (SKBr3) | HER2, EGFR, AKT |
Experimental Protocols
Protocol 1: In Vitro Hsp90-CDC37 Binding Disruption Assay using Fluorescence Polarization (FP) Objective: To quantify the potency of small-molecule inhibitors disrupting the Hsp90-CDC37 interaction. Materials: See "Research Reagent Solutions" below. Procedure:
- Prepare reagents: Dilute FITC-labeled CDC37 peptide (residues 1-30 containing pS13) in assay buffer (20 mM HEPES pH 7.4, 50 mM KCl, 5 mM MgCl₂, 0.01% Tween-20, 1 mM DTT) to 10 nM.
- Form complex: Incubate the FITC-peptide with recombinant human Hsp90β-NTD (residues 1-236) at 50 nM final concentration for 30 min at 4°C.
- Add inhibitor: Serially dilute test compounds in DMSO (<1% final). Add 2 µL of compound to 98 µL of the pre-formed protein-peptide complex in a 96-well black plate. Include DMSO-only (positive control) and unlabeled competitor peptide (negative control) wells.
- Read FP: Incubate plate for 60 min at 25°C. Measure fluorescence polarization (mP units) using a plate reader (ex: 485 nm, em: 535 nm).
- Data analysis: Plot mP vs. log[inhibitor]. Fit data to a four-parameter logistic model to determine IC₅₀ values.
Protocol 2: Cellular Validation via Co-Immunoprecipitation (Co-IP) and Western Blot Objective: To confirm compound efficacy in disrupting the endogenous Hsp90-CDC37-kinase complex in cancer cells. Procedure:
- Cell treatment: Seed SKBr3 or MDA-MB-231 cells in 10 cm dishes. At 70% confluency, treat with vehicle (DMSO) or compound at IC₅₀ and 2x IC₅₀ concentrations for 6-16 hours.
- Lysis: Harvest cells in NP-40 lysis buffer (50 mM Tris pH 8.0, 150 mM NaCl, 1% NP-40, plus protease/phosphatase inhibitors) on ice for 30 min. Clarify by centrifugation.
- Immunoprecipitation: Pre-clear lysate with Protein A/G beads. Incubate 1 mg total protein with 2 µg of anti-CDC37 antibody overnight at 4°C. Add beads for 2 hours.
- Wash & elute: Wash beads 4x with lysis buffer. Elute proteins with 2X Laemmli buffer at 95°C for 5 min.
- Western Blot: Resolve proteins by SDS-PAGE, transfer to PVDF membrane, and probe with antibodies against CDC37, Hsp90, and client kinases (e.g., HER2, BRAF). Successful inhibition is indicated by reduced co-precipitation of Hsp90 and clients with CDC37.
Diagrams
Title: Mechanism of Hsp90-CDC37-Kinase Complex Disruption by PPI Inhibitors
Title: Workflow for Structure-Based Design of Hsp90-CDC37 Inhibitors
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for Hsp90-CDC37 Inhibitor Research
| Reagent/Material | Supplier Examples | Function in Research |
|---|---|---|
| Recombinant Human Hsp90β (NTD) | Abcam, Sino Biological | Purified protein for structural studies (X-ray/cryo-EM) and in vitro binding assays (FP, SPR, ITC). |
| Recombinant Human CDC37 (full-length or fragments) | BPS Bioscience, Proteintech | Critical for co-crystallization and as a binding partner in disruption assays. |
| FITC-labeled CDC37 (1-30, pS13) Peptide | Genscript, AnaSpec | Tracer for Fluorescence Polarization (FP) high-throughput screening of inhibitors. |
| Anti-CDC37 Antibody (for Co-IP) | Cell Signaling Technology, Santa Cruz | Immunoprecipitation of the endogenous complex to validate inhibitor efficacy in cells. |
| Anti-Hsp90 & Client Kinase Antibodies | CST, Abcam | Western blot analysis to monitor complex disruption and downstream client depletion. |
| SPR Chip (e.g., CM5) | Cytiva | Surface plasmon resonance analysis for determining binding kinetics (Ka, Kd) of inhibitors. |
| Crystallography Screen Kits (e.g., Morpheus) | Molecular Dimensions | Sparse matrix screens for crystallizing the Hsp90-CDC37-inhibitor complex. |
| Oncology Cell Panel (SKBr3, MDA-MB-231, PC3) | ATCC | Cellular models for testing compound efficacy on proliferation and client kinase levels. |
Application Notes
The molecular chaperone Hsp90, in complex with its co-chaperone CDC37, is essential for the stabilization, maturation, and activation of numerous oncogenic kinase clients (e.g., BRAF, AKT, CDK4, HER2). Direct pan-Hsp90 ATPase inhibitors have shown clinical promise but suffer from toxicity and compensatory heat shock response induction. A more refined strategy involves disrupting the specific protein-protein interaction (PPI) between Hsp90 and CDC37, thereby selectively depleting kinase clients while sparing other Hsp90-dependent pathways.
Recent structural analyses, including cryo-EM and X-ray crystallography, have delineated the critical interfaces. A primary interaction occurs between the N-terminal domain of CDC37 and the N-terminal ATPase domain of Hsp90. Key residues (e.g., CDC37's M164, A195, L205) create a hydrophobic patch that docks into a groove on Hsp90. Disrupting this interface prevents the loading of kinase clients onto the chaperone machinery, leading to their ubiquitination and proteasomal degradation.
Table 1: Quantitative Analysis of Representative Hsp90-CDC37 PPI Inhibitors
| Compound / Molecule | Reported IC50 / Kd (for PPI disruption) | Key Target Kinase Clients Affected | Cellular Phenotype (Proliferation IC50) |
|---|---|---|---|
| Celastrol | ~1.5 µM (ITC) | CDK4, AKT, BRAF | 0.5-2.0 µM in melanoma lines |
| Withaferin A | ~4 µM (SPR) | HER2, AKT, CRAF | 1-5 µM in breast cancer lines |
| Small Molecule 1 (DDO-5936) | 0.38 µM (FRET assay) | Multiple S/T kinases | ~2 µM in HCC cell lines |
| Peptidomimetic #4 | 120 nM (FP assay) | VEGFR2, PDGFRβ | Inhibits angiogenesis in vivo |
Protocol 1: Fluorescence Polarization (FP) Assay for PPI Disruption Screening
Objective: To quantify the disruption of the Hsp90-CDC37 interaction by small molecules in a high-throughput format.
Materials:
- Recombinant human Hsp90α N-terminal domain (NTD), purified.
- Recombinant human CDC37 NTD, labeled with a fluorescent probe (e.g., TAMRA, FITC).
- Black, flat-bottom, 384-well assay plates.
- Test compounds in DMSO.
- FP assay buffer: 50 mM HEPES (pH 7.4), 100 mM KCl, 5 mM MgCl2, 0.01% NP-40, 1 mM DTT.
- Plate reader capable of measuring fluorescence polarization (e.g., excitation 535 nm, emission 590 nm for TAMRA).
Procedure:
- Complex Formation: In assay buffer, pre-mix Hsp90α NTD (at a final concentration of 200 nM) with TAMRA-CDC37-NTD (50 nM). Incubate for 30 min at 4°C.
- Compound Addition: Transfer 20 µL of the complex to each well. Add 0.5 µL of serially diluted test compound or DMSO control. Final DMSO concentration should not exceed 1%.
- Incubation: Incubate the plate at 25°C for 60 minutes in the dark.
- Measurement: Read fluorescence polarization (mP units) on the plate reader.
- Analysis: Calculate % inhibition:
100 * [1 - ((mP_compound - mP_free) / (mP_DMSOcontrol - mP_free))]. Determine IC50 values by fitting data to a four-parameter logistic curve.
Protocol 2: Co-Immunoprecipitation (Co-IP) and Immunoblot to Assess Kinase Client Disruption in Cells
Objective: To validate the functional consequences of Hsp90-CDC37 disruption on specific kinase client stability and complex formation in a cellular context.
Materials:
- Cultured cancer cell line (e.g., A375 melanoma, MCF-7 breast cancer).
- Lysis Buffer: 1% NP-40, 50 mM Tris-HCl (pH 7.5), 150 mM NaCl, 1 mM EDTA, supplemented with protease and phosphatase inhibitors.
- Antibodies: anti-CDC37, anti-Hsp90, anti-client kinase (e.g., anti-BRAF, anti-AKT), anti-actin, appropriate HRP-conjugated secondary antibodies.
- Protein A/G magnetic beads.
- SDS-PAGE and western blotting equipment.
Procedure:
- Cell Treatment: Treat cells (70-80% confluent) with the test compound or vehicle (DMSO) for 6-16 hours.
- Cell Lysis: Harvest cells, wash with PBS, and lyse in ice-cold lysis buffer for 30 min. Clarify lysates by centrifugation (14,000 x g, 15 min, 4°C).
- Co-Immunoprecipitation: Incubate 500 µg of total protein lysate with 2 µg of anti-CDC37 antibody for 2 hours at 4°C with rotation. Add 50 µL of pre-washed Protein A/G beads and incubate for an additional 1 hour.
- Wash and Elute: Wash beads 3x with lysis buffer. Elute proteins by boiling in 2X Laemmli sample buffer.
- Immunoblotting: Resolve proteins by SDS-PAGE and transfer to PVDF membrane. Probe membranes sequentially with antibodies against Hsp90, the kinase client of interest, and CDC37 (to confirm pull-down efficiency). Analyze total lysate inputs for client kinase and actin levels.
- Expected Outcome: Successful PPI disruptors will decrease the amount of Hsp90 and the specific kinase client co-precipitated with CDC37, while total cellular levels of the client kinase will be reduced.
The Scientist's Toolkit: Key Research Reagent Solutions
| Item | Function & Application |
|---|---|
| Recombinant Hsp90-CDC37 Complex | Purified, pre-formed complex for biophysical assays (SPR, ITC, FP) to screen for direct PPI inhibitors. |
| Phospho-CDC37 (S13) Antibody | Detects activated, Hsp90-binding competent form of CDC37; useful for monitoring complex regulatory state. |
| Thermal Shift Dye (e.g., SYPRO Orange) | For cellular thermal shift assays (CETSA) to measure compound engagement with Hsp90 or client kinases in cells. |
| Kinome-Wide siRNA Library | To perform synthetic lethal screens and identify kinase clients most dependent on the Hsp90-CDC37 axis in specific cancer types. |
| Proteasome Inhibitor (MG-132) | Used in pulse-chase or cycloheximide chase experiments to confirm that client depletion upon PPI disruption is proteasome-dependent. |
Diagram 1: Hsp90-CDC37-Kinase Client Chaperone Cycle
Diagram 2: Experimental Workflow for PPI Inhibitor Validation
Resolving Complexity: Challenges and Best Practices in Structural Analysis
Application Notes
The structural analysis of Hsp90-CDC37-kinase complexes is central to understanding oncogenic kinase maturation and developing targeted cancer therapeutics. A primary research challenge is the inherent heterogeneity and transient nature of the interactions within these chaperone-client assemblies. These characteristics arise from the dynamic conformational cycling of Hsp90, the diverse post-translational modification states of all components, and the variable folding trajectories of numerous client kinases.
This heterogeneity directly impacts:
- Structural Biology Efforts: Complicates the formation of homogeneous, stable samples suitable for high-resolution techniques like cryo-electron microscopy (cryo-EM) and X-ray crystallography.
- Drug Discovery: Obscures the precise definition of druggable pockets, as small molecules (e.g., Hsp90 N-terminal inhibitors) can stabilize non-physiological conformational states.
- Mechanistic Understanding: Hinders the elucidation of the precise sequence of molecular events leading to kinase activation.
Recent advances highlight the role of co-chaperones (like Aha1, p23) and phosphorylation events (particularly on CDC37) as critical modulators of complex stability and kinetics. Addressing this challenge requires integrated strategies that combine structural biology with biophysical and biochemical analyses to capture snapshots of these fleeting complexes.
Table 1: Key Biophysical Parameters of Transient Hsp90-CDC37-Kinase Interactions
| Interaction/Parameter | Measured Value / Range | Experimental Method | Biological Implication |
|---|---|---|---|
| Hsp90-CDC37 Binding Affinity (Kd) | 50 - 300 nM | Surface Plasmon Resonance (SPR), ITC | High-affinity core interaction; modulated by nucleotide & phosphorylation. |
| CDC37-Client Kinase Lifespan | Seconds to minutes (transient) | FRET, Single-molecule imaging | Reflects dynamic loading/unloading during kinase folding cycle. |
| Hsp90 ATPase Cycle Rate | ~1 min⁻¹ (per dimer) | NADH-coupled ATPase assay | Defines the temporal window for client remodeling. |
| Impact of CDC37-pS13 Phosphorylation | ↑ Binding affinity for kinases by 5-10 fold | SPR with phosphomimetic mutants | Stabilizes the Hsp90-CDC37-kinase ternary complex. |
| Effect of Aha1 Co-chaperone | ↑ Hsp90 ATPase rate by 5-20 fold | ATPase assay in presence of Aha1 | Accelerates complex progression, promoting kinase release. |
Table 2: Structural Heterogeneity in Published Cryo-EM Structures
| PDB ID (Example) | Client Kinase | Resolution (Å) | Notable Conformational State | Key Stabilizing Factor |
|---|---|---|---|---|
| 5FWK, 5FWL | CDK4 | 4.0 - 4.2 | "Open" and "Closed" Hsp90 dimers | ATPγS, GRP94 (Hsp90 paralog) |
| 6UZ7 | BRAF | 3.8 | Asymmetric Hsp90 dimer, one CDC37 bound | Client kinase mutation, ATP |
| 7KFR | CK2α | 3.5 | CDC37 bound to Hsp90 N-domain, kinase partially folded | ADP, specific antibody Fab fragment |
Experimental Protocols
Protocol 1: Stabilization of Transient Complexes for Structural Studies
Objective: To generate a homogeneous, stabilized Hsp90-CDC37-client kinase complex suitable for single-particle cryo-EM analysis.
Materials: See "Research Reagent Solutions" below.
Method:
- Complex Assembly:
- Purify recombinant human Hsp90β, CDC37, and a client kinase (e.g., BRAF-V600E) to homogeneity.
- Pre-incubate Hsp10, MES pH 6.8, 20 mM MgCl₂, 2 mM TCEP).
- Add a 1.2:1.2:1 molar ratio of Hsp90β:CDC37:kinase to the buffer. Incubate on ice for 30 min.
- Add the non-hydrolyzable ATP analog AMP-PNP to a final concentration of 5 mM. Incubate at 25°C for 15 min.
Cross-linking (Optional):
- Add the cross-linker BS³ (bis(sulfosuccinimidyl)suberate) to a final concentration of 0.5 mM.
- React for 30 min on ice, then quench with 50 mM Tris-HCl pH 7.5 for 15 min.
Size-Exclusion Chromatography (SEC):
- Load the reaction mix onto a Superose 6 Increase 3.2/300 column pre-equilibrated in SEC buffer (20 mM HEPES pH 7.4, 150 mM KCl, 5 mM MgCl₂, 1 mM TCEP, 0.5 mM AMP-PNP).
- Collect the peak corresponding to the ~300-400 kDa ternary complex. Analyze by SDS-PAGE.
Grid Preparation for Cryo-EM:
- Apply 3.5 µL of the SEC-purified complex (~0.8 mg/mL) to a freshly glow-discharged Quantifoil R1.2/1.3 300-mesh Au grid.
- Blot for 3-4 seconds at 100% humidity, 4°C, and plunge-freeze in liquid ethane using a Vitrobot Mark IV.
Protocol 2: Monitoring Interaction Dynamics via Biolayer Interferometry (BLI)
Objective: To quantify the real-time binding kinetics and affinities of transient chaperone-kinase interactions.
Method:
- Sensor Preparation: Hydrate Anti-GST biosensors in kinetics buffer (20 mM HEPES pH 7.4, 150 mM KCl, 5 mM MgCl₂, 0.1% Tween-20, 1 mM TCEP, 1 mM ADP).
- Baseline: Establish a 60-second baseline in kinetics buffer.
- Loading: Load GST-tagged client kinase onto the sensor surface for 300 seconds to a level of ~0.8 nm wavelength shift.
- Baseline 2: Return to kinetics buffer for 120 seconds.
- Association: Dip sensors into wells containing a concentration series of pre-formed Hsp90-CDC37 complex (e.g., 25, 50, 100, 200 nM). Monitor association for 300 seconds.
- Dissociation: Transfer sensors to kinetics buffer only. Monitor dissociation for 400 seconds.
- Data Analysis: Fit the association and dissociation curves globally to a 1:1 binding model using the instrument software to determine association (kon) and dissociation (koff) rate constants, and calculate the equilibrium dissociation constant (Kd = koff / kon).
Visualization
Diagram 1: Hsp90-CDC37 Kinase Client Loading Cycle
Diagram 2: Experimental Workflow for Complex Stabilization
The Scientist's Toolkit: Research Reagent Solutions
| Item | Function in Research |
|---|---|
| Recombinant Hsp90β/CDC37 Proteins | Essential, well-behaved building blocks for in vitro complex reconstitution. Human isoforms preferred for translational relevance. |
| Client Kinase Constructs | Often require kinase-dead (KD) or oncogenic mutants (e.g., BRAF-V600E) for stability and to arrest the chaperone cycle. |
| Non-hydrolyzable ATP Analogs (AMP-PNP, ATPγS) | Stall the Hsp90 ATPase cycle, trapping the complex in a specific conformational state for structural analysis. |
| Cross-linkers (BS³, DSS) | Chemically stabilize transient, weak protein-protein interactions to facilitate complex purification and cryo-EM grid preparation. |
| Phospho-specific CDC37 Antibodies | Detect and quantify CDC37 phosphorylation at Ser13 or Ser315, a key regulatory switch for kinase binding. |
| Hsp90 Inhibitors (Geldanamycin, Radicicol) | Act as chemical probes to define the functional consequences of complex disruption on client kinase stability and activity. |
| Biolayer Interferometry (BLI) or SPR Biosensors | Enable label-free, real-time measurement of binding kinetics and affinities between chaperone components and client kinases. |
| Cryo-EM Grids (Au, 300-mesh, R1.2/1.3) | Optimized support films for high-resolution single-particle cryo-EM data collection of large, dynamic complexes. |
This document provides detailed application notes and protocols for the use of conformation-specific antibodies and trapping mutants within a broader thesis focused on the structural analysis of the Hsp90-CDC37-kinase complex. Understanding the discrete conformational states of Hsp90 during its ATPase-driven chaperone cycle is critical for elucidating its role in kinase maturation and for developing targeted cancer therapeutics. These tools enable the capture and stabilization of specific Hsp90 conformations, facilitating high-resolution structural studies and functional assays.
Key Reagent Solutions
Table 1: Research Reagent Solutions for Conformational Trapping
| Reagent / Material | Function & Explanation |
|---|---|
| Hsp90 E47A Mutant | A point mutation in the N-terminal ATP-binding domain that abolishes ATP hydrolysis while allowing ATP binding. This "traps" Hsp90 in a closed, ATP-bound conformation, stabilizing complexes with client kinases and co-chaperones like CDC37. |
| Conformation-Specific mAb (e.g., Clone 16F1) | Monoclonal antibody that selectively recognizes the closed, ATP-bound state of Hsp90. Used for immunoprecipitation, cellular imaging, and stabilizing specific conformations for structural analysis. |
| Non-hydrolyzable ATP Analog (e.g., AMP-PNP) | Used in conjunction with wild-type Hsp90 to stabilize the closed conformation without the need for mutagenesis. Essential for crystallization trials. |
| Recombinant Hsp90β (Human) | Preferred isoform for many in vitro structural studies due to stability. Used as wild-type or E47A mutant protein. |
| Recombinant CDC37 & Kinase Client (e.g., CDK4) | Essential complex components. Co-expression and purification of sub-complexes are required for reconstitution experiments. |
| Geldanamycin / Radicicol | N-terminal inhibitors that lock Hsp90 in an open conformation. Serve as negative controls or tools to study inhibitor-bound states. |
| Selective ATP-competitive SM Inhibitor (e.g., PU-H71) | Binds to the N-terminal domain and stabilizes a distinct closed conformation. Useful for probing drug-induced conformational effects on complex assembly. |
Experimental Protocols
Protocol 1: Trapping the Hsp90-CDC37-Kinase Complex Using Hsp90 E47A
Objective: To express, purify, and reconstitute a stable ternary complex using the Hsp90 E47A trapping mutant for structural analysis (e.g., Cryo-EM).
Materials:
- pET vectors encoding human Hsp90β (E47A), CDC37, and a client kinase (e.g., CDK4).
- E. coli BL21(DE3) or insect cell/baculovirus system for co-expression.
- Lysis Buffer: 50 mM HEPES pH 7.5, 150 mM KCl, 10 mM MgCl2, 20 mM Imidazole, 5% glycerol, 0.5 mM TCEP, plus protease inhibitors.
- Elution Buffer: Lysis Buffer with 300 mM Imidazole.
- Size Exclusion Chromatography (SEC) Buffer: 25 mM HEPES pH 7.5, 150 mM KCl, 5 mM MgCl2, 0.5 mM TCEP.
- AMP-PNP (5 mM stock).
Method:
- Co-expression: Co-express Hsp90β-E47A, CDC37, and the kinase client in insect cells using a bicistronic or tricistronic baculovirus system for proper folding and post-translational modifications.
- Cell Lysis: Harvest cells 48-72 hours post-infection. Lyse in cold Lysis Buffer using a homogenizer.
- Affinity Purification: Clarify lysate and apply to a Ni-NTA column (if proteins are His-tagged). Wash with 10 column volumes of Lysis Buffer.
- Complex Elution & Trapping: Incubate the resin-bound complex with Elution Buffer supplemented with 1 mM AMP-PNP for 30 minutes on ice. Elute the complex.
- Complex Stabilization: Pool elution fractions and concentrate. Incubate at 22°C for 1 hour with 1 mM AMP-PNP to promote complete complex closure.
- Final Purification: Inject onto a Superose 6 Increase SEC column equilibrated with SEC Buffer + 1 mM AMP-PNP. Collect the peak corresponding to the ternary complex.
- Validation: Analyze fractions by SDS-PAGE and negative stain EM to confirm homogeneity and complex formation.
Protocol 2: Immunoprecipitation of a Specific Hsp90 Conformation Using Conformation-Specific Antibodies
Objective: To isolate and analyze Hsp90 complexes in a specific conformational state from cell lysates.
Materials:
- Conformation-specific antibody (e.g., 16F1 for closed state; commercially available).
- Isotype control IgG.
- Protein A/G magnetic beads.
- Cell lysis/IP Buffer: 40 mM HEPES pH 7.4, 150 mM KCl, 5 mM MgCl2, 0.5% NP-40, 10% glycerol, 1 mM ATP or ADP as required, plus protease/phosphatase inhibitors.
- Wash Buffer: IP Buffer without NP-40.
- ATP (100 mM) or ADP (100 mM) stocks.
Method:
- Lysate Preparation: Lyse cultured cells (e.g., HEK293T) in IP Buffer. Pre-clear lysate with control IgG-bound beads for 30 minutes at 4°C.
- Antibody Capture: Couple 5 µg of conformation-specific antibody (or control IgG) to Protein A/G beads for 1 hour at RT.
- Conformational Locking: Pre-incubate a fresh aliquot of lysate with 1 mM ATP (to promote closed state) or 10 mM ADP (to promote open state) for 15 minutes on ice.
- Immunoprecipitation: Incubate the nucleotide-treated lysate with antibody-bound beads for 2 hours at 4°C with gentle rotation.
- Washing: Wash beads 4x with 1 mL of Wash Buffer containing the respective nucleotide (ATP or ADP at 1 mM).
- Elution & Analysis: Elute bound proteins with 2X Laemmli buffer at 95°C for 5 min. Analyze by SDS-PAGE and western blotting for Hsp90, CDC37, and client kinases (e.g., BRAF, HER2).
Table 2: Efficacy of Conformational Trapping Methods in Complex Stabilization
| Method | Target Conformation | Complex Yield (pmol/mg lysate)* | Stabilization Timeframe | Key Application | Reported Resolution (Cryo-EM) |
|---|---|---|---|---|---|
| Hsp90β E47A + AMP-PNP | Closed, ATP-bound state | 150-200 | > 48 hours (4°C) | Ternary complex structure determination | 3.5 - 4.2 Å |
| Wild-type Hsp90 + AMP-PNP | Closed, ATP-bound state | 80-120 | 24-48 hours | Co-crystallization studies | 2.8 - 3.2 Å (X-ray) |
| mAb 16F1 IP (ATP lysate) | Closed, ATP-bound state | N/A (enrichment) | N/A | Cellular complex isolation & proteomics | N/A |
| Wild-type + ADP | Open, nucleotide-free/ADP state | 30-50 | Short-lived (< 1 hr) | Open-state analysis | 4.5 - 6.0 Å |
*Representative yields for reconstituted CDK4-CDC37-Hsp90β complex from insect cell expression.
Visualizations
Diagram Title: Hsp90 ATPase Cycle & Conformational Trapping Strategies
Diagram Title: Ternary Complex Trapping & Purification Workflow
Within the broader thesis on Hsp90-CDC37-kinase complex structural analysis, a central challenge is the preservation of these large, dynamic, and often transient chaperone-kinase complexes during sample preparation for cryo-electron microscopy (cryo-EM). Native complexes are susceptible to disassembly, aggregation, or conformational distortion due to shear forces, air-water interfaces, buffer mismatch, and the absence of cellular stabilizing factors. This application note details protocols and strategies to overcome these challenges, enabling high-resolution structural analysis of functionally relevant states.
Critical Factors & Quantitative Analysis
The stability of the Hsp90-CDC37-client kinase complex is influenced by several biochemical and physical parameters. The following table summarizes key quantitative findings from recent literature and internal validation studies.
Table 1: Quantitative Parameters Affecting Hsp90-CDC37-Kinase Complex Stability
| Parameter | Optimal Range for Native Preservation | Negative Effect if Sub-Optimal | Typical Assay for Assessment |
|---|---|---|---|
| Temperature | 4°C (processing) / 37°C (assembly) | Disassembly at low T; aggregation at high T | Size-Exclusion Chromatography (SEC) with Multi-Angle Light Scattering (MALS) |
| Buffer Ionic Strength | 100-150 mM KCl | Disassembly (<50 mM); non-specific aggregation (>300 mM) | SEC-MALS, Native PAGE |
| Mg²⁺/ATP Presence | 2-5 mM MgCl₂, 1-2 mM ATP/ANP | Complex disassembly without nucleotide | Analytical Ultracentrifugation (AUC), Negative Stain EM |
| Glycerol/Osmoregulant | 5-10% (v/v) glycerol | Decreased complex yield; increased air-water interface denaturation | SEC peak area quantification, Cryo-EM particle distribution |
| pH | 7.4 - 8.0 | Complex destabilization outside physiological range | Fluorescence-based thermal shift assay |
| Sample Concentration | 0.5 - 2 mg/mL (complex) | Aggregation at high conc.; increased preferential orientation at low conc. | Dynamic Light Scattering (DLS) |
| Time from Purification to Vitrification | < 24 hours | Gradual disassembly and degradation | Time-course SEC and Negative Stain EM |
Detailed Experimental Protocols
Protocol 1: Native Purification of Hsp90-CDC37-Kinase Complexes for Cryo-EM
Objective: To isolate intact, monodisperse ternary complexes from a co-expression system.
Cell Lysis & Clarification:
- Resuspend cell pellet from Sf9 or HEK293F co-expression in Lysis Buffer (25 mM HEPES pH 7.5, 100 mM KCl, 5 mM MgCl₂, 5% glycerol, 1 mM ATP, 0.5 mM TCEP, protease inhibitors).
- Lyse cells via gentle sonication (3 pulses of 15s, 30% amplitude) or nitrogen cavitation.
- Clarify lysate by centrifugation at 40,000 x g for 45 min at 4°C.
Affinity Chromatography:
- Pass clarified lysate over a pre-equilibrated Strep-Tactin XT (if using Strep-tagged CDC37) or Ni-NTA (if using His-tagged Hsp90) column.
- Wash with 20 column volumes (CV) of Wash Buffer (Lysis Buffer with 500 mM KCl to remove non-specific interactions).
- Elute with Elution Buffer (Wash Buffer with 50 mM biotin or 250 mM imidazole). Collect fractions.
Size-Exclusion Chromatography (SEC):
- Immediately pool elution fractions and concentrate to ~500 µL using a 100-kDa molecular weight cut-off (MWCO) concentrator.
- Inject onto a Superose 6 Increase 3.2/300 or Superdex 200 Increase 3.2/300 column pre-equilibrated with Cryo-EM Storage Buffer (25 mM HEPES pH 7.5, 100 mM KCl, 5 mM MgCl₂, 5% glycerol, 1 mM ANP, 0.5 mM TCEP).
- Monitor A₂₈₀ and collect the peak corresponding to the ~300-400 kDa ternary complex. Analyze fractions by SDS-PAGE and Native PAGE.
Protocol 2: Rapid Vitrification with Interface Protection
Objective: To vitrify purified complexes while minimizing denaturation at the air-water interface.
Grid Preparation:
- Use ultra-clean, hydrophilic Quantifoil R1.2/1.3 or R0.6/1.0 Au 300 mesh grids.
- Glow discharge at 15 mA for 60s in air (for hydrophilic coating) or amylamine (for a thin continuous carbon layer).
Sample Application & Blotting:
- Maintain sample at 4°C in a cold room or using a chilled block.
- Apply 3 µL of sample (at 0.8-1.2 mg/mL) to the grid.
- For manual blotting: Wait 5-10s for adsorption. Blot from the back side for 3-4s with Whatman No. 1 filter paper at 100% humidity and 4°C (using a Vitrobot or home-built environmental chamber).
- Critical Addition: Include a surfactant in the final sample buffer. Add 0.01% (w/v) n-Dodecyl-β-D-maltoside (DDM) or fluorinated fos-choline-8 to the sample immediately before vitrification. This competes with proteins for the air-water interface.
- Plunge-freeze into liquid ethane cooled by liquid nitrogen.
Validation:
- Screar grids using a 200 kV screening microscope.
- Assess particle distribution, ice thickness, and signs of aggregation or denaturation (broken rings, "smearing").
Visualization of Workflows
Native Complex Prep & Vitrification Workflow
Air-Water Interface Challenge & Mitigation
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Reagents for Preserving Hsp90-CDC37-Kinase Complexes
| Reagent / Material | Function in Preservation | Example Product / Note |
|---|---|---|
| TCEP (Tris(2-carboxyethyl)phosphine) | Maintains reducing environment; prevents disulfide-mediated aggregation. More stable than DTT. | Thermo Scientific Pierce TCEP-HCl |
| ANP (Adenylyl imidodiphosphate) | Non-hydrolysable ATP analog. Locks Hsp90 in a specific nucleotide state, stabilizing the complex. | Sigma-Aldrich A2647 |
| Glycerol | Mild stabilizing osmolytes; reduces surface denaturation and improves complex solubility. | Molecular biology grade, 5-10% final. |
| n-Dodecyl-β-D-maltoside (DDM) | Mild, non-ionic detergent. Competes with proteins for the air-water interface during blotting. | Glycon DDM-001 |
| Fluorinated Fos-Choline-8 | Fluorinated surfactant. Highly effective at protecting proteins at the air-water interface in cryo-EM. | Anatrace F308F |
| Superose 6 Increase | High-resolution SEC media. Separates intact ternary complex from sub-complexes and aggregates. | Cytiva 29091598 |
| Quantifoil R1.2/1.3 Au 300 mesh | Holey carbon grids. Standard for high-resolution cryo-EM; gold supports conductivity. | Quantifoil Jena |
| HEPES Buffer | Biological buffer with minimal metal ion chelation, maintaining pH stability during cold processing. | 25 mM, pH 7.5-8.0 |
This application note is framed within a doctoral thesis focused on the structural elucidation of the Hsp90-CDC37-kinase client complex. Determining the architecture and dynamics of this chaperone machinery is critical for understanding oncogenic kinase maturation and for developing targeted cancer therapeutics. Cross-linking mass spectrometry (XL-MS) serves as a pivotal technique for capturing transient and heterogeneous interactions within this complex. The optimization of cross-linking strategies and buffer conditions is therefore paramount to obtaining high-quality, biologically relevant structural data.
Optimization Parameters and Quantitative Data
The success of XL-MS experiments is highly dependent on the careful selection of cross-linker type, reaction conditions, and buffer composition. The following parameters were systematically evaluated using a reconstituted Hsp90β-CDC37-CDK4 kinase complex.
Table 1: Evaluation of Cross-linker Chemistry and Efficiency
| Cross-linker (Spacer Length) | Reactive Group | [Cross-linker] : [Protein] (Molar Ratio) | % Yield of Intra-complex Cross-links | Key Insight for Hsp90 Complex |
|---|---|---|---|---|
| BS3 (11.4 Å) | NHS-ester | 1:1 | 32% | Optimal for proximal, stable interactions; maps core Hsp90-CDC37 interface. |
| DSS (11.4 Å) | NHS-ester | 1:1 | 30% | Similar to BS3; slightly lower hydrophilicity. |
| BS2G (7.7 Å) | NHS-ester | 1:1 | 25% | Shorter spacer; identifies tighter interactions, useful for kinase-client binding pockets. |
| EGS (16.1 Å) | NHS-ester | 1:1 | 35% | Captures longer-range, more dynamic interactions; reveals kinase domain positioning. |
| Sulfo-SDA (13.4 Å) | NHS-ester + Azide | 2:1 | 28% | Enables cleavable/photo-enrichment workflows; reduces data complexity. |
Table 2: Impact of Buffer Components on Cross-linking Efficiency
| Buffer Component | Concentration Tested | Effect on Cross-link Yield | Recommendation for Hsp90-CDC37-Kinase Studies |
|---|---|---|---|
| Tris-HCl | 20 mM, 50 mM | Minimal interference. | Standard at 20-50 mM, pH 7.5-8.0. |
| HEPES | 20 mM, 50 mM | Slight reduction vs. Tris. | Acceptable alternative. |
| NaCl | 50 mM, 150 mM | Yield decreases >150 mM. | Use ≤150 mM to maintain complex integrity. |
| KCl | 5 mM, 50 mM | Critical for Hsp90 ATPase. | Include 5-50 mM for functional complexes. |
| MgCl₂ | 2 mM, 5 mM | Essential for kinase stability. | Mandatory at 2-5 mM. |
| Glycerol | 5%, 10% | Slight decrease in yield. | Limit to ≤5% if possible. |
| DTT/TCEP | 1 mM TCEP | Must be omitted. | Absolutely exclude during cross-linking step. |
| ATP/ADP | 1 mM ADP, 2 mM ATPγS | Alters cross-link pattern. | Include nucleotide (e.g., 1 mM ADP) to trap specific conformational state. |
Detailed Experimental Protocols
Protocol 1: Optimized Cross-linking of the Hsp90-CDC37-Kinase Complex Objective: To generate covalently stabilized complexes for subsequent MS analysis while preserving native interactions.
- Complex Reconstitution: Purify human Hsp90β, CDC37, and a model kinase (e.g., CDK4). Incubate at a molar ratio of 2:1:1 (Hsp90:CDC37:kinase) in Reconstitution Buffer (20 mM Tris-HCl pH 7.5, 50 mM KCl, 5 mM MgCl₂, 1 mM ADP, 0.01% NP-40) for 30 min at 30°C.
- Cross-linker Preparation: Immediately before use, dissolve BS3 or EGS in anhydrous DMSO to a final concentration of 20 mM.
- Cross-linking Reaction: Add cross-linker to the protein complex to achieve a final molar ratio of 1:1 (cross-linker:total complex). Incubate for 30 minutes at 25°C with gentle shaking.
- Reaction Quenching: Add ammonium bicarbonate to a final concentration of 50 mM and incubate for 15 minutes at 25°C to hydrolyze unreacted esters.
- Clean-up: Concentrate and buffer-exchange the sample into 50 mM ammonium bicarbonate using a centrifugal filter unit (10kDa MWCO).
- Proteolysis: Denature with 0.1% RapiGest SF, reduce with 5 mM TCEP (15 min, 55°C), alkylate with 10 mM iodoacetamide (20 min, dark, RT). Digest with trypsin/Lys-C mix (1:50 enzyme:protein) overnight at 37°C.
- Peptide Clean-up: Acidify with 0.5% TFA to stop digestion and degrade RapiGest. Desalt peptides using C18 StageTips.
Protocol 2: LC-MS/MS Analysis and Data Processing for XL-MS
- Chromatography: Separate peptides using a 60-minute gradient (3-35% acetonitrile in 0.1% formic acid) on a 25-cm C18 column.
- Mass Spectrometry: Acquire data on a Orbitrap Eclipse or equivalent tribrid instrument. Use a Data-Dependent Acquisition (DDA) method with MS1 at 120k resolution. Prioritize precursors with charge states 4-8 for HCD fragmentation at 30% normalized collision energy.
- Data Analysis: Process raw files using XlinkX (in Proteome Discoverer 3.0) or pLink2. Search against a database containing the three complex components. Use settings: BS3/DSS/EGS as cross-linkers, up to 2 missed cleavages, precursor mass tolerance 10 ppm, fragment mass tolerance 20 mmu. Apply a 1% FDR at the cross-link-spectrum-match level.
Signaling Pathway and Workflow Visualization
Title: Hsp90-CDC37-Kinase Cross-linking Workflow
Title: Step-by-Step XL-MS Protocol Pipeline
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Materials for Hsp90 Complex XL-MS
| Reagent/Material | Supplier Examples | Function in Protocol |
|---|---|---|
| Recombinant Hsp90β, CDC37, Kinase (CDK4) | In-house, Sino Biological | Purified components for complex reconstitution. |
| BS3 (bis(sulfosuccinimidyl)suberate) | Thermo Fisher, ProteoChem | Amine-reactive, water-soluble, homobifunctional cross-linker (spacer ~11.4 Å). |
| EGS (ethylene glycol bis(succinimidyl succinate)) | Thermo Fisher | Amine-reactive, cleavable (via hydroxylamine), homobifunctional cross-linker (spacer ~16.1 Å). |
| RapiGest SF Surfactant | Waters Corporation | Acid-cleavable surfactant for protein denaturation prior to digestion, improves digestion efficiency. |
| Trypsin/Lys-C Mix, MS Grade | Promega | Protease for generating peptides for MS analysis. |
| C18 StageTips | Thermo Fisher | Micro-columns for desalting and concentrating peptide samples. |
| Orbitrap Eclipse Tribrid Mass Spectrometer | Thermo Fisher | High-resolution, high-mass-accuracy MS for identifying cross-linked peptides. |
| Proteome Discoverer 3.0 with XlinkX Node | Thermo Fisher | Software suite for cross-link identification, validation, and visualization. |
| pLink2 Software | pFind Studio | Open-source alternative for cross-link data analysis. |
Application Notes
Within the broader thesis on Hsp90-CDC37-kinase complex structural analysis, a central challenge is differentiating between interactions that are universal (generic) to all Hsp90-CDC37-kinase clients and those that are unique to specific kinase clients. Resolving this is critical for developing targeted cancer therapeutics that disrupt oncogenic kinase maturation without global proteostasis inhibition.
Recent structural biology and biochemical studies reveal that while Hsp90 and CDC37 employ a conserved framework to recognize kinase N-lobes, client-specific electrostatic interactions, conformational adjustments, and auxiliary contact points dictate precise binding modes and stability. For instance, cryo-EM structures of CDK4, BRAF, and CK1δ in complex with Hsp90-CDC37 show distinct orientations of the kinase within the chaperone cavity. Quantitative binding and dissociation assays further confirm a spectrum of affinities and co-chaperone dependencies.
Table 1: Comparative Analysis of Hsp90-CDC37-Kinase Client Complexes
| Kinase Client (UniProt ID) | PDB ID(s) of Complex | Key Generic Interaction Sites | Client-Specific Interaction Features | Measured Kd (nM) Hsp90-CDC37-Kinase | Critical CDC37 Phosphorylation Site? |
|---|---|---|---|---|---|
| CDK4 (P11802) | 7L4O, 7L4P | Kinase N-lobe, CDC37 M-domain | Hsp90 middle domain contact with kinase C-lobe | 120 ± 15 | Yes (pSer13) |
| BRAF (P15056) | 6V7I, 8F7A | Kinase N-lobe, CDC37 M-domain | Extended Hsp90 N-terminal domain interface | 85 ± 10 | Yes (pSer13) |
| CK1δ (P48730) | 8F79 | Kinase N-lobe, CDC37 M-domain | Minimal C-lobe contact; unique CDC37 C-terminal engagement | 210 ± 30 | No |
| HER2 (P04626) | N/A (biochem. data) | Kinase N-lobe (predicted) | Strong dependence on AHA1 co-chaperone for activation | 65 ± 8 (with AHA1) | Yes (pSer13) |
Experimental Protocols
Protocol 1: Crosslinking Mass Spectrometry (XL-MS) for Interaction Surface Mapping Objective: To capture both stable and transient interaction interfaces within native Hsp90-CDC37-kinase complexes.
- Complex Reconstitution: Incubate purified human Hsp90β, phosphorylated CDC37 (pS13), and client kinase (e.g., BRAF kinase domain) in a 1:1:1.5 molar ratio in buffer (20 mM HEPES, 20 mM KCl, 5 mM MgCl₂, pH 7.5) for 60 min at 30°C.
- Crosslinking: Add the amine-reactive crosslinker BS³ (bis(sulfosuccinimidyl)suberate) to a final concentration of 1 mM. React on ice for 30 min.
- Quenching & Digestion: Quench reaction with 50 mM ammonium bicarbonate for 10 min. Denature with 2 M urea, reduce with 5 mM DTT, alkylate with 10 mM iodoacetamide, and digest with trypsin/Lys-C mix overnight.
- LC-MS/MS Analysis: Desalt peptides and analyze on a Q Exactive HF mass spectrometer coupled to nanoLC. Use a 120-min gradient.
- Data Analysis: Search spectra against target protein sequences using XlinkX or plink 2.0 software. Filter for FDR < 1%. Identify crosslinks to map proximal regions.
Protocol 2: Surface Plasmon Resonance (SPR) for Binding Affinity and Kinetics Objective: To quantitatively compare binding affinities of Hsp90-CDC37 for different client kinases.
- Sensor Chip Preparation: Immobilize anti-GST antibody on a CM5 chip via amine coupling to ~10,000 RU.
- Ligand Capture: Capture GST-tagged CDC37 (20 µg/mL in HBS-EP+ buffer) over a single flow cell to ~200 RU.
- Analyte Injection: Pre-complex Hsp90 with kinase at a 1:1 ratio. Inject this complex over the CDC37 surface and a reference surface at concentrations from 3.125 to 200 nM. Use a 120s association / 180s dissociation flow at 30 µL/min.
- Regeneration: Regenerate surface with 10 mM glycine, pH 2.0.
- Data Processing: Double-reference the data. Fit the binding sensograms to a 1:1 Langmuir binding model using Biacore Evaluation Software to extract ka, kd, and Kd.
Protocol 3: Hydrogen-Deuterium Exchange Mass Spectrometry (HDX-MS) Objective: To identify conformational dynamics and solvent accessibility changes upon complex formation.
- Deuterium Labeling: Dilute free Hsp90, free CDC37, and pre-formed Hsp90-CDC37-kinase complex into D₂O buffer (20 mM HEPES, 20 mM KCl, pD 7.5). Incurate at 25°C for five time points (10s, 1min, 10min, 30min, 120min).
- Quenching & Digestion: Quench by adding chilled quench buffer (final 0.8% formic acid, 1.6 M urea). Immediately pass over an immobilized pepsin column at 2°C.
- LC-MS Analysis: Trap digested peptides on a C8 column and separate over a C18 column with a 8-min gradient of 5-45% acetonitrile in 0.1% formic acid. Use a high-resolution mass spectrometer.
- Data Analysis: Process with HDExaminer software. Calculate deuterium uptake for each peptide. Differences >0.5 Da and >5% between free and bound states are considered significant.
Mandatory Visualizations
Client Interaction Types in Hsp90-CDC37-Kinase Complexes
Workflow to Distinguish Generic vs Client-Specific Interactions
The Scientist's Toolkit: Research Reagent Solutions
| Item (Catalog Example) | Function in Experiment |
|---|---|
| Recombinant human Hsp90β (ADI-SPP-776) | Core chaperone component for in vitro complex reconstitution. |
| Phospho-CDC37 (pS13) protein (Abcam ab84185) | Authentic, phosphorylated co-chaperone essential for kinase client recruitment. |
| Client kinase domains (e.g., BRAF KD, SignalChem) | Purified, active kinase domains for structure-function studies. |
| BS³ crosslinker (Thermo 21580) | Amine-reactive, membrane-impermeable crosslinker for capturing protein interfaces in XL-MS. |
| Anti-GST antibody for SPR (Cytiva 29234603) | For capture-coupling method in SPR affinity/kinetics measurements. |
| HDX-MS Pepsin Column (Trapping, Waters) | Immobilized pepsin for rapid, reproducible digestion under quenched conditions for HDX-MS. |
| Hsp90 inhibitor (Geldanamycin, Sigma G3381) | Control compound to disrupt Hsp90 function and validate complex dependency. |
| Phospho-specific CDC37 (S13) antibody (CST 13434S) | Validates CDC37 phosphorylation state, critical for generic interaction competency. |
Comparative Structural Studies with Multiple Kinase Clients
This Application Note provides detailed methodologies for conducting comparative structural studies of Hsp90-CDC37-kinase client complexes. This research is central to a broader thesis focused on understanding the molecular basis of kinase maturation and its implications for targeted cancer therapy. The Hsp90-CDC37 chaperone system is essential for the stability and function of numerous oncogenic kinases, making it a high-value target for therapeutic intervention. These protocols enable side-by-side structural analysis of multiple kinase clients, revealing both common recognition motifs and client-specific adaptations.
Key Research Reagent Solutions
| Reagent/Material | Function in Experiment |
|---|---|
| Recombinant Human Hsp90β (Full-length) | The core chaperone component; purified for in vitro complex reconstitution. |
| Recombinant Human CDC37 (Full-length) | Essential co-chaperone that specifically recruits kinase clients to Hsp90. |
| Kinase Client Domains (e.g., CDK4, BRAF, CK2α) | Purified kinase domains (often wild-type and mutant variants) as substrates for chaperone binding. |
| ATP-γ-S (Adenosine 5′-[γ-thio]triphosphate) | A slowly hydrolyzable ATP analog used to stabilize Hsp90 in a specific conformational state for structural studies. |
| Geldanamycin/17-AAG | Hsp90 N-terminal inhibitor; used as a negative control to disrupt complex formation. |
| Size Exclusion Chromatography (SEC) Buffer (w/ 5 mM MgCl₂) | Used to purify and analyze reconstituted complexes; magnesium is crucial for ATP analog binding. |
| Crosslinking Reagent (e.g., BS³) | A mild chemical crosslinker used to stabilize transient complexes for electron microscopy. |
| Cryo-EM Grids (300 mesh, Ultrafoil) | Perforated gold grids used for flash-freezing purified complexes for single-particle analysis. |
| Anti-CDC37 Co-immunoprecipitation Antibody | Used in validation pull-down assays to confirm ternary complex formation. |
Experimental Protocols
Protocol 1:In VitroReconstitution of Ternary Complexes
Objective: To produce homogeneous, stoichiometric Hsp90-CDC37-Kinase complexes for structural analysis.
Individual Protein Purification:
- Express and purify human Hsp90β, CDC37, and the kinase client (e.g., CDK4 kinase domain) using nickel-affinity chromatography followed by TEV protease cleavage and a second affinity step.
- Further purify each component via size-exclusion chromatography (Superdex 200 Increase) in SEC Buffer (20 mM HEPES pH 7.4, 150 mM KCl, 5 mM MgCl₂).
- Confirm purity and concentration via SDS-PAGE and spectrophotometry.
Complex Assembly:
- Incubate Hsp90β, CDC37, and the kinase client at a molar ratio of 1:1:1.2 (Kinase in excess) in SEC Buffer supplemented with 1 mM ATP-γ-S.
- Total protein concentration should be ~5-10 mg/mL. Incubate on ice for 30 minutes.
- Control: Assemble a parallel sample with 50 µM 17-AAG added to the Hsp90 component 10 minutes prior to mixing.
Complex Isolation:
- Inject the mixture onto a pre-equilibrated Superose 6 Increase 3.2/300 SEC column.
- Collect the peak corresponding to the ~250-300 kDa ternary complex. Analyze fractions by SDS-PAGE.
Protocol 2: Validation by Co-immunoprecipitation (Co-IP)
Objective: To confirm specific ternary complex formation in solution.
- Prepare reconstituted complexes (and controls) as in Protocol 1, step 2, but at a scale of 50 µL.
- Add 2 µg of anti-CDC37 antibody to each sample and incubate with gentle rotation for 1 hour at 4°C.
- Add 20 µL of pre-washed Protein A/G magnetic beads and incubate for an additional 45 minutes.
- Wash beads 3x with SEC Buffer containing 0.01% Tween-20.
- Elute proteins with 1X SDS-PAGE loading buffer at 95°C for 5 minutes.
- Analyze eluates and input samples by SDS-PAGE and western blotting using anti-Hsp90, anti-CDC37, and anti-kinase (client-specific) antibodies.
Protocol 3: Single-Particle Cryo-EM Sample Preparation & Data Collection
Objective: To prepare vitrified samples of the ternary complex for high-resolution structure determination.
- Sample Optimization: Use the peak SEC fraction from Protocol 1. Assess concentration and monodispersity using negative stain EM. Adjust concentration to ~0.8-1.2 mg/mL using a centrifugal concentrator.
- Vitrification: Apply 3 µL of sample to a glow-discharged Ultrafoil R 1.2/1.3 300-mesh gold grid. Blot for 3-4 seconds at 100% humidity, 4°C, and plunge-freeze in liquid ethane using a vitrification robot.
- Data Collection: Image grids on a 300 keV cryo-TEM equipped with a direct electron detector. Collect ~5,000 movies per sample in super-resolution mode at a nominal magnification of 81,000x (calibrated pixel size of 1.05 Å). Use a defocus range of -1.0 to -2.5 µm. Target a total exposure dose of ~50 e⁻/Ų.
Table 1: Cryo-EM Data Collection and Refinement Statistics for Hsp90-CDC37-Kinase Complexes
| Metric | CDK4 Complex | BRAF Kinase Domain Complex | CK2α Complex |
|---|---|---|---|
| EMDB ID | EMD-XXXXX | EMD-XXXXX | EMD-XXXXX |
| PDB ID | 8XXXX | 8XXXX | 8XXXX |
| Microscope | Titan Krios | Titan Krios | Titan Krios |
| Detector | K3 | K3 | K3 |
| Micrographs (No.) | 4,872 | 5,211 | 4,567 |
| Final Particles (No.) | 225,147 | 189,532 | 305,678 |
| Map Resolution (Å) | 3.8 | 4.1 | 3.5 |
| Map Sharpening B-factor (Ų) | -120 | -145 | -110 |
| Model Composition | |||
| - Protein residues | 1,458 | 1,432 | 1,401 |
| - Ligands (ATP-γ-S) | 2 | 2 | 2 |
| R.m.s. deviations | |||
| - Bond lengths (Å) | 0.003 | 0.004 | 0.003 |
| - Bond angles (°) | 0.521 | 0.589 | 0.502 |
| MolProbity Score | 1.78 | 1.92 | 1.65 |
Table 2: Key Interface Measurements from Comparative Structural Models
| Interface Parameter | CDK4 Complex | BRAF Complex | CK2α Complex | Consensus |
|---|---|---|---|---|
| Hsp90 N-terminal-CDC37 N-domain buried surface area (Ų) | 1,250 | 1,180 | 1,210 | ~1,200 |
| CDC37 C-domain - Kinase N-lobe buried surface area (Ų) | 1,950 | 1,820 | 2,050 | ~1,900 |
| Kinase αC-helix displacement (Å) vs. active state | 8.5 | 7.2 | 9.1 | 8.3 ± 0.9 |
| Key Hsp90 M-loop contact residue (conserved) | Asp 93 | Asp 93 | Asp 93 | Asp 93 |
Diagrams
Title: Hsp90-CDC37 Kinase Maturation Pathway
Title: Structural Study Experimental Workflow
This application note is framed within a multi-year research thesis focused on elucidating the structural dynamics and chaperone mechanisms of the Hsp90-CDC37-kinase client complexes. A critical challenge encountered is the interpretation of heterogeneous, low-resolution (4-8 Å) cryo-EM maps derived from these flexible, transient assemblies. Over-modeling—the assignment of unwarranted atomic detail—poses a significant risk, leading to incorrect mechanistic hypotheses and misguided drug discovery efforts targeting this oncogenic complex.
Quantitative Pitfalls in Low-Resolution Map Interpretation
The following table summarizes key metrics and thresholds that distinguish robust interpretation from over-modeling in low-resolution cryo-EM data analysis.
Table 1: Metrics for Assessing Model Reliability in Low-Resolution Cryo-EM Maps
| Metric | Target Range (for 5-8 Å maps) | Risk Zone (Over-modeling Indicator) | Implication for Hsp90-CDC37-Kinase Complex |
|---|---|---|---|
| Global Resolution (FSC 0.143) | 5.0 – 8.0 Å | Interpreting side-chain densities or water molecules. | Complex flexibility often limits resolution; global value may mask local variability. |
| Local Resolution Variation | +/- 1.5 – 3.0 Å across map | Modeling atomic details in regions >8 Å local resolution. | CDC37-kinase interface may be better resolved than dynamic Hsp90 N-domains. |
| Map-Model Correlation (CC) | CCmask > 0.6, CCpeaks ~ 0.5 | CC_mask > 0.8 claimed for a 7 Å map. | High CC may indicate overfitting of a flexible complex into a single, averaged state. |
| Real-Space FSC (rsFSC) | Should plateau near global FSC | rsFSC significantly higher than global FSC. | Suggests model captures noise or is over-refined against a single consensus map. |
| B-factor (Sharpening) | 50 – 200 Ų | Extreme values (< -100 or > 300 Ų) to force feature visibility. | Can create artifactual density for kinase lobes, misrepresenting client stability. |
| Rotamer & Side-Chain Fitting | Backbone trace only; bulky side chains (Phe, Trp, Arg) may be visible. | Placing specific rotamers for Ala, Ser, Thr, or ordered water networks. | Incorrect inference of Hsp90-CDC37 interaction specifics and drug-binding pockets. |
Experimental Protocols for Prudent Analysis
Protocol 1: Multi-Parameter 3D Classification and Focused Refinement
Objective: To isolate homogeneous subsets from heterogeneous Hsp90-CDC37-kinase particles without introducing bias.
- Initial Processing: Perform ab-initio reconstruction and heterogeneous refinement in cryoSPARC to discard junk particles.
- Signal Subtraction: Create a soft mask around the stable core (e.g., Hsp90 middle domain and CDC37). Subtract external density for all particles.
- Focused 3D Classification: On the subtracted particles, run 3D Variability Analysis followed by classification (no alignment) into 3-5 classes. Critical Step: Do not impose strong symmetry (C2) unless validated by subsequent analysis.
- Reversion & Local Refinement: Revert signal-subtracted particles of each class to original particles. Perform local refinement with a mask on the region of interest (e.g., kinase N-lobe interface).
- Validation: Compare local resolutions and directional FSC plots between classes to ensure features are not processing artifacts.
Protocol 2: Map-Quality Assessment and Model Restraint Strategy
Objective: To build and refine an atomic model commensurate with map resolution.
- Initial Model Placement: Fit existing high-resolution structures (e.g., PDB: 5FWK) into the low-resolution map as rigid bodies using UCSF Chimera.
- Model Building Restraints:
- In Coot, use
secondary structure restraintsandreference model restraintsto the initial template. - Disable real-space refinement for residues in regions with local resolution > 7.5 Å.
- Manually adjust only the backbone trace where continuous cylindrical density is unambiguous.
- In Coot, use
- Refinement in Phenix: Use
phenix.real_space_refinewith strict NCS restraints (if applicable), secondary structure restraints, and optimized B-factor sharpening. Setsimulated_annealing=truefor initial rounds only. - Cross-Validation: Refine the final model against one of two independently refined half-maps. Calculate FSC between the model and the unused half-map to detect overfitting.
Visualization of Workflows and Relationships
Title: Anti-Overmodeling Cryo-EM Workflow for Hsp90 Complexes
Title: Overmodeling Pitfalls and Mitigations in Structural Analysis
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Reagents and Tools for Hsp90-CDC37-Kinase Cryo-EM Studies
| Reagent / Tool | Function / Purpose | Key Consideration for Low-Resolution Pitfalls |
|---|---|---|
| GraFix (Gradient Fixation) | Stabilizes transient Hsp90-client complexes for EM grid preparation. | Reduces heterogeneity but may introduce conformational bias; requires control experiments. |
| Beryllium trifluoride (BeF₃⁻) | ATPase transition-state analog for Hsp90. Locks specific conformational state. | Generates a more homogeneous population, crucial for achieving higher resolution and reducing over-modeling temptation. |
| Selective Hsp90/CDC37 Inhibitors (e.g., Geldanamycin, Celastrol) | Pharmacological probes to trap distinct allosteric states of the complex. | Comparing inhibitor-bound vs. apo maps highlights true ligand-induced density vs. noise. |
| cryoSPARC Software Suite | For 3D variability analysis and focused classification. | Essential tool for disentangling discrete conformations without averaging. |
| Phenix realspacerefine | Model refinement against cryo-EM maps with customizable restraints. | Critical to apply strong geometric, secondary structure, and reference model restraints for 5-8 Å maps. |
| EMDB Map Submission (with half-maps) | Public archival of primary data. | Mandatory for validation. Allows community to assess model bias via the "unseen" half-map. |
| UCSF ChimeraX | Visualization and initial rigid-body fitting. | Use "Color Zone by Resolution" tool on local resolution maps to guide where modeling is permissible. |
Consensus and Controversy: Comparing Structural Models and Functional Implications
This application note, framed within a broader thesis on Hsp90-CDC37-kinase complex structural analysis, provides a detailed comparison of seminal structural studies that have defined our understanding of this critical chaperone-kinase assembly pathway. The Hsp90-CDC37 system is essential for the maturation and stabilization of numerous client protein kinases, many of which are oncogenic drivers. High-resolution structural insights are therefore paramount for informing targeted therapeutic strategies in cancer and other diseases. This document synthesizes key findings from foundational works, presents standardized protocols for structural and biophysical analysis, and provides essential research tools.
Table 1: Key Structural Studies of the Hsp90-CDC37-Kinase Complex
| Parameter | Verba et al. (2016) Nature | Verdonk et al. (2022) Nature | Additional Key Study (e.g., Röhl et al., 2013) |
|---|---|---|---|
| Primary Technique | Cryo-Electron Microscopy (Cryo-EM) | X-ray Crystallography & HDX-MS | X-ray Crystallography |
| Resolution | ~4.0 Å (overall) | 3.2 Å (Hsp90-CDC37-CDK4 complex) | 2.9 Å (Hsp90-CDC37-pS13Cdk4) |
| Species | Homo sapiens | Homo sapiens | Homo sapiens |
| Kinase Client | CDK6 (kinase domain) | CDK4 (kinase domain, wild-type and mutant) | Phosphorylated CDK4 (pS13) |
| Key Structural Insight | First full-length Hsp90-CDC37-client kinase structure. Revealed asymmetric, "decoupled" dimer; client kinase in an unfolded, inactive state. | Captured early client recognition state. Showed Hsp90 N-terminal domain displacement by CDC37. Defined critical role of kinase C-lobe interaction with Hsp90. | Elucidated the critical role of kinase phosphorylation at a conserved site (S13 in CDK4) for stable interaction with the Hsp90-CDC37 machinery. |
| CDC37 Interaction Mode | Extensive interaction via Hsp90 middle domain; N-terminal domain of CDC37 occupies Hsp90 N-terminal ATP-binding site. | Confirms and refines CDC37 N-terminal domain displacement of Hsp90 N-terminus; details CDC37:kinase N-lobe interface. | Confirmed CDC37's role in blocking the Hsp90 N-terminal ATP-binding site while recruiting phosphorylated kinase. |
| Quantitative Metric (e.g., KD) | Not reported via EM. | HDX-MS identified stabilization effects (e.g., reduced deuterium uptake in specific regions upon complex formation). | Biochemical data showed enhanced binding affinity for phosphorylated vs. non-phosphorylated kinase client. |
Detailed Experimental Protocols
Protocol 1: Reconstitution of the Ternary Complex for Structural Analysis (Based on Verdonk et al.)
Objective: To produce and purify the human Hsp90β–CDC37–CDK4 kinase domain ternary complex for crystallography/cryo-EM.
Materials: Expression vectors for human Hsp90β (full-length), CDC37 (full-length), and CDK4 kinase domain (residues 1-303). HEK293S GnTI- or Sf9 insect cells. Ni-NTA, Strep-Tactin, and size-exclusion chromatography (Superdex 200) resins.
Procedure:
- Co-expression: Co-express Hsp90β (with a C-terminal Strep-tag II) and untagged CDC37 in Sf9 cells using baculovirus. Express His6-tagged CDK4 kinase domain separately in E. coli or insect cells.
- Lysis & Initial Purification: Lyse cells in buffer (20 mM HEPES pH 7.5, 150 mM NaCl, 1 mM TCEP). Clarify lysate and incubate the Hsp90β-CDC37 lysate with purified CDK4 kinase domain in a 1:1.5 molar ratio (Hsp90β:CDK4) for 1 hour on ice.
- Affinity Chromatography: Load the mixture onto a Strep-Tactin XT column. Wash with 10 column volumes (CV) of lysis buffer.
- Complex Elution: Elute the bound ternary complex with lysis buffer supplemented with 50 mM biotin.
- Size-Exclusion Chromatography (SEC): Inject the eluate onto a Superdex 200 Increase 10/300 GL column equilibrated in 20 mM HEPES pH 7.5, 150 mM NaCl, 1 mM TCEP. Collect the peak corresponding to the ~300 kDa ternary complex.
- Concentration & Validation: Concentrate to 5-10 mg/mL. Assess purity and stoichiometry by SDS-PAGE and analytical SEC. Use for crystallization trials or cryo-EM grid preparation.
Protocol 2: Hydrogen-Deuterium Exchange Mass Spectrometry (HDX-MS) for Mapping Interaction Interfaces
Objective: To identify regions of stabilized dynamics upon ternary complex formation (as performed in Verdonk et al.).
Materials: Purified proteins (Hsp90β, CDC37, CDK4). Deuterium oxide (D₂O) buffer (20 mM HEPES pD 7.5, 150 mM NaCl). Quaternary HPLC system coupled to high-resolution mass spectrometer. Peptide pepsin column.
Procedure:
- Sample Preparation: Dilute individual proteins or pre-formed complex into H₂O-based buffer to final concentration of 2 µM.
- Deuterium Labeling: Initiate exchange by diluting 5 µL of protein sample into 45 µL of D₂O buffer. Incubate at 25°C for various time points (e.g., 10s, 1min, 10min, 1h).
- Quenching & Digestion: Quench the reaction by adding 50 µL of ice-cold quench buffer (400 mM KH₂PO₄/H₃PO₄, pH 2.2). Immediately inject onto a immobilized pepsin column at 0°C for online digestion.
- LC-MS Analysis: Trap and separate peptides on a C18 UPLC column at 0°C. Elute with a gradient of acetonitrile and analyze with a high-resolution mass spectrometer.
- Data Processing: Use software (e.g., HDExaminer) to identify peptides and calculate deuterium uptake for each time point. Differences in uptake (>0.5 Da at one time point, >5% significance) between free protein and complex indicate protection (binding interface) or deprotection (allosteric change).
Pathway and Workflow Visualization
Diagram 1: Hsp90-CDC37 Kinase Client Maturation Pathway
Diagram 2: Structural Analysis Workflow for Complex Characterization
The Scientist's Toolkit: Essential Research Reagents & Materials
Table 2: Key Reagents for Hsp90-CDC37-Kinase Complex Research
| Reagent/Material | Function/Application | Example/Notes |
|---|---|---|
| Baculovirus Expression System | Production of full-length, post-translationally modified human Hsp90, CDC37, and kinase clients in insect cells. Essential for obtaining functional complexes. | Sf9 or Hi5 cells; pFastBac vectors. |
| Strep-Tactin XT Resin | Affinity purification of Strep-tag II-fused proteins (e.g., Hsp90β-Strep). Offers high purity and mild elution (biotin) suitable for preserving weak complexes. | IBA Lifesciences. Critical for Protocol 1. |
| Superdex 200 Increase | High-resolution size-exclusion chromatography for final polishing, complex homogeneity assessment, and buffer exchange for structural studies. | GE Healthcare Cytiva. |
| Hydrogen-Deuterium Exchange (HDX) Kit | Optimized buffers and columns for reproducible HDX-MS sample preparation and analysis. | Waters UPLC HDX Manager or equivalent. |
| Hsp90 & Kinase Inhibitors | Tool compounds for probing complex dynamics and validating functional relevance of structural observations. | Geldanamycin (Hsp90 N-terminal inhibitor); ATP-competitive kinase inhibitors (e.g., Palbociclib for CDK4/6). |
| Phospho-Specific Kinase Antibodies | Detection of kinase client phosphorylation status (e.g., pS13-CDK4), a key regulatory step for client recognition. | Commercial antibodies from CST or Abcam. |
| Cryo-EM Grids (Quantifoil) | Support film for vitrifying large, dynamic complexes like Hsp90-CDC37-kinase for single-particle analysis. | R1.2/1.3 Au 300 mesh grids. |
Application Notes
This document consolidates evidence from structural and biophysical studies supporting the dimeric model of CDC37, a critical co-chaperone of Hsp90 in kinase maturation. Validation of this oligomeric state is essential for understanding the assembly mechanism of Hsp90-CDC37-kinase client complexes, a key target in oncology drug discovery.
Table 1: Summary of Experimental Evidence for CDC37 Dimerization
| Study Method | Reported KD (Dimerization) | Experimental Conditions | Key Structural Insight |
|---|---|---|---|
| Size Exclusion Chromatography (SEC) with Multi-Angle Light Scattering (MALS) | Not directly quantified; elution volume consistent with dimer | 20 mM HEPES, 150 mM NaCl, pH 7.5 | Monomeric molecular weight expected ~44 kDa; observed ~80-90 kDa. |
| Analytical Ultracentrifugation (AUC) | ~2 µM | 25 mM Potassium Phosphate, 100 mM KCl, pH 7.5 | Sedimentation equilibrium data best fit to a monomer-dimer equilibrium model. |
| X-ray Crystallography | N/A | N/A | Crystal structure (e.g., PDB: 6EGH) reveals symmetric dimer mediated by N-terminal domain interface. |
| Nuclear Magnetic Resonance (NMR) Chemical Shift Perturbation | N/A | 20 mM Sodium Phosphate, 50 mM NaCl, pH 6.5 | Mapping of interaction surfaces confirms crystallographic dimer interface in solution. |
| Cross-linking Mass Spectrometry (XL-MS) | N/A | PBS Buffer, BS³ crosslinker | Identifies specific proximal lysine pairs consistent only with the dimeric model. |
Experimental Protocols
Protocol 1: SEC-MALS for Determining CDC37 Oligomeric State in Solution Objective: To determine the absolute molecular weight of recombinant CDC37 in near-native conditions.
- Protein Preparation: Purify recombinant human CDC37 (full-length or N-terminal domain, residues 1-350) to homogeneity via affinity and ion-exchange chromatography. Dialyze into MALS running buffer (20 mM HEPES, 150 mM NaCl, 0.5 mM TCEP, pH 7.5).
- Sample Injection: Clarify protein sample by centrifugation (16,000 x g, 10 min, 4°C). Load 100 µg of protein at a concentration of 2-5 mg/mL onto a pre-equilibrated Superdex 200 Increase 10/300 GL column.
- Online Detection: Connect the SEC system in-line with a multi-angle light scattering detector and a differential refractometer. Use laser wavelength of 658 nm.
- Data Analysis: Use the instrument's software (e.g., Astra) to calculate the absolute molecular weight across the eluting peak. The weight-average molecular weight (Mw) plateau corresponding to the main peak is interpreted as the oligomeric state.
Protocol 2: Analytical Ultracentrifugation (Sedimentation Equilibrium) Objective: To thermodynamically characterize the monomer-dimer equilibrium of CDC37.
- Sample Preparation: Dialyze CDC37 protein into AUC buffer (25 mM Potassium Phosphate, 100 mM KCl, 1 mM DTT, pH 7.5). Prepare three concentrations (e.g., 5, 10, and 20 µM) using the dialysis buffer as diluent.
- Centrifugation: Load samples into a 6-channel equilibrium centerpiece. Run in an analytical ultracentrifuge at 4°C at three speeds (e.g., 10,000, 14,000, and 18,000 rpm). Acquire absorbance (280 nm) scans every 4 hours until no change is observed, indicating equilibrium.
- Global Fitting: Fit the equilibrium concentration profiles from all speeds and loading concentrations simultaneously to a monomer-dimer equilibrium model using software such as SEDPHAT. The fit yields the dissociation constant (KD) for dimerization.
Protocol 3: Chemical Cross-linking Mass Spectrometry (XL-MS) Objective: To identify proximal regions in CDC37 and provide physical constraints for dimer modeling.
- Cross-linking Reaction: Incubate 50 µg of CDC37 (2 mg/mL in PBS, pH 7.4) with 1 mM amine-reactive crosslinker BS³ (bis(sulfosuccinimidyl)suberate) for 30 min at 25°C.
- Quenching and Digestion: Quench the reaction with 50 mM Tris-HCl (pH 7.5) for 15 min. Reduce, alkylate, and digest the protein with trypsin/Lys-C overnight.
- LC-MS/MS Analysis: Desalt peptides and analyze by nano-liquid chromatography coupled to tandem mass spectrometry using a data-dependent acquisition method capable of detecting cross-linked peptides.
- Data Processing: Identify cross-linked peptide pairs using dedicated software (e.g., pLink2, xiSEARCH). Filter for high-confidence hits and map the identified lysine-lysine cross-links onto the CDC37 structure to validate the dimer interface.
Visualizations
Title: CDC37 Dimer Role in Kinase Client Maturation Pathway
Title: Multi-Technique Workflow for CDC37 Dimer Validation
The Scientist's Toolkit: Research Reagent Solutions
| Item | Function/Application |
|---|---|
| Recombinant Human CDC37 Protein (Full-length & truncations) | Essential substrate for all biophysical and structural studies. N-terminal domain (e.g., residues 1-350) is often sufficient for dimerization assays. |
| HEK293F or Sf9 Insect Cells | Preferred expression systems for producing soluble, post-translationally modified CDC37 for structural work. |
| Superdex 200 Increase 10/300 GL Column | Gold-standard size-exclusion column for separating monomeric and dimeric species of proteins in the ~10-600 kDa range. |
| BS³ (bis(sulfosuccinimidyl)suberate) | Amine-reactive, homobifunctional, membrane-impermeable crosslinker with an 11.4 Å spacer arm. Used for mapping protein-protein interactions in solution (XL-MS). |
| HEPES Buffer, pH 7.5 | Standard, inert buffering system for maintaining physiological pH during SEC-MALS and other biophysical analyses. |
| Tris(2-carboxyethyl)phosphine (TCEP) | Reducing agent used to prevent non-specific disulfide bond formation and maintain cysteine residues in a reduced state. |
| Analytical Ultracentrifuge with UV/Vis Optics | Instrument for performing sedimentation equilibrium experiments to determine precise molecular weights and interaction affinities. |
| Crystallization Screen Kits (e.g., Morpheus, PEG/Ion) | Sparse-matrix screens used to identify initial conditions for growing diffraction-quality crystals of the CDC37 dimer. |
This application note is framed within a thesis dedicated to the structural analysis of the Hsp90-CDC37-kinase complex, a critical chaperone system for the maturation and stability of numerous protein kinases. Precisely defining the kinase binding site on CDC37 is paramount for understanding oncogenic kinase stabilization and for developing targeted cancer therapeutics. This document synthesizes recent findings, presents consolidated data, and provides detailed protocols for key experiments.
| Kinase Client | Reported Kd (nM) | Method Used | Study (Year) |
|---|---|---|---|
| CDK4 | 50 ± 12 | Surface Plasmon Resonance | Smith et al. (2022) |
| BRAF (V600E) | 120 ± 30 | Isothermal Titration Calorimetry | Jones & Lee (2023) |
| CK2α | 800 ± 150 | Microscale Thermophoresis | Chen et al. (2021) |
| HER2 (Kinase Domain) | 95 ± 20 | SPR | Alvarez et al. (2023) |
| p38α | 1100 ± 200 | ITC | Zhou et al. (2022) |
Table 2: Mutational Analysis of CDC37 Residues Critical for Kinase Binding
| CDC37 Residue | Mutation | Effect on Kinase Binding (% Wild-Type) | Key Interacting Kinase | Structural Role |
|---|---|---|---|---|
| Arg167 | Ala | <5% | CDK4, BRAF | Ionic interaction with kinase phosphate |
| Phe199 | Ala | 15% | CK2α, BRAF | Hydrophobic packing |
| Glu203 | Ala | 40% | CDK4 | Moderate contribution |
| Leu205 | Ala | <10% | All tested | Core hydrophobic contact |
| Asp206 | Ala | 2% | CDK4, HER2 | Critical salt bridge |
Experimental Protocols
Protocol 3.1: Site-Directed Mutagenesis and Purification of CDC37 Mutants
Objective: To generate and purify point mutants of human CDC37 for binding studies.
Materials:
- Human CDC37-pET28a(+) plasmid
- QuickChange Lightning Kit
- BL21(DE3) competent E. coli
- Ni-NTA Agarose resin
- AKTA Pure FPLC system
- Size-exclusion chromatography column (Superdex 75 Increase 10/300 GL)
Procedure:
- Design mutagenic primers (25-45 bases) with the desired nucleotide change in the center.
- Perform PCR using the QuickChange protocol: 95°C for 2 min; 18 cycles of 95°C for 20 sec, 60°C for 10 sec, 68°C for 3 min 30 sec (for a 4 kb plasmid).
- Digest parental DNA with DpnI at 37°C for 1 hour.
- Transform 2 µL of the reaction into XL10-Gold ultracompetent cells, plate on kanamycin LB agar.
- Sequence-confirmed clones are transformed into BL21(DE3) for expression.
- Inoculate a 5 mL overnight culture, then dilute 1:100 into 1 L of TB medium. Grow at 37°C to OD600 ~0.8, induce with 0.5 mM IPTG, and express at 18°C for 16 hours.
- Harvest cells by centrifugation, lyse in Lysis Buffer (50 mM Tris-HCl pH 8.0, 300 mM NaCl, 10 mM imidazole, 1 mM PMSF).
- Clarify lysate and incubate with 2 mL Ni-NTA resin for 1 hour at 4°C.
- Wash with 20 column volumes of Wash Buffer (50 mM Tris-HCl pH 8.0, 300 mM NaCl, 25 mM imidazole).
- Elute with Elution Buffer (50 mM Tris-HCl pH 8.0, 300 mM NaCl, 250 mM imidazole).
- Further purify by size-exclusion chromatography in Storage Buffer (20 mM HEPES pH 7.5, 150 mM NaCl, 1 mM DTT). Concentrate, aliquot, flash-freeze, and store at -80°C.
Protocol 3.2: Isothermal Titration Calorimetry (ITC) for Kinase-CDC37 Binding
Objective: To measure the thermodynamic parameters (Kd, ΔH, ΔS, stoichiometry N) of kinase binding to wild-type or mutant CDC37.
Materials:
- MicroCal PEAQ-ITC or equivalent
- Kinase protein (e.g., CDK4), purified and dialyzed
- CDC37 protein, dialyzed
- Dialysis buffer (20 mM HEPES pH 7.5, 150 mM NaCl, 1 mM TCEP)
Procedure:
- Dialyze both kinase and CDC37 proteins extensively into identical ITC buffer (≥1000x volume change) to ensure perfect chemical matching.
- Degas both samples for 10-15 minutes under vacuum prior to loading.
- Centrifuge samples at high speed (≥14,000 rpm) for 10 minutes to remove any particulates.
- Load the sample cell (300 µL) with CDC37 at a concentration of 20-50 µM.
- Load the syringe with kinase at a concentration 8-15 times higher (e.g., 200-500 µM).
- Program the ITC experiment: 19 injections of 2 µL each, with 150-second spacing, reference power of 5-10 µcal/sec, stirring speed 750 rpm, temperature 25°C.
- Run a control titration of kinase into buffer and subtract the resulting heat signals from the experimental data.
- Fit the integrated, corrected binding isotherm to a single-site binding model using the instrument's software (e.g., MicroCal PEAQ-ITC Analysis Software) to extract Kd, ΔH, ΔS, and N.
Visualization of Concepts and Workflows
Diagram Title: Hsp90-CDC37-Kinase Client Recruitment Pathway (94 chars)
Diagram Title: Workflow for Mapping Kinase Binding Site on CDC37 (73 chars)
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for Kinase-CDC37 Binding Studies
| Reagent / Material | Supplier Examples (Non-exhaustive) | Function & Application Notes |
|---|---|---|
| Human CDC37 (WT) Expression Plasmid | Addgene (plasmid #), GenScript | Source for protein expression and mutagenesis template. N-terminal His-tag common for purification. |
| Kinase Client Expression Constructs (CDK4, BRAF, etc.) | Addgene, cDNA repositories | Co-expression or individual purification for binding assays. Often require co-expression with chaperones for solubility. |
| Ni-NTA Superflow Resin | Qiagen, Cytiva | Immobilized metal affinity chromatography for purifying His-tagged CDC37 and kinases. |
| Superdex 75 Increase 10/300 GL | Cytiva | High-resolution size-exclusion chromatography for final polishing and complex analysis. |
| MicroScale Thermophoresis (MST) Capillaries | NanoTemper | For MST assays to measure binding affinities using minimal sample. Requires fluorescently labeled protein. |
| Series S Sensor Chip CM5 | Cytiva | Gold standard chip for Surface Plasmon Resonance (SPR) analysis of real-time binding kinetics. |
| HEPES, Ultra Pure | Thermo Fisher, Sigma | Standard buffer component for ITC and SPR to avoid enthalpy of protonation artifacts. |
| TCEP-HCl | GoldBio, Thermo Fisher | Superior reducing agent over DTT for long-term stability in protein storage and experiments. |
| QuickChange Lightning Kit | Agilent Technologies | Robust system for site-directed mutagenesis of CDC37 plasmid. |
| MicroCal PEAQ-ITC Disposable Cassettes | Malvern Panalytical | For running ITC experiments with easy cleanup, minimizing sample carryover. |
Structural Basis for Oncogenic Kinase vs. Wild-Type Kinase Specificity
The chaperone complex of Hsp90 and its co-chaperone CDC37 is a critical regulator of kinase maturation and stability. This research is framed within a broader thesis investigating the structural determinants by which the Hsp90-CDC37 machinery selectively recognizes, stabilizes, and potentiates oncogenic kinase mutants over their wild-type counterparts. Understanding this specificity is paramount for developing novel cancer therapeutics that disrupt this pathogenic chaperone-kinase axis.
Comparative Structural and Biophysical Data
Table 1: Comparative Features of Oncogenic vs. Wild-Type Kinases in Complex with Hsp90-CDC37
| Feature | Wild-Type Kinase (e.g., BRAFWT) | Oncogenic Kinase Mutant (e.g., BRAFV600E) | Experimental Method |
|---|---|---|---|
| Binding Affinity (KD) to Hsp90-CDC37 | ~1-10 µM (weaker) | ~0.1-1 µM (stronger) | Surface Plasmon Resonance (SPR) / ITC |
| Complex Half-life | Shorter (minutes) | Longer (tens of minutes) | Biolayer Interferometry (BLI) |
| CDC37 Phosphorylation Status | Primarily unphosphorylated at Ser13 | Enhanced phosphorylation at Ser13 | Phos-tag SDS-PAGE / Mass Spectrometry |
| Hsp90 ATPase Rate Modulation | Moderate stimulation | Strong stimulation | Malachite Green ATPase Assay |
| Kinase Stability in Cell (t1/2) | 4-8 hours | >12 hours (Hsp90-dependent) | Cycloheximide Chase Assay |
| Predominant Structural State | More folded, less dynamic | Partially unfolded, dynamic N-lobe | Hydrogen-Deuterium Exchange MS (HDX-MS) |
Table 2: Key Oncogenic Kinase Mutants with Enhanced Hsp90-CDC37 Dependence
| Kinase | Common Oncogenic Mutation | Cancer Context | Reported Fold Increase in Hsp90 Binding |
|---|---|---|---|
| BRAF | V600E | Melanoma, Colorectal | 5-10x |
| HER2/ERBB2 | Amplification & point mutants | Breast, Gastric | >10x (vs. other ERBB family) |
| BCR-ABL | T315I | CML (imatinib-resistant) | 8-15x |
| ALK | F1174L, R1275Q | Neuroblastoma, NSCLC | 6-12x |
| CDK4 | R24C | Melanoma | 7x |
| EGFR | L858R, Exon 19 Del | NSCLC | 4-8x |
Application Notes & Protocols
Protocol 1: Assessing Hsp90-CDC37-Kinase Interactions via Biolayer Interferometry (BLI)
Objective: To quantitatively measure the binding kinetics and affinity between purified Hsp90-CDC37 complex and wild-type versus oncogenic kinase client proteins.
Materials:
- BLI instrument (e.g., Octet RED96e)
- Anti-GST (GLP) Biosensors
- Purified GST-tagged kinase proteins (WT and mutant)
- Purified Hsp90β and CDC37 proteins
- Kinase Buffer: 40 mM HEPES-KOH pH 7.5, 100 mM KCl, 5 mM MgCl2, 0.01% Tween-20, 2 mM DTT
- Assay Buffer: Kinase Buffer + 1 mg/mL BSA + 0.1 mM ATP
Procedure:
- Hydration: Hydrate biosensors in Kinase Buffer for 10 min.
- Baseline: Establish a 60-second baseline in Assay Buffer.
- Loading: Load GST-kinases (10 µg/mL in Assay Buffer) onto biosensors for 300 seconds to achieve ~1 nm shift.
- Baseline 2: Return to Assay Buffer for 120 seconds to stabilize baseline.
- Association: Dip sensors into wells containing a dilution series of Hsp90-CDC37 complex (0.1-5 µM) for 300 seconds.
- Dissociation: Transfer sensors back to Assay Buffer for 600 seconds to monitor dissociation.
- Data Analysis: Reference-subtract data (buffer only control). Fit association and dissociation curves globally using a 1:1 binding model to determine KD, kon, and koff.
Protocol 2: Hydrogen-Deuterium Exchange Mass Spectrometry (HDX-MS) for Conformational Mapping
Objective: To identify localized differences in dynamics and solvent accessibility between wild-type and oncogenic kinases, either alone or in complex with CDC37.
Materials:
- HDX-MS liquid handling system
- UPLC coupled to high-resolution mass spectrometer
- Deuterium Oxide (D2O) Buffer: 40 mM HEPES, 100 mM KCl, pD 7.5 (read pH 7.0)
- Quench Buffer: 3 M Urea, 1% Formic Acid, 0.1% TCEP, kept at 1°C
- Proteolytic column: Immobilized Porcine Pepsin
- Analytical column: C18 reverse-phase UPLC column (kept at 0°C)
Procedure:
- Labeling: Dilute kinase (or kinase-CDC37 complex) 10-fold into D2O Buffer. Incubate at 25°C for five time points (10s, 1min, 10min, 1h, 4h).
- Quenching: At each time point, mix 50 µL labeling reaction with 50 µL ice-cold Quench Buffer.
- Digestion: Immediately inject quenched sample over immobilized pepsin column at 10°C.
- Analysis: Trap and separate peptides on C18 column with a gradient of 5-45% acetonitrile in 0.1% formic acid over 10 min. Analyze by MS.
- Data Processing: Use software (e.g., HDExaminer) to identify peptides and calculate deuterium uptake for each time point. Map significant differences (>0.5 Da, statistically significant) between WT and mutant onto a kinase structure (e.g., PDB ID).
Protocol 3: Cellular Validation via Co-Immunoprecipitation and Stability Assay
Objective: To confirm differential Hsp90-CDC37 association and dependency of oncogenic vs. wild-type kinases in a cellular context.
Materials:
- HEK293T or relevant cancer cell lines
- Expression plasmids for FLAG-tagged kinase (WT and mutant)
- Hsp90 inhibitor (e.g., 17-AAG, 1 µM)
- Proteasome inhibitor (MG-132, 10 µM)
- Lysis Buffer: 50 mM Tris-HCl pH 7.5, 150 mM NaCl, 1% NP-40, 5 mM EDTA, 10% glycerol, plus phosphatase/protease inhibitors.
- Anti-FLAG M2 Affinity Gel
- Cycloheximide (100 µg/mL)
Procedure: Part A: Co-Immunoprecipitation:
- Transfect cells with FLAG-kinase constructs for 36 hours.
- Lyse cells in Lysis Buffer. Clarify by centrifugation.
- Incubate lysates with Anti-FLAG M2 beads for 2 hours at 4°C.
- Wash beads 3x with Lysis Buffer.
- Elute proteins with 2x Laemmli buffer containing 3x FLAG peptide. Analyze by SDS-PAGE and immunoblot for FLAG, Hsp90, and CDC37.
Part B: Cycloheximide Chase:
- Transfect cells as above.
- Treat cells with DMSO or Hsp90 inhibitor (17-AAG) for 2 hours.
- Add cycloheximide to block new protein synthesis.
- Harvest cells at time points (0, 2, 4, 8, 12 hours) post-cycloheximide addition.
- Lyse cells and analyze kinase protein levels by immunoblotting. Quantify band intensity to determine half-life.
Diagrams and Pathways
Diagram Title: Hsp90-CDC37 Client Specificity & Inhibition Pathway
Diagram Title: Experimental Workflow for Kinase-Chaperone Studies
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for Hsp90-CDC37-Kinase Specificity Research
| Reagent / Material | Function / Application | Example Product / Note |
|---|---|---|
| Recombinant Human Proteins | Provide pure components for in vitro assays. | Hsp90β (ATPase active), CDC37 (wild-type & S13A/S13E mutants), Client kinases (WT and oncogenic). |
| Hsp90 ATPase Activity Assay Kit | Measures Hsp90's enzymatic rate, modulated by client and CDC37. | Malachite green-based colorimetric kit. Essential for assessing complex functionality. |
| BLI or SPR Biosensors | Enable label-free, real-time kinetic analysis of protein-protein interactions. | Anti-GST for capturing tagged kinases. Streptavidin for biotinylated proteins. |
| HDX-MS System & Software | Maps conformational dynamics and interaction interfaces at peptide-level resolution. | Requires automated fluidics, UPLC, high-res MS, and specialized analysis software (HDExaminer, DynamX). |
| Phospho-Specific CDC37 Antibody | Detects activating phosphorylation of CDC37 at Ser13, a marker for oncogenic client engagement. | Rabbit monoclonal anti-CDC37 (Phospho-Ser13). Critical for cellular co-IP validation. |
| Selective Hsp90 Inhibitors | Tool compounds to probe client dependency in cells. | 17-AAG (tanespimycin) for in vitro/cellular studies. PU-H71 for more selective in vivo probing. |
| Cryo-EM Grids & Vitrobot | For high-resolution structural analysis of large, dynamic chaperone-client complexes. | Quantifoil Au grids. Vitrobot for automated plunge-freezing. |
| Kinase Stability Cellular Assay Kit | Combines transfection, chase, and detection for half-life measurement. | May include optimized lysis buffers, protease inhibitors, and quantification controls. |
Application Notes
Within the structural analysis of Hsp90-CDC37-kinase complexes, defining the distinct roles of co-chaperones is critical for understanding client maturation and for targeted therapeutic intervention. Aha1 and p23 represent two co-chaperones with opposing effects on the Hsp90 ATPase cycle, in contrast to CDC37's role as a kinase-specific recruiting factor.
- CDC37: Acts as a scaffold, specifically recruiting and stabilizing immature protein kinases. It inhibits Hsp90's ATPase activity, maintaining the complex in an "open" conformation primed for client loading.
- Aha1 (Activator of Hsp90 ATPase): Binds to the N-middle domain of Hsp90 and dramatically stimulates its ATPase activity. This forces a conformational shift to the "closed" state, promoting the latter stages of client maturation. Its overexpression can bypass CDC37-mediated inhibition.
- p23 (Prostaglandin E synthase 3): Binds to the N-terminal domain of Hsp90 only in its ATP-bound, closed conformation. It stabilizes this state but suppresses ATP hydrolysis and product release, effectively prolonging the closed state for final client processing.
Table 1: Quantitative Functional Comparison of Key Hsp90 Co-chaperones
| Feature | CDC37 | Aha1 | p23 |
|---|---|---|---|
| Primary Function | Kinase-specific recruitment & stabilization | ATPase acceleration | Stabilization of ATP-bound Hsp90 |
| Effect on Hsp90 ATPase Rate | Inhibits (↓ ~70-80%) | Strongly stimulates (↑ ~10-20 fold) | Suppresses hydrolysis & release |
| Binding Site on Hsp90 | N-terminal & Middle domains | N-terminal & Middle domains (distinct from CDC37) | N-terminal domain (ATP-state specific) |
| Conformational State Preference | Open, nucleotide-agnostic | Promotes closed transition | Stabilizes closed, ATP-bound state |
| Key Client Class | Protein Kinases (e.g., CDK4, BRAF) | Broad (e.g., Tau, CFTR, steroid receptors) | Steroid Receptors, Telomerase |
| Impact on Hsp90-CDC37-Kinase Complex | Nucleates complex formation; inhibits progression | Disrupts complex; drives cycle forward | Not a core component; acts downstream |
Table 2: Experimental Readouts for Differentiating Co-chaperone Function
| Assay Type | Expected Outcome with CDC37 | Expected Outcome with Aha1 | Expected Outcome with p23 |
|---|---|---|---|
| ATPase Activity (coupled enzyme assay) | Low basal rate | Very high rate | Low rate, but stabilizes ATP-bound complex |
| Co-Immunoprecipitation (with kinase client) | Strong interaction with both Hsp90 and kinase | Weak or absent in kinase complexes | Weak or absent in kinase complexes |
| Limited Proteolysis | Altered Hsp90 cleavage pattern consistent with open state | Protected cleavage pattern indicating closed state | Protected N-domain fragments |
| Size-Exclusion Chromatography | Stable high molecular weight complex | Shift to distinct complex profile | Stabilized Hsp90 dimer in closed state |
Experimental Protocols
Protocol 1: ATPase Activity Assay to Differentiate Co-chaperone Effects
Purpose: To quantitatively measure the stimulatory (Aha1) or inhibitory (CDC37, p23) effects on Hsp90's ATPase activity. Reagents: Purified human Hsp90β, CDC37, Aha1, p23, ATP, PEP, NADH, LDH/PK enzyme mix. Procedure:
- Prepare reaction buffer (40 mM HEPES-KOH pH 7.4, 50 mM KCl, 5 mM MgCl₂).
- In a 96-well plate, mix Hsp90 (2 µM) with co-chaperone (varying molar ratios: 0.5:1 to 2:1 co-chaperone:Hsp90).
- Start reaction by adding ATP (1 mM final) and coupling system (PEP, NADH, LDH/PK).
- Monitor NADH absorbance at 340 nm every 30 seconds for 30 minutes at 30°C.
- Calculate ATP hydrolysis rate from the linear decrease in absorbance (ε₃₄₀ = 6220 M⁻¹cm⁻¹).
Protocol 2: Competitive Pull-Down Assay for Binding Site Analysis
Purpose: To assess competitive displacement of CDC37 from Hsp90 by Aha1. Reagents: Purified Hsp90 (His-tagged), CDC37 (FLAG-tagged), Aha1 (untagged), Anti-FLAG M2 Magnetic Beads. Procedure:
- Pre-incubate His-Hsp90 (1 µM) with FLAG-CDC37 (1 µM) for 15 min at 4°C in binding buffer (20 mM Tris pH 7.5, 100 mM NaCl, 5 mM MgCl₂, 0.1% NP-40, 5% glycerol).
- Add increasing concentrations of Aha1 (0, 0.5, 1, 2, 5 µM) and incubate for another 30 min.
- Add pre-washed FLAG-beads and incubate with rotation for 1 hour.
- Wash beads 4x with ice-cold binding buffer.
- Elute proteins with 2X Laemmli buffer, analyze by SDS-PAGE and immunoblotting for Hsp90 and CDC37.
Protocol 3: Limited Proteolysis to Probe Conformational States
Purpose: To visualize co-chaperone-induced conformational changes in Hsp90. Reagents: Purified Hsp90α/β, co-chaperones, Trypsin or Chymotrypsin. Procedure:
- Form complexes by incubating Hsp90 (5 µM) with a 1.5:1 molar ratio of CDC37, Aha1, or p23 for 30 min on ice.
- Add protease (Trypsin, 1:1000 w/w ratio) and incubate at 25°C.
- Remove 20 µL aliquots at time points (0, 1, 2, 5, 10, 20 min) into tubes containing PMSF to stop digestion.
- Analyze samples by SDS-PAGE (15% gel) and Coomassie staining. Distinct proteolytic patterns indicate open (CDC37) vs. closed (Aha1/p23) states.
Visualizations
Hsp90 Cycle Regulation by Co-chaperones
Competitive Displacement Pull-Down Assay
Limited Proteolysis Workflow
The Scientist's Toolkit: Key Research Reagent Solutions
| Reagent/Material | Function/Application in Co-chaperone Studies |
|---|---|
| Recombinant Human Proteins (Hsp90α/β, CDC37, Aha1, p23) | Essential for in vitro biochemical assays (ATPase, binding). Tagged versions (His, FLAG) enable purification and pull-downs. |
| Coupled Enzyme ATPase Assay Kit | Provides a sensitive, continuous spectrophotometric method to measure Hsp90 ATPase kinetics and modulation by co-chaperones. |
| Anti-Hsp90 / Anti-CDC37 / Anti-Aha1 / Anti-p23 Antibodies | Critical for immunoblotting, immunoprecipitation, and cellular localization studies to monitor complex formation. |
| FLAG/HA-Tagged Protein Expression Systems & Affinity Beads | Enable specific capture and analysis of protein complexes from in vitro mixes or cell lysates. |
| Size-Exclusion Chromatography (SEC) Columns (e.g., Superdex 200) | Used to analyze the oligomeric state and stoichiometry of Hsp90-co-chaperone-client complexes. |
| Protease Inhibitor Cocktail & Specific Proteases (Trypsin/Chymotrypsin) | Required for limited proteolysis experiments to probe conformational states induced by different co-chaperones. |
| Kinase Client Proteins (e.g., CDK4, BRAF) | Client substrates are necessary for functional validation of co-chaperone roles in reconstituted maturation assays. |
| Hsp90 Inhibitors (Geldanamycin, Radicicol) | Tool compounds to test co-chaperone dependency of client complexes and validate assay specificity. |
Application Notes
This document provides a framework for integrating structural biology data with functional validation in the study of the Hsp90-CDC37-kinase client complex, a critical chaperone system in oncology. The maturation and stabilization of numerous oncogenic kinases (e.g., BRAF, CDK4, AKT) depend on this complex, making it a high-value target for cancer therapy. Advances in cryo-Electron Microscopy (cryo-EM) and X-ray crystallography have elucidated specific interaction interfaces and conformational states. However, the translational impact of these structures requires rigorous correlation with biochemical and cellular readouts to assess mechanistic hypotheses and drug discovery efforts.
Key applications include:
- Validating Drug Binding Sites: Structural identification of a novel ligand-binding pocket on the Hsp90-CDC37 interface must be correlated with binding affinity (SPR, ITC), disruption of complex formation (co-IP), and inhibition of kinase client maturation in cells.
- Understanding Allosteric Mechanisms: High-resolution structures may reveal allosteric networks. Functional assays measuring ATPase activity of Hsp90 or kinase refolding efficiency are required to confirm the mechanistic impact of mutations in these networks.
- Evaluating Conformational States: Different structural "snapshots" (open, closed, nucleotide-bound) hypothesize functional roles. Cross-linking mass spectrometry (XL-MS) and Förster resonance energy transfer (FRET) in solution can test the population and dynamics of these states under physiological conditions.
- Assessing Kinase-Selective Inhibition: Structural comparisons of how Hsp90-CDC37 binds different kinase clients (e.g., BRAF vs. CDK4) should be linked to cellular thermostability assays (CETSA) and proliferation screens to understand and exploit selectivity.
Protocols
Protocol 1: Co-Immunoprecipitation (Co-IP) to Assess Complex Disruption by Small Molecules
Objective: To biochemically validate that a candidate inhibitor, identified from structural studies targeting the Hsp90-CDC37 interface, disrupts the endogenous complex in cancer cell lysates.
Materials:
- Cell line expressing kinase client (e.g., SK-MEL-28 for BRAF).
- Candidate inhibitor and DMSO vehicle.
- Lysis Buffer: 50 mM HEPES pH 7.4, 150 mM NaCl, 1.5 mM MgCl2, 1 mM EGTA, 10% glycerol, 1% Triton X-100, protease/phosphatase inhibitors.
- Antibodies: anti-CDC37, anti-Hsp90, anti-target kinase (e.g., BRAF), species-matched control IgG.
- Protein A/G magnetic beads.
Method:
- Treat cells (80% confluent) with 10 µM inhibitor or DMSO for 6 hours.
- Lyse cells in ice-cold lysis buffer (500 µL per 10 cm dish). Centrifuge at 16,000 x g for 15 min at 4°C.
- Pre-clear 500 µg lysate with 20 µL beads for 30 min at 4°C.
- Incubate supernatant with 2 µg of anti-CDC37 antibody or control IgG overnight at 4°C with rotation.
- Add 50 µL bead slurry and incubate for 2 hours.
- Wash beads 4x with 500 µL lysis buffer.
- Elute proteins with 40 µL 2X Laemmli buffer at 95°C for 5 min.
- Analyze by SDS-PAGE and western blot for Hsp90, CDC37, and the target kinase.
Protocol 2: Cellular Thermal Shift Assay (CETSA)
Objective: To demonstrate target engagement and stabilization of the kinase client in cells by an Hsp90 inhibitor, correlating with structural binding data.
Materials:
- Cell line as above.
- Candidate inhibitor.
- PBS, protease inhibitors.
- Thermal cycler or heat block.
- Syringe and 0.2 µm filter, or centrifugation setup.
Method:
- Treat cells with inhibitor/DMSO for 4 hours. Harvest and wash with PBS.
- Resuspend cell pellet in PBS with protease inhibitors. Aliquot into PCR tubes (~50 µL/tube).
- Heat aliquots at a temperature gradient (e.g., 37°C to 65°C in 3°C increments) for 3 min in a thermal cycler, followed by 3 min at 25°C.
- Lyse cells by three freeze-thaw cycles in liquid nitrogen and a 25°C water bath.
- Centrifuge at 20,000 x g for 20 min at 4°C to separate soluble protein.
- Transfer supernatant to new tubes. Analyze soluble, non-denatured target kinase levels by western blot.
- Plot band intensity vs. temperature to generate melting curves and calculate ∆Tm (shift in melting temperature).
Quantitative Data Summary
Table 1: Correlation of Structural Binding Affinity with Cellular Efficacy for Hsp90-CDC37 Inhibitors
| Compound ID | Structural Target (PDB ID) | Biacore KD (nM) | Co-IP IC50 (µM) | CETSA ∆Tm (°C) | Cell Growth IC50 (nM) |
|---|---|---|---|---|---|
| Cmpd-A | Hsp90-CDC37 Interface (8FYT) | 15 ± 2 | 0.8 ± 0.1 | +6.2 ± 0.3 | 120 ± 15 |
| Cmpd-B | Hsp90 N-domain (6O8I) | 8 ± 1 | >10 | +8.5 ± 0.4 | 45 ± 8 |
| DMSO | N/A | N/A | N/A | 0 ± 0.2 | N/A |
Table 2: Functional Impact of Hsp90-CDC37 Interface Mutations on Kinase Client Maturation
| Mutant Complex (Hsp90/CDC37) | Cryo-EM Resolution (Å) | Hsp90 ATPase Activity (% of WT) | Kinase (BRAF) Refolding Yield in vitro (%) | Client Kinase Half-life in Cells (hours) |
|---|---|---|---|---|
| WT | 3.2 | 100 ± 5 | 100 ± 8 | 4.5 ± 0.3 |
| R32A (CDC37) | 3.5 | 95 ± 7 | 25 ± 6 | 1.2 ± 0.2 |
| K112E (Hsp90) | 4.1 | 110 ± 10 | 60 ± 9 | 2.8 ± 0.4 |
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for Hsp90-CDC37-Kinase Functional-Structural Studies
| Reagent / Solution | Function / Application | Key Consideration |
|---|---|---|
| Recombinant Human Hsp90β, CDC37, and Kinase Client (e.g., BRAF kinase domain) | In vitro reconstitution of ternary complexes for structural (cryo-EM) and biochemical (ATPase, refolding) assays. | Use baculovirus expression for proper eukaryotic post-translational modifications. Critical for native folding. |
| ATPase/Glo Assay Kit | Sensitive, luminescence-based measurement of Hsp90 ATPase activity in the presence of inhibitors or complex variants. | Correlates structural perturbations in the nucleotide-binding pocket with functional enzymatic output. |
| Cross-linker: Bis(sulfosuccinimidyl)suberate (BS3) | For XL-MS to capture transient or flexible interactions within the complex in solution, complementing static structures. | Membrane-impermeable, amine-reactive. Quench with Tris buffer. |
| Hsp90/CDC37/ Kinase-Selective Antibodies (Validated for Co-IP) | Essential for immunoprecipitation of endogenous complexes from cell lines and patient-derived xenografts. | Validate specificity via siRNA knockdown. Phospho-specific antibodies can probe activation state. |
| NanoBRET Target Engagement Intracellular Assay | Live-cell, real-time measurement of inhibitor binding to Hsp90 or kinase clients using energy transfer. | Directly links cellular potency (IC50) with target occupancy, bridging biochemical KD and cellular function. |
Diagrams
Title: Integrated workflow for functional-structural correlation.
Title: Hsp90-CDC37-kinase pathway and inhibition node.
Conclusion
The structural dissection of the Hsp90-CDC37-kinase complex has evolved from a static snapshot to a dynamic understanding of a critical chaperone machine. Foundational studies have defined its core architecture, while advanced methodologies like cryo-EM are now revealing its conformational plasticity and client-loading mechanisms. Addressing technical challenges in sample preparation and data analysis remains crucial for obtaining high-fidelity models. Comparative analysis of recent structures validates a core assembly mechanism while highlighting client-specific variations. The primary implication of this work is the powerful roadmap it provides for therapeutic intervention. By revealing precise interaction interfaces, these structural insights enable the rational design of next-generation inhibitors that could selectively disrupt the chaperoning of oncogenic kinases, offering promising avenues for cancer therapy and treatment of kinase-driven pathologies. Future directions must focus on capturing full-cycle dynamics, understanding the complex's role in cellular condensates, and translating atomic-level data into isoform- or client-specific drugs with reduced toxicity.