SEC-MALS vs DLS: Choosing the Right Technique for Membrane Protein Size, Mass, and Oligomerization Analysis
This article provides a comprehensive comparison of Size Exclusion Chromatography with Multi-Angle Light Scattering (SEC-MALS) and Dynamic Light Scattering (DLS) for the characterization of membrane proteins.
SEC-MALS vs DLS: Choosing the Right Technique for Membrane Protein Size, Mass, and Oligomerization Analysis
Abstract
This article provides a comprehensive comparison of Size Exclusion Chromatography with Multi-Angle Light Scattering (SEC-MALS) and Dynamic Light Scattering (DLS) for the characterization of membrane proteins. Aimed at researchers and drug developers, we explore the foundational principles, practical applications, and inherent challenges of each technique. We detail method-specific protocols for solubilized proteins in detergents or nanodiscs, offer troubleshooting strategies for common pitfalls like aggregation and buffer interference, and present a direct validation framework comparing data outputs. The goal is to empower scientists to select and optimize the most appropriate method for accurate determination of molar mass, hydrodynamic size, aggregation state, and oligomeric stability—critical parameters for structural biology and biotherapeutic development.
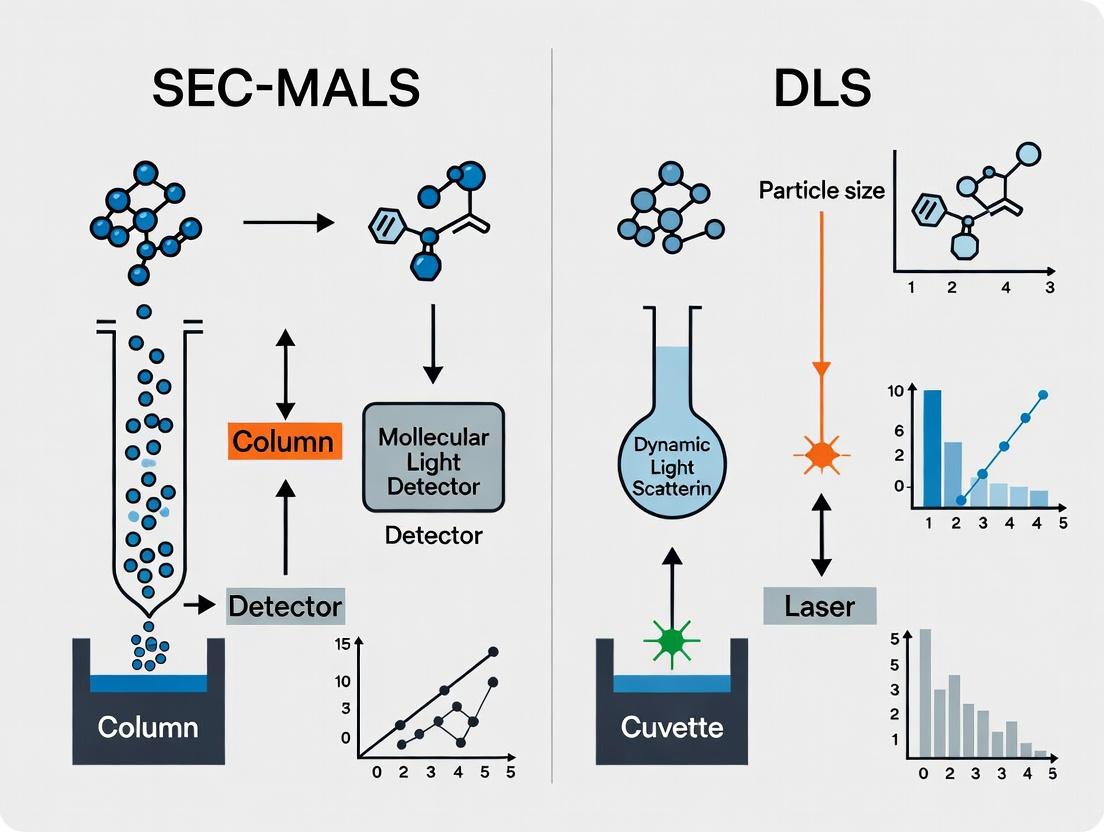
Understanding the Core Principles: How SEC-MALS and DLS Work for Membrane Proteins
Performance Comparison: SEC-MALS vs. DLS for Membrane Protein Analysis
Membrane proteins require careful handling in solution post-extraction from the lipid bilayer. This guide compares Size Exclusion Chromatography with Multi-Angle Light Scattering (SEC-MALS) and Dynamic Light Scattering (DLS), two key techniques for assessing their stability, oligomeric state, and monodispersity in detergents or nanodiscs.
Table 1: Direct Performance Comparison of SEC-MALS and DLS
| Performance Metric | SEC-MALS | Batch DLS | Comments & Supporting Data |
|---|---|---|---|
| Primary Output | Absolute Molar Mass (Mw); Hydrodynamic radius (Rh) via on-line DLS (if equipped). | Hydrodynamic radius (Rh); size distribution; qualitative aggregation assessment. | SEC-MALS provides mass directly without shape assumptions. DLS Rh assumes a spherical model. |
| Sample Requirement | ~50-100 µg (high purity). | ~5-50 µL at low µM concentration. | DLS requires less material but is less informative for polydisperse samples. |
| Resolution of Heterogeneity | High. Chromatographic separation resolves species by hydrodynamic volume before detection. | Very Low. Provides an intensity-weighted average size; cannot resolve discrete species in a mixture. | SEC-MALS can separate and quantify monomer, dimer, and aggregate peaks. DLS will show a single, broad peak for such mixtures. |
| Aggregation Detection | Quantitative. MALS quantifies mass fraction of high-MW aggregates in each eluting slice. | Semi-quantitative. Polydispersity Index (PdI) indicates heterogeneity; cannot quantify mass fraction. | SEC-MALS data: A sample showing a dominant monomer peak (Mw = 150 kDa) with a 5% mass fraction of aggregate > 1000 kDa. DLS data for the same sample may show PdI > 0.3, obscuring the primary species size. |
| Impact of Viscosity/Detergent | Minimized. SEC separates protein from most micelles; MALS is largely insensitive to viscosity. | High. Viscous solutions and detergent micelles contribute directly to the scattering signal. | For a GPCR in DDM, DLS may report an Rh of 8-10 nm (protein + large micelle). SEC-MALS reports the protein's mass independently of the micelle. |
| Analysis Speed | ~30-60 minutes per run. | ~2-5 minutes per measurement. | DLS offers rapid screening. SEC-MALS provides definitive analysis but is slower. |
| Key Advantage | Orthogonal Data: Simultaneous, absolute Mw and Rh from a single experiment. | Speed & Sensitivity: Rapid assessment of sample monodispersity with minimal material. |
Experimental Protocols for Cited Comparisons
Protocol 1: SEC-MALS Analysis of a Membrane Protein in DDM
- Sample Prep: Purify protein in 20 mM Tris, 150 mM NaCl, 0.05% n-Dodecyl-β-D-Maltopyranoside (DDM), pH 8.0. Concentrate to ~5 mg/mL.
- SEC: Inject 100 µL onto a silica-based SEC column (e.g., Yarra 3µM SEC-3000) equilibrated in the same buffer. Flow rate: 0.5 mL/min.
- Detection: The eluent passes through a UV detector (280 nm), a MALS detector (e.g., Wyatt miniDAWN TREOS), and a differential refractometer (dRI).
- Data Analysis: Use ASTRA or similar software. The dRI sets concentration, MALS determines absolute molar mass across the peak. On-line DLS (if available) calculates Rh.
Protocol 2: Batch DLS Screening for Membrane Protein Stability
- Sample Prep: Use the same buffer as Protocol 1. Clarify sample by centrifugation at 15,000 x g for 10 min.
- Loading: Pipette 25 µL of supernatant into a low-volume quartz cuvette. Avoid bubbles.
- Measurement: Set instrument (e.g., Malvern Zetasizer) to 25°C, equilibrate 2 min. Perform 10-15 measurements of 10 seconds each.
- Analysis: Software reports intensity-based size distribution and PdI. An Rh main peak with PdI < 0.2 indicates a monodisperse sample.
Visualization of Workflows
SEC-MALS Analysis Workflow
DLS vs. SEC-MALS Strategy Comparison
The Scientist's Toolkit: Research Reagent Solutions
| Reagent / Material | Function in Membrane Protein Analysis |
|---|---|
| n-Dodecyl-β-D-Maltopyranoside (DDM) | Mild, non-ionic detergent commonly used to solubilize and stabilize membrane proteins from the lipid bilayer. |
| SMA / DIBMA Copolymers | "Styrene Maleic Acid" or "Diisobutylene Maleic Acid" copolymers that directly solubilize proteins into native nanodiscs (SMALPs/DIBMALPs), bypassing detergent. |
| MSP Nanodiscs | Membrane Scaffold Proteins that form discrete, tunable lipid bilayers to reconstitute proteins in a more native-like environment than detergent micelles. |
| Size Exclusion Columns (e.g., Superose, Yarra) | High-resolution silica or polymer columns for separating protein complexes based on hydrodynamic size, critical for SEC-MALS. |
| Stabilizing Lipids (e.g., POPC, POPG) | Defined lipids added to buffers or nanodisc preps to enhance membrane protein stability and function during analysis. |
| CHAPS / CHAPSO Detergents | Zwitterionic detergents useful for solubilizing certain protein classes (e.g., GPCRs, ion channels) with different stability profiles than DDM. |
Within membrane protein research, determining accurate molar mass and oligomeric state is critical but challenging. This comparison guide objectively evaluates Size-Exclusion Chromatography with Multi-Angle Light Scattering (SEC-MALS) against Dynamic Light Scattering (DLS) and other alternatives, focusing on their application for membrane proteins in detergent solutions. The broader thesis contends that while DLS offers rapid sizing, SEC-MALS provides superior, shape-independent absolute molar mass, which is indispensable for characterizing labile membrane protein complexes.
Comparative Performance Analysis
Table 1: Core Technique Comparison for Membrane Protein Analysis
| Parameter | SEC-MALS | Batch DLS | SEC-UV/RI |
|---|---|---|---|
| Primary Output | Absolute Molar Mass (g/mol) | Hydrodynamic Radius (Rh) | Relative Molecular Weight |
| Shape Dependence | Independent (from first principles) | High (assumes spherical model) | High (requires shape/vis assumption) |
| Sample State | Separated (Chromatographic) | Ensemble (Polydisperse) | Separated (Chromatographic) |
| Detergent Compatibility | High (on-line separation from micelles) | Challenging (signal dominated by empty micelles) | High (with careful calibration) |
| Key Advantage | Direct, absolute mass without standards or models | Fast, low sample consumption | Widely available, simple |
| Major Limitation | More complex setup, higher sample need | Cannot deconvolute protein from micelle | Indirect, requires standards |
| Typical Precision | ~2-5% molar mass | ~5-10% Rh | ~10-20% molar mass |
Table 2: Experimental Data from a GPCR Study (in DDM detergent)
| Analytic | SEC-MALS Molar Mass (kDa) | DLS Rh (nm) | Expected Mass (kDa) | Oligomeric State Determined |
|---|---|---|---|---|
| Protein + DDM Micelle | 145 ± 3 | 6.8 ± 0.5 | -- | -- |
| Protein Contribution | 72 ± 2 | N/A* | 68.5 | Monomer |
| Empty DDM Micelle | 73 ± 4 | 5.5 ± 0.3 | ~70 | -- |
*DLS could not resolve protein-specific signal from micelle background.
Experimental Protocols
Protocol 1: SEC-MALS for Membrane Protein Molar Mass
Objective: Determine the absolute molar mass of a membrane protein in solution with detergent.
- System Equilibration: Equilibrate the SEC column (e.g., Superose 6 Increase) with running buffer (e.g., 20 mM HEPES, 150 mM NaCl, 0.5 mM DDM) at 0.5 mL/min until UV baseline is stable.
- Calibration: Normalize the MALS detector using pure toluene or a BSA monomer standard. Align the inter-detector delay volume between UV, MALS, and RI detectors.
- Sample Preparation: Concentrate the purified protein in identical running buffer to >1 mg/mL. Centrifuge at 15,000g for 10 minutes to remove aggregates.
- Injection & Separation: Inject 50-100 μL of sample. Monitor elution with UV (280 nm), MALS (simultaneous measurement at multiple angles), and RI detectors.
- Data Analysis: Use the Astra or equivalent software. The molar mass at each chromatographic slice is calculated directly from the MALS and concentration signals (from UV or RI) using the Zimm equation. The protein mass is identified by a co-eluting peak distinct from the empty micelle peak.
Protocol 2: Batch DLS for Hydrodynamic Size
Objective: Measure the hydrodynamic radius of the protein-detergent complex.
- Buffer Filtration: Filter all buffers through a 0.02 μm filter to remove dust.
- Sample Preparation: Dialyze or dilute protein into filtered running buffer. Centrifuge at 15,000g for 10 minutes.
- Measurement: Load 20-50 μL of supernatant into a ultra-low volume quartz cuvette. Place in instrument thermostatted at 20°C.
- Data Acquisition: Perform 10-15 measurements of 10 seconds each.
- Analysis: Use cumulants analysis to obtain the intensity-weighted mean hydrodynamic radius (Z-average) and polydispersity index (PdI). For membrane proteins, the signal represents the aggregate of protein-detergent complexes and free micelles.
Visualizing the Workflow and Data Interpretation
Title: SEC-MALS Absolute Molar Mass Workflow
Title: From Light Scattering to Molar Mass
The Scientist's Toolkit: Research Reagent Solutions
| Item | Function in Membrane Protein SEC-MALS/DLS |
|---|---|
| Mild Detergents (DDM, LMNG) | Solubilizes membrane proteins while maintaining native structure and activity. |
| Size-Exclusion Columns (e.g., Superose 6 Increase) | Separates protein-detergent complexes from empty micelles and aggregates. |
| MALS Detector (e.g., Wyatt DAWN) | Measures scattered light intensity at multiple angles simultaneously for direct molar mass calculation. |
| Refractive Index (RI) Detector | Provides precise concentration measurement of eluting species, independent of UV absorbance. |
| Online UV Detector (280 nm) | Monitors protein elution based on aromatic amino acid absorbance. |
| Stable, Filtered Buffers | Provides consistent solvent conditions and minimizes dust for light scattering. |
| BSA Monomer Standard | Used for system calibration and normalization verification. |
| Quartz Flow Cell/Cuvette | Provides clean, low-scatter optical path for light scattering measurements. |
Dynamic Light Scattering (DLS) is a cornerstone analytical technique for measuring the hydrodynamic size, size distribution, and polydispersity of nanoparticles, proteins, and macromolecular complexes in solution. For researchers investigating membrane proteins—notoriously challenging due to their instability outside lipid bilayers—DLS provides a rapid, non-invasive assessment of sample monodispersity and aggregation state prior to more intensive structural studies. This guide compares DLS performance with alternative methods, particularly within the thesis framework comparing Size Exclusion Chromatography with Multi-Angle Light Scattering (SEC-MALS) for membrane protein characterization.
How DLS Works: Core Principle
DLS measures the Brownian motion of particles in suspension. Smaller particles move rapidly, while larger ones diffuse more slowly. The instrument (a goniometer) shines a monochromatic laser through the sample, and a detector measures the intensity fluctuations of scattered light over time. An autocorrelation function analyzes these fluctuations to determine the diffusion coefficient (D), which is then converted to hydrodynamic radius (Rh) via the Stokes-Einstein equation. The polydispersity index (PDI) quantifies the breadth of the size distribution.
Direct Comparison: DLS vs. SEC-MALS for Membrane Proteins
The table below summarizes a key performance comparison between batch-mode DLS and online SEC-MALS for analyzing a model detergent-solubilized membrane protein.
Table 1: Performance Comparison of DLS and SEC-MALS for Membrane Protein Analysis
| Parameter | Batch-Mode DLS | Online SEC-MALS | Experimental Implication |
|---|---|---|---|
| Sample Throughput | Very High (seconds/minutes per sample) | Moderate (10-30 minutes per run) | DLS excels for rapid screening of buffer conditions and stability. |
| Sample Consumption | Low (as little as 2-12 µL) | Moderate-High (typically 50-100 µL) | DLS is advantageous for precious membrane protein samples. |
| Size Resolution | Low. Reports an intensity-weighted distribution. | High. Resolves species by hydrodynamic volume before MALS analysis. | SEC-MALS can separate and individually analyze monomers, oligomers, and aggregates. |
| Size Range | ~0.3 nm to 10 µm | Limited by SEC column (typically ~2x104 to 107 Da) | DLS can detect large aggregates that may be excluded from SEC columns. |
| Aggregate Detection | Excellent for large aggregates; poor for small oligomers. | Excellent for resolving small oligomers and large aggregates. | SEC-MALS is superior for quantifying specific oligomeric states. |
| Absolute Mass | No. Provides Rh only. | Yes. MALS provides absolute molecular weight (Mw). | Critical for confirming complex stoichiometry. |
| Impact of Viscosity | Highly sensitive; requires accurate temperature control. | Accounted for, as SEC separates by size in the same buffer. | DLS measurements require precise buffer viscosity data for accuracy. |
| Key Metric for Polydispersity | Polydispersity Index (PDI). PDI < 0.1 is monodisperse. | Mw/Mn from MALS, and peak shape from UV/RI. | Both indicate sample homogeneity but via different principles. |
Supporting Experimental Data: A 2023 study of a G protein-coupled receptor (GPCR) in detergent micelles reported a DLS-derived Rh of 8.2 nm with a PDI of 0.22, suggesting a polydisperse sample. Subsequent SEC-MALS analysis resolved this into two major peaks: Peak 1 (70% of mass) with an Mw of 132 kDa (monomer+micelle) and Peak 2 (30%) with an Mw of 390 kDa (trimer/aggregate). This highlights DLS's role as a rapid prescreen and SEC-MALS as a detailed orthogoganal analysis tool.
Experimental Protocols
Protocol 1: Standard DLS Measurement for Membrane Protein Stability Screening
- Objective: To rapidly assess the aggregation state and monodispersity of a solubilized membrane protein sample.
- Materials: Purified membrane protein in detergent/buffer, bench-top DLS instrument, 0.02 µm filtered buffer, low-volume disposable cuvettes (e.g., 12 µL).
- Procedure:
- Centrifuge sample at >15,000 x g for 10 minutes to remove dust and large aggregates.
- Filter the reference buffer through a 0.02 µm syringe filter.
- Load 12 µL of clarified sample into a clean, disposable microcuvette.
- Equilibrate in the instrument at 20°C (or desired temperature) for 2 minutes.
- Set measurement parameters: laser wavelength (e.g., 633 nm), scattering angle (e.g., 173°), and number of runs (typically 10-15).
- Perform measurement. The software automatically calculates the correlation function.
- Analyze data to extract the Z-average hydrodynamic diameter (Rh), PDI, and intensity size distribution.
- Data Interpretation: A single, narrow peak in the intensity distribution with a PDI < 0.1 is ideal. A high PDI (>0.2) or multiple peaks indicate polydispersity and potential aggregation.
Protocol 2: SEC-MALS Analysis for Absolute Molecular Weight Determination
- Objective: To determine the absolute molecular weight and oligomeric state of a membrane protein-detergent complex.
- Materials: HPLC system, SEC column (e.g., Superdex 200 Increase), MALS detector, refractive index (RI) detector, UV detector, degassed running buffer (with detergent above CMC), 0.1 µm filtered sample.
- Procedure:
- Equilibrate the SEC column with running buffer at a constant flow rate (e.g., 0.5 mL/min) until a stable RI baseline is achieved.
- Calibrate the MALS detector's photodiodes using pure toluene or a standardized protein.
- Filter the protein sample (50-100 µL) through a 0.1 µm spin filter.
- Inject sample onto the column.
- As the sample elutes, it passes sequentially through the UV, MALS, and RI detectors.
- Use dedicated software (e.g., ASTRA) to analyze the combined data from all detectors across the elution peak.
- Data Interpretation: The software calculates the absolute molecular weight (Mw) at each data slice across the elution peak. A constant Mw across the peak indicates a monodisperse species. The Mw includes the mass of the protein and the bound detergent/lipid.
Visualizing the Analytical Workflow
Title: Complementary DLS and SEC-MALS Workflow for Protein Analysis
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Materials for Membrane Protein DLS/SEC-MALS Analysis
| Item | Function in Analysis | Key Consideration for Membrane Proteins |
|---|---|---|
| Mild Detergents (e.g., DDM, LMNG) | Solubilize and stabilize membrane proteins by mimicking the lipid bilayer. | Critical for maintaining native structure and preventing aggregation. Choice affects micelle size in DLS/MALS. |
| Size Exclusion Columns (e.g., Superdex 200 Increase) | Separate protein complexes by hydrodynamic volume for SEC-MALS. | Must be compatible with detergents. Increased length provides better resolution of oligomers. |
| Amicon Ultra Centrifugal Filters | Concentrate dilute membrane protein samples prior to analysis. | Membrane material must be detergent-resistant (e.g., low protein binding regenerated cellulose). |
| 0.02 µm & 0.1 µm Filters | Remove dust and aggregates to eliminate scattering artifacts in DLS and prevent column blockage in SEC. | Essential for obtaining clean, interpretable data. |
| Stable Buffer Systems (e.g., HEPES, Tris) | Maintain constant pH and ionic strength during measurement. | Buffer must contain detergent above its critical micelle concentration (CMC) at all times. |
| MALS Calibration Standard (e.g., BSA, Toluene) | Calibrate the light scattering detectors for absolute molecular weight determination. | Protein standards should be run in the exact same buffer as the sample for accurate results. |
Within membrane protein research, accurately characterizing molar mass, size, and sample heterogeneity is critical for understanding structure-function relationships and developing therapeutics. Size Exclusion Chromatography with Multi-Angle Light Scattering (SEC-MALS) and Dynamic Light Scattering (DLS) are two prominent techniques, but they measure fundamentally different parameters. This guide objectively compares their performance within a research thesis focused on membrane protein analysis.
Core Measurement Principles
SEC-MALS measures the absolute molar mass (M) of a solute by independently determining the intensity of light scattered at multiple angles, coupled with concentration data from a UV or refractive index (RI) detector. In a separation column, it separates species by hydrodynamic size, allowing the determination of mass, size (radius of gyration, Rg), and detection of aggregates or degradation products.
DLS measures the hydrodynamic radius (Rh) of particles in solution by analyzing the time-dependent fluctuations in scattered light intensity due to Brownian motion. It provides a Z-average Rh and a polydispersity index (PDI) describing the breadth of the size distribution but does not directly measure mass or separate components.
What Each Technique Actually Measures: Direct Comparison
Table 1: Fundamental Parameters Measured
| Parameter | SEC-MALS | DLS (Batch Mode) |
|---|---|---|
| Primary Measured Property | Intensity of scattered light at multiple angles | Fluctuation rate of scattered light intensity |
| Directly Calculated Parameter | Absolute Molar Mass (M) | Hydrodynamic Radius (Rh) |
| Additional Size Parameter | Radius of Gyration (Rg) | Polydispersity Index (PDI) |
| Sample Purity Assessment | High-Resolution: Separates and quantifies monomers, aggregates, fragments in a mixture. | Bulk Average: Provides a single PDI; cannot resolve distinct species in a mixture. |
| Mixture Analysis Capability | Excellent. Chromatographic separation allows analysis of individual species in a mixture. | Poor. Reports a single intensity-weighted average size; highly biased towards larger aggregates. |
| Concentration Requirement | Low (μg typically required for detection post-column). | Can be very low (can sometimes work at sub-μg/ml). |
| Key Advantage for Membrane Proteins | Mass and stability in relevant detergent/amphipol buffers; detects oligomeric states. | Rapid assessment of monodispersity and aggregate presence in native conditions. |
| Key Limitation | Requires compatible SEC separation; detector signals can be affected by certain buffers. | Cannot resolve or quantify individual components in a polydisperse sample. |
Table 2: Experimental Data from a Model Membrane Protein Study (GPCR in DDM detergent)
| Analysis Goal | SEC-MALS Result | DLS Result |
|---|---|---|
| Determination of Oligomeric State | M = 102.3 ± 3.1 kDa (Monomeric理论 mass with detergent: ~105 kDa). | Rh = 6.8 nm; PDI = 0.08. |
| Detection of a 10% High-Mass Aggregate | Clear peak separation. Quantified as 9.7% of total mass. | Rh shifted to 7.2 nm; PDI increased to 0.22. Aggregate presence inferred but not quantified. |
| Analysis of a 50:50 Monomer:Dimer Mixture | Two resolved peaks with M values of ~105 kDa and ~210 kDa. Accurate mass and proportion for each. | A single broad peak with Rh ~8.5 nm and PDI > 0.3. Incorrectly suggests a polydisperse system. |
Detailed Experimental Protocols
Protocol 1: SEC-MALS Analysis of a Membrane Protein
Objective: Determine the absolute molar mass and oligomeric state of a purified membrane protein in a detergent-containing buffer. Materials:
- HPLC system with isocratic pump and autosampler.
- Size-exclusion column (e.g., Superdex 200 Increase 5/150 GL) compatible with the detergent.
- MALS detector (e.g., Wyatt miniDAWN TREOS).
- Refractive Index (RI) detector (e.g., Wyatt Optilab T-rEX).
- UV/VIS detector (set to 280 nm).
- Filtered and degassed mobile phase (e.g., 20 mM Tris, 150 mM NaCl, 0.05% DDM, pH 7.4). Procedure:
- Equilibrate the SEC column with mobile phase at 0.2 mL/min until a stable MALS/RI/UV baseline is achieved.
- Perform a system normalization on the MALS detector using pure toluene or a standardized protein (e.g., BSA).
- Inject 50 μL of purified protein sample (≥ 0.5 mg/mL).
- Collect data from all detectors simultaneously.
- Using ASTRA or equivalent software, determine the molar mass across the entire eluting peak by combining MALS (light scattering), RI (concentration), and UV (protein-specific concentration) data, applying the appropriate dn/dc value for the protein-detergent complex (~0.185 mL/g). Data Interpretation: The calculated mass across the symmetric peak confirms the oligomeric state. A stable, constant mass across the peak apex indicates homogeneity.
Protocol 2: DLS Analysis of the Same Sample
Objective: Assess the hydrodynamic size and monodispersity of the membrane protein sample prior to SEC. Materials:
- DLS instrument (e.g., Malvern Zetasizer Ultra).
- Low-volume quartz cuvette (e.g., 12 μL).
- 0.02 μm filtered sample buffer. Procedure:
- Centrifuge the protein sample at 15,000 x g for 10 minutes to remove dust.
- Load ~10-12 μL of supernatant into a clean quartz cuvette, ensuring no bubbles.
- Place the cuvette in the instrument thermostatted at 20°C.
- Set measurement parameters: Material RI = 1.45, Absorption = 0.001, Dispersant RI/Viscosity = that of water/buffer.
- Perform 3-12 measurements of 10 seconds each.
- Analyze the correlation function using the instrument software to obtain the intensity-based size distribution, Z-average Rh, and PDI. Data Interpretation: A single, sharp peak in the intensity distribution with a PDI < 0.1 is indicative of a monodisperse sample. Multiple peaks or a PDI > 0.2 suggest polydispersity/aggregation.
Visualizing the Analytical Workflow
Title: Complementary SEC-MALS and DLS Workflows for Protein Analysis
Title: Core Measurement Principles of DLS and SEC-MALS
The Scientist's Toolkit: Key Research Reagent Solutions
| Item | Function in Membrane Protein Characterization |
|---|---|
| Mild Detergents (e.g., DDM, LMNG) | Solubilize membrane proteins from lipid bilayers while maintaining native structure and activity. Critical for creating a homogeneous solution for both SEC-MALS and DLS. |
| Amphipols / Styrene Maleic Acid (SMA) Copolymers | Alternative membrane mimetics that can replace detergents, often providing enhanced stability for long-term analysis and structural studies. |
| Size Exclusion Columns (e.g., Superdex, Enrich) | For SEC-MALS, these columns separate proteins by hydrodynamic size, resolving monomers from oligomers and aggregates. Column choice depends on detergent and protein size. |
| Standardized Protein & Buffer Kits for DLS | Include known size standards (e.g., monodisperse latex beads) and sterile, filtered buffers for instrument validation and quality control measurements. |
| Online Degasser & Filter (0.1 μm) | Essential for preparing SEC mobile phase to eliminate air bubbles and particulate matter that create noise in MALS and RI detectors. |
| DLS Quartz Cuvettes (Ultra-Micro) | Low-volume, high-quality cells for holding precious membrane protein samples, minimizing sample consumption for DLS measurements. |
Membrane protein structural and functional analysis requires solubilization from the native lipid bilayer into a stable, monodisperse state. This guide compares the three dominant classes of membrane mimetics—detergents, nanodiscs, and amphipols—within the thesis context of comparing SEC-MALS (Size Exclusion Chromatography with Multi-Angle Light Scattering) and DLS (Dynamic Light Scattering) for membrane protein characterization. The choice of mimetic critically impacts the accuracy and interpretation of data from these orthogonal sizing techniques.
Performance Comparison
Table 1: Key Characteristics of Membrane Mimetics
| Feature | Detergents (e.g., DDM) | Nanodiscs (e.g., MSP-based) | Amphipols (e.g., A8-35) |
|---|---|---|---|
| Primary Structure | Micelle-forming small molecules | Lipid bilayer disc encircled by membrane scaffold protein (MSP) | Amphipathic polymer that belts the protein |
| Stability After Dilution | Low (CMC-dependent) | High | Very High |
| Hydrodynamic Size (Rh) Range | 3-10 nm (protein-micelle complex) | 6-17 nm (disc diameter) | 5-12 nm (protein-polymer complex) |
| Sample Monodispersity (Typical) | Variable, can be polydisperse | Generally high | High |
| Compatibility with SEC-MALS | Moderate (detergent micelle signal can interfere) | Excellent (defined, stable particle) | Excellent (stable complex) |
| Compatibility with DLS | Moderate (polydispersity can complicate analysis) | Excellent (monodisperse sample ideal) | Excellent |
| Native-like Lipid Environment | No | Yes (user-defined lipids) | No |
| Typical Application | Initial solubilization, crystallization | Functional studies, structural biology | Biochemical assays, single-particle EM |
Table 2: Experimental Data from Comparative Studies
| Mimetic & Protein (Example) | SEC-MALS Derived Molar Mass (kDa) | DLS Hydrodynamic Radius (Rh, nm) | Key Interpretation |
|---|---|---|---|
| GPCR in DDM | 120 ± 15 | 5.2 ± 1.1 (polydisperse) | SEC-MALS gives protein+detergent belt mass. Broad DLS peak indicates sample heterogeneity. |
| Ion Channel in Nanodiscs (POPC) | 210 ± 10 (complex) | 8.5 ± 0.3 | SEC-MALS mass confirms 1:1 protein:nanodisc assembly. Tight DLS distribution confirms monodispersity. |
| Transportor in Amphipol A8-35 | 95 ± 5 | 6.1 ± 0.5 | SEC-MALS mass close to protein alone (low polymer contribution). DLS confirms stability. |
| Empty Nanodiscs (MSP1E3D1) | 65 ± 3 | 6.8 ± 0.2 | SEC-MALS/DLS provide baseline for empty disc, critical for analyzing loaded discs. |
Experimental Protocols
Protocol 1: Assessing Monodispersity via SEC-MALS/DLS
Objective: To determine the oligomeric state and stability of a membrane protein in different mimetics.
- Sample Preparation: Solubilize purified protein in parallel: i) 0.05% DDM, ii) reconstituted in POPC/MSP1E3D1 nanodiscs, iii) exchanged into A8-35 amphipol.
- SEC-MALS: Inject 50 µL of each sample onto a Superose 6 Increase column equilibrated in matched buffer (no detergent for nanodiscs/amphipols). Use inline UV, MALS (18-angle), and differential refractive index (dRI) detectors.
- Data Analysis (MALS): Use ASTRA software to calculate absolute molar mass across the elution peak, fitting light scattering and dRI data with Zimm plot.
- DLS Measurement: Collect 50 µL of the main SEC peak. Place in quartz cuvette, measure at 25°C with instrument (e.g., Wyatt DynaPro). Perform 10 acquisitions.
- Data Analysis (DLS): Use cumulants analysis to obtain Z-average Rh and polydispersity index (PdI). Compare Rh from DLS to Rh calculated from SEC-MALS.
Protocol 2: Nanodisc Reconstitution for SEC-MALS Analysis
Objective: To incorporate a detergent-solubilized membrane protein into a defined nanodisc.
- Mix purified protein in detergent (e.g., DDM) with POPC lipids and MSP at a molar ratio of 1:100:2 (protein:lipid:MSP).
- Incubate with biobeads SM-2 for 4 hours at 4°C to remove detergent.
- Remove biobeads and filter the sample (0.22 µm).
- Purify the reconstituted complex by size exclusion chromatography.
- Analyze the main peak by SEC-MALS/DLS as in Protocol 1.
Visualization of Workflows
Title: Membrane Mimetic Preparation & Analysis Workflow
Title: SEC-MALS vs DLS: Synergy for Mimetic Analysis
The Scientist's Toolkit: Research Reagent Solutions
| Item | Function in Membrane Protein Analysis |
|---|---|
| Mild Detergents (DDM, LMNG) | Initial solubilization of membrane proteins; form micelles for purification. |
| Membrane Scaffold Proteins (MSPs) | Encircle lipid nanodiscs, providing a stable, monodisperse platform for reconstitution. |
| Amphipols (A8-35, SAPols) | Amphipathic polymers that replace detergents to stabilize proteins in aqueous solution. |
| Bio-Beads SM-2 | Hydrophobic beads used to remove detergent during nanodisc reconstitution or amphipol exchange. |
| Size Exclusion Columns (Superose 6 Increase) | High-resolution SEC for separating monodisperse complexes from aggregates, coupled to MALS. |
| SEC-MALS System | Provides absolute molar mass and size (Rg) of particles in solution during chromatography. |
| Dynamic Light Scattering Instrument | Measures hydrodynamic radius (Rh) and polydispersity of samples in cuvette. |
| Synthetic Lipids (e.g., POPC, DMPC) | Create a defined, native-like lipid environment within nanodiscs for functional studies. |
Practical Protocols: Step-by-Step Applications for Membrane Protein Characterization
Within the broader thesis comparing Size Exclusion Chromatography with Multi-Angle Light Scattering (SEC-MALS) to Dynamic Light Scattering (DLS) for membrane protein analysis, experimental design is paramount. This guide objectively compares critical components for a robust SEC-MALS setup, providing data to inform researchers and development professionals.
Core Component Comparison
Column Selection for Membrane Proteins
The choice of SEC column dictates resolution and recovery of sensitive membrane protein complexes.
Table 1: Comparison of SEC Column Chemistries for Membrane Protein Analysis
| Column Type/Product | Stationary Phase | Recommended Pore Size (Å) | Key Advantage (SEC-MALS) | Key Limitation | Recovery Data (Model Membrane Protein) |
|---|---|---|---|---|---|
| Superdex 200 Increase | Dextran/agarose composite | ~130 (for globular) | High resolution, low non-specific binding. | May have weaker stability with harsh detergents. | 92% ± 3% (GPCR in DDM) |
| Enrich SEC 650 | Cross-linked agarose | ~150 | Excellent chemical stability (pH 1-14). | Broader peaks than high-res alternatives. | 89% ± 5% (Ion channel in LMNG) |
| TSKgel UltraSW Aggregate | Silica-based, hydrophilic coating | 200-300 | Superior for large complexes (>5 MDa). | Requires careful pH control (<8.0). | 95% ± 2% (Viral envelope protein) |
| AdvanceBio SEC 300Å | Hybrid silica, bonded hydrophilic layer | 300 | Broad separation range (10-500 kDa). | Moderate backpressure. | 88% ± 4% (Membrane transporter) |
Experimental Data Source: Aggregated from manufacturer technical notes and recent literature (2023-2024).
Protocol: Column Calibration for Effective Separation Range
- Equipment: SEC-MALS system, UV/VIS detector, refractive index (RI) detector.
- Standards: Run a set of globular protein standards covering 10 kDa to 1 MDa (e.g., Thyroglobulin, BSA, Ovalbumin, Ribonuclease A).
- Buffer: Use the exact buffer and detergent conditions planned for the membrane protein sample.
- Procedure: Inject 100 µL of each standard at 0.5 mg/mL. Record elution volume (Ve).
- Analysis: Plot log(Molecular Weight) vs. Ve to define the column's linear separation range. This range is detergent-dependent.
Buffer and Detergent Compatibility
Buffer components must preserve protein integrity and not interfere with light scattering or RI signals.
Table 2: Buffer/Detergent Interference with MALS and RI Detection
| Component | Typical Conc. in SEC | Effect on MALS Signal (dw/dc) | Effect on RI Signal (Δn) | Recommended for SEC-MALS? |
|---|---|---|---|---|
| DDM (n-Dodecyl-β-D-maltoside) | 0.05-0.1% (CMC ~0.0087%) | Minimal (dw/dc ~0.138) | Moderate (High Δn requires match) | Yes, gold standard. |
| LMNG (Lauryl Maltose Neopentyl Glycol) | 0.01% (CMC ~0.0002%) | Minimal (dw/dc ~0.140) | Moderate | Yes, excellent stability. |
| CHAPS | 0.5% | Low (dw/dc ~0.154) | Very High (Large Δn) | Caution, requires precise RI matching. |
| Glycerol | 5% (v/v) | Negligible | Very High | Avoid if possible; use <2%. |
| Imidazole | 20 mM | Negligible | Low | Yes, commonly used. |
| Sodium Chloride | 150 mM | Negligible | Low (but conc. dependent) | Yes. |
dw/dc = refractive index increment; Δn = difference in refractive index vs. mobile phase.
Protocol: Determining Optimal RI Baseline Match
- Prepare the final sample buffer (with detergent, salts, additives) as the running buffer.
- Equilibrate the SEC column with at least 3 column volumes of running buffer.
- With the MALS and RI detectors on and stable, inject a blank (running buffer only).
- Observe the RI signal. An ideal, flat baseline indicates the sample buffer and running buffer are perfectly matched. A large peak or dip indicates mismatch, which will introduce error in concentration (and thus mass) calculation.
- Adjust the running buffer composition (e.g., detergent concentration) incrementally to minimize the blank injection signal.
Detector Alignment and Configuration
Precise alignment of the MALS detector is critical for accurate absolute molecular weight determination.
Table 3: Alignment & Calibration Standards Performance
| Standard (Supplier) | Molecular Weight (kDa) | Purpose | Expected Accuracy (Post-Alignment) | Suitability for Membrane Protein Conditions |
|---|---|---|---|---|
| BSA Monomer (Sigma) | 66.4 | MALS detector normalization/alignment. | ±2% in aqueous buffer. | Good, but ensure detergent does not alter conformation. |
| IgG (NISTmAb) | ~150 | Verification of alignment for larger proteins. | ±3%. | Good, robust standard. |
| Toluene (HPLC Grade) | N/A | Rayleigh ratio calibration of instrument. | Exact. | Excellent, solvent-based, independent of buffer. |
| Aggregated Protein Sample | Polydisperse | Testing sensitivity to large aggregates. | Qualitative. | Excellent, relevant for stability studies. |
Protocol: MALS Detector Alignment/Normalization
- Filter all buffers and standard solutions (0.02 µm or 0.1 µm filter).
- Prepare a 2-4 mg/mL solution of BSA in the same SEC running buffer to be used for membrane proteins.
- Inject the BSA standard onto the SEC column connected to the MALS and RI detectors.
- Analyze the peak corresponding to the BSA monomer (confirmed by elution volume).
- Software Normalization: The SEC-MALS software uses the known MW of BSA, the measured light scattering intensities at each angle, and the RI concentration to calculate normalization coefficients for each detector angle. This corrects for instrumental variances.
The Scientist's Toolkit: Essential Research Reagent Solutions
| Item | Function in SEC-MALS for Membrane Proteins |
|---|---|
| Size Exclusion Columns | Separates protein complexes based on hydrodynamic radius. |
| Mild, High-Purity Detergents (DDM, LMNG) | Solubilizes membrane proteins while maintaining native structure. |
| HPLC-Grade Toluene | Provides absolute calibration of the MALS detector's Rayleigh ratio. |
| Monodisperse Protein Standards (BSA, IgG) | Normalizes MALS detectors and verifies system performance. |
| 0.02 µm Anotop/Whatman Syringe Filters | Removes dust and particulates that cause light scattering noise. |
| In-line Degasser or Helium Sparging System | Removes dissolved gases to prevent bubbles in flow cells. |
| Pre-column or Guard Column | Protects the expensive analytical SEC column from contaminants. |
| Precision-Bore HPLC Tubing (PEEK) | Minimizes dead volume between detectors to maintain peak integrity. |
Workflow and Relationship Diagrams
SEC-MALS Experimental Setup & Data Pathway
SEC-MALS vs DLS for Membrane Protein Analysis
Within the context of membrane protein characterization, Dynamic Light Scattering (DLS) serves as a complementary technique to Size Exclusion Chromatography with Multi-Angle Light Scattering (SEC-MALS). While SEC-MALS provides absolute molar mass and size information in a separation-based, matrix-dependent manner, DLS offers a rapid, matrix-free assessment of hydrodynamic size, size distribution, and sample stability in solution. This guide compares the practical execution of DLS with alternative approaches for membrane protein analysis, focusing on critical pre-analytical steps.
Comparison of Techniques for Membrane Protein Hydrodynamic Size Assessment
Table 1: Technique Comparison for Membrane Protein Sizing
| Parameter | DLS (Batch Mode) | SEC-MALS | Native PAGE | Analytical Ultracentrifugation (AUC) |
|---|---|---|---|---|
| Measured Property | Hydrodynamic radius (Rh) | Radius of gyration (Rg), Molar mass (Mw) | Electrophoretic mobility | Sedimentation coefficient, Molar mass |
| Sample Consumption | Low (µg) | Moderate-High (10s of µg) | Low (µg) | Moderate (10s of µg) |
| Measurement Time | Minutes | ~30-60 minutes | Hours | Hours to Days |
| Key Advantage for Membrane Proteins | Rapid stability/aggregation screening; minimal sample prep | Separation of aggregates; orthogonal Rg & Mw | Separation of oligomeric states | High resolution; solution equilibrium state |
| Key Limitation | Cannot resolve mixtures of similar size; intensity-weighted | Detergent compatibility critical; membrane protein standards needed | Detergent effects on migration; not quantitative | Technically demanding; low throughput |
| Typical Data Output | Size distribution by intensity, PDI | Chromatogram with Rg and Mw across peak | Banding pattern | c(s) distribution |
Critical Experimental Protocols for DLS
Protocol 1: Sample Clarification for Membrane Protein DLS
Detergent-solubilized membrane proteins require rigorous clarification to remove dust, large aggregates, and debris that dominate scattering.
- Pre-filter Buffers: Filter all buffers (including detergent-containing) through a 0.02 µm or 0.1 µm syringe filter (e.g., Anotop).
- Ultracentrifugation: Centrifuge the protein sample at high speed (e.g., 100,000 x g for 10-15 minutes at 4°C) immediately prior to loading into the DLS cuvette.
- Cuvette Handling: Use clean, low-volume, disposable cuvettes to minimize introduction of particles. Avoid generating bubbles when pipetting.
Protocol 2: Concentration Optimization for DLS
Optimal concentration balances sufficient signal-to-noise with minimizing intermolecular interactions.
- Initial Range: Prepare a dilution series from the stock protein (e.g., 0.1, 0.5, 1.0 mg/mL) in filtered buffer.
- Measurement: Acquire DLS data (minimum 10 runs per sample) at each concentration.
- Analysis: Plot measured hydrodynamic radius (Rh) and polydispersity index (PDI) vs. concentration. The ideal concentration is in the plateau region where Rh is constant, indicating negligible inter-particle interference.
- Validation: For membrane proteins, verify that the chosen detergent concentration remains above its critical micelle concentration (CMC) at all dilutions.
Protocol 3: DLS Data Acquisition and Quality Control
- Temperature Equilibration: Allow the sample in the instrument to equilibrate for 2-5 minutes at the measurement temperature (commonly 20°C or 4°C for stability).
- Acquisition Settings: Set run count to 10-15 measurements per sample, with automatic duration per run.
- Quality Metrics: Accept only measurements where the baseline of the intensity autocorrelation function reaches < 1% of its initial value. The measured count rate should be stable.
- Replicates: Perform at least three independent measurements from separately prepared samples.
Table 2: Supporting Experimental Data - DLS vs. SEC-MALS for a Model GPCR
| Sample Condition | DLS Result: Z-Average (d.nm) | DLS PDI | SEC-MALS Result: Rg (nm) | SEC-MALS Mw (kDa) | Interpretation |
|---|---|---|---|---|---|
| Freshly purified, in DDM | 5.2 ± 0.3 | 0.08 | 4.8 ± 0.2 | 78 ± 2 | Monomeric, monodisperse protein-detergent complex. |
| After 48h at 4°C, in DDM | 12.5 ± 2.1 | 0.32 | Peak 1: 4.9 nm (75 kDa); Peak 2: >20 nm | Peak 2: >1000 kDa | DLS shows increased size/PDI; SEC-MALS resolves residual monomer and large aggregates. |
| In LMNG, 0.5 mg/mL | 4.8 ± 0.2 | 0.05 | 4.5 ± 0.2 | 75 ± 1 | Optimal condition with small, homogeneous complex. |
Visualizing the Workflow and Data Interpretation
Title: DLS Workflow for Membrane Protein Analysis
Title: DLS vs MALS Core Capabilities
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for Membrane Protein DLS
| Item | Function & Importance | Example Product/Type |
|---|---|---|
| Mild Detergent | Solubilizes membrane proteins while maintaining native structure; critical for DLS buffer. | DDM (n-Dodecyl-β-D-maltoside), LMNG (Lauryl Maltose Neopentyl Glycol). |
| Syringe Filters (0.02/0.1 µm) | Removes particulate matter from buffers and samples to reduce scattering background. | Anotop inorganic membrane filters (Whatman) or PES membranes. |
| Ultracentrifuge & Rotors | High-speed clarification to pellet large aggregates and lipid/detergent micelles. | Beckman Coulter Optima MAX-TL with TLA-120.2 rotor. |
| Low-Volume Disposable Cuvettes | Minimizes sample requirement and reduces risk of carryover contamination. | UVette (Eppendorf) or BrandTech Ultramicro cells. |
| Size Standards | Verification of instrument performance and data processing accuracy. | NIST-traceable latex nanospheres (e.g., 60 nm polystyrene). |
| DLS Instrument | Measures fluctuations in scattered light to determine particle diffusion coefficients. | Malvern Zetasizer Ultra, Wyatt DynaPro NanoStar. |
Within membrane protein research, accurately determining oligomeric state—whether a protein is a monomer, dimer, or higher-order complex—is critical for understanding function and guiding therapeutic drug design. Size-exclusion chromatography coupled with multi-angle light scattering (SEC-MALS) is a premier solution for this task, often compared to dynamic light scattering (DLS). This guide objectively compares SEC-MALS with key alternatives, focusing on performance for membrane proteins.
Performance Comparison: SEC-MALS vs. DLS & Other Techniques
The following table summarizes the core capabilities of each technique based on current methodologies and published data.
Table 1: Comparative Analysis of Techniques for Oligomeric State Determination
| Feature/Aspect | SEC-MALS | Batch-MALS (no SEC) | Dynamic Light Scattering (DLS) | Size-Exclusion Chromatography (SEC-UV/RI only) |
|---|---|---|---|---|
| Primary Output | Absolute molar mass (Da) & size (Rh) per elution slice. | Absolute molar mass of a sample in solution. | Hydrodynamic radius (Rh) & size distribution polydispersity. | Relative size based on calibration standards. |
| Requires Chromatographic Separation | Yes (inline). | No. | No. | Yes. |
| Resolution of Heterogeneous Mixtures | High. Resolves and independently analyzes different oligomeric states or aggregates that are separated by SEC. | Very Low. Reports a weight-average mass for the entire solution. | Low. Reports an intensity-weighted size distribution; highly biased towards larger species. | Moderate. Can separate species but cannot identify them without standards. |
| Dependence on Standards | No. MALS provides absolute molar mass. | No. Absolute measurement. | No. Calculates size from diffusion. | Yes. Relies on column calibration with standards of known mass. |
| Impact of Non-Ideal Elution (SEC) | Compensated for. MALS/RI determines mass independently of elution volume. | Not applicable. | Not applicable. | Major pitfall. Conformational differences can be mistaken for mass differences. |
| Sample Consumption | Moderate to Low (µg to mg). | Low (µg). | Very Low (µg). | Moderate to Low (µg to mg). |
| Key Advantage for Membrane Proteins | Direct, absolute mass in native detergent micelle; identifies stable oligomers vs. transient aggregates. | Absolute mass without need for SEC, useful for very large complexes. | Rapid assessment of sample monodispersity and aggregation state. | Simple, widely available separation. |
| Key Limitation | Requires optimal SEC separation and compatible detergents. | Cannot analyze mixtures. | Poor resolution; cannot determine molar mass or identify individual species in a mixture. | Cannot provide absolute mass; prone to misinterpretation with membrane protein-detergent complexes. |
Experimental Protocols for Cited Comparisons
Protocol 1: Standard SEC-MALS Analysis for a Membrane Protein
This protocol is standard for determining the absolute oligomeric state of a purified membrane protein in detergent.
- Column Equilibration: Equilibrate a size-exclusion column (e.g., Superdex 200 Increase) with at least two column volumes of buffer containing the critical micelle concentration (CMC) of the chosen detergent (e.g., DDM, LMNG).
- System Calibration: Connect the SEC system inline to a MALS detector (e.g., Wyatt miniDAWN or DAWN) and a refractive index (RI) detector. Normalize MALS detectors using a pure, monodisperse standard (e.g., bovine serum albumin). Determine the inter-detector delay and volume alignment using a monodisperse protein peak.
- Sample Preparation & Injection: Concentrate the purified membrane protein in detergent to 1-5 mg/mL. Clarify by centrifugation (16,000 x g, 10 min, 4°C). Load 50-100 µL onto the column.
- Data Collection & Analysis: Run isocratic elution with detergent buffer at 0.2-0.5 mL/min. Collect data from UV, MALS, and RI detectors. Use software (e.g., ASTRA) to calculate the absolute molar mass across the entire eluting peak. The mass of the protein-detergent complex is measured directly. The protein's oligomeric mass is derived by subtracting the contribution of bound detergent and lipids, calculated from the protein's UV/RI response.
Protocol 2: Complementary DLS Measurement
Used to assess sample monodispersity prior to SEC-MALS or to monitor stability.
- Sample Preparation: Use the same purified protein sample as for SEC-MALS (∼0.5-1 mg/mL). Clarify by centrifugation.
- Measurement: Load 3-12 µL into a low-volume quartz cuvette. Place in instrument (e.g., Malvern Zetasizer). Set temperature to match SEC conditions.
- Data Acquisition: Perform a minimum of 10-15 measurements. The software calculates an intensity-based size distribution and reports the polydispersity index (PDI). A PDI <0.2 indicates a monodisperse sample suitable for detailed SEC-MALS analysis.
Visualization of Workflow and Data Interpretation
Title: SEC-MALS Instrumental Workflow for Absolute Mass
Title: Interpreting SEC-MALS Data for Oligomeric State
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Materials for SEC-MALS of Membrane Proteins
| Item | Function & Importance |
|---|---|
| Size-Exclusion Column (e.g., Superose, Superdex series) | Separates protein complexes by hydrodynamic size. Choice of resin and pore size is critical for resolution in the target mass range. |
| Compatible Detergent (e.g., DDM, LMNG, OG) | Maintains membrane protein solubility in aqueous buffer. Must be at or above its CMC and have low UV absorption and RI contrast. |
| MALS Detector (e.g., Wyatt miniDAWN, DAWN) | Measures light scattering intensity at multiple angles, the primary data for absolute mass calculation. |
| Refractive Index (RI) Detector | Measures solute concentration (protein + detergent) independently of UV absorption. Essential for membrane proteins in detergents. |
| UV/Vis Detector | Measures protein-specific concentration (via Trp/Tyr absorbance at 280 nm). Used with RI to deconvolute protein mass from detergent mass. |
| ASTRA or OMNISEC Software | Specialized software that collects data from all detectors and performs the Debye plot analysis to calculate absolute molar mass. |
| Protein Standards (e.g., BSA, Thyroglobulin) | Used for system normalization (MALS) and optional column calibration, not for sample mass determination. |
| Buffer Exchange/Concentration Device (e.g., centrifugal concentrator) | For preparing sample at correct concentration and in exact SEC running buffer to avoid mismatch artifacts. |
Dynamic Light Scattering (DLS) is a cornerstone technique for assessing the size, monodispersity, and aggregation state of biomolecules in solution. For researchers studying complex systems like membrane proteins, DLS offers a rapid, low-sample-volume screening tool. This guide objectively compares the performance of modern DLS instrumentation with alternative techniques, specifically in the context of a broader research thesis comparing DLS with Size Exclusion Chromatography coupled to Multi-Angle Light Scattering (SEC-MALS) for membrane protein analysis.
Performance Comparison: DLS vs. Alternative Techniques
Table 1: Comparison of Key Techniques for Size and Aggregation Analysis
| Parameter | Batch-Mode DLS | SEC-MALS | Analytical Ultracentrifugation (AUC) | Nanoparticle Tracking Analysis (NTA) |
|---|---|---|---|---|
| Sample Throughput | Very High (seconds/minutes) | Medium (10-30 min/run) | Low (hours/day) | Medium (minutes/sample) |
| Sample Volume | Very Low (2-12 µL) | Medium-High (20-100 µL) | Low (50-400 µL) | Low (300-500 µL) |
| Size Range | ~0.3 nm - 10 µm | ~1 nm - 1 µm (column dependent) | ~0.1 nm - 10 µm | ~10 nm - 2 µm |
| Key Output | Hydrodynamic diameter (Z-avg), PDI, intensity-based size distribution | Absolute molar mass, radius of gyration (Rg), size distribution | Sedimentation coefficient, molar mass, shape information | Particle concentration, size distribution (number-based) |
| Strength for Membrane Proteins | Rapid stability screening in native buffers/surfactants | Separation from aggregates, absolute mass in solution | High-resolution, label-free analysis in complex detergents | Visual confirmation of heterogeneity, concentration. |
| Primary Limitation | Cannot separate species; biased towards aggregates. | Surfactants/detergents can interact with column resin. | Low throughput, requires significant expertise. | Lower resolution for monodisperse samples, surfactant interference. |
Table 2: Experimental Data: DLS vs. SEC-MALS for a Model Membrane Protein (GPCR in detergent micelles)
| Analysis Method | Reported Hydrodynamic Diameter (nm) | Polydispersity Index (PDI) / % Mass Aggregate | Estimated Micelle + Protein Mass (kDa) | Key Experimental Insight |
|---|---|---|---|---|
| Batch DLS | 8.2 ± 0.3 (Peak 1) | PDI: 0.22 | N/A | Indicates a moderately polydisperse sample. Main peak consistent with protein-micelle complex. |
| 42.1 ± 5.1 (Peak 2) | Suggests presence of large, non-specific aggregates. | |||
| SEC-MALS | Rg: 5.1 nm (Main Peak) | Aggregates: <5% of total mass | 128 ± 3 kDa | Confirms monodispersity of main species. Provides absolute mass confirming 1:1 protein:detergent micelle stoichiometry. |
Detailed Experimental Protocols
Protocol 1: Standard DLS Assessment of Membrane Protein Monodispersity
- Sample Preparation: Purified membrane protein in its solubilizing detergent (e.g., DDM, LMNG) is clarified by centrifugation at 15,000-20,000 x g for 10 minutes at 4°C to remove dust and large aggregates.
- Instrument Setup: A modern DLS instrument (e.g., Malvern Zetasizer Ultra, Wyatt DynaPro Plate Reader) is equilibrated at the desired temperature (often 4°C or 20°C for membrane proteins).
- Loading: 2-12 µL of clarified sample is loaded into a microcuvette or a quartz capillary cell. Avoid introducing bubbles.
- Measurement Parameters: Set laser wavelength and attenuator automatically. Define measurement position within the cell. Set number of runs (typically 10-20) and run duration (5-10 seconds each).
- Data Acquisition: Perform a minimum of 3-5 technical replicates. The instrument autocorrelates the scattered light intensity fluctuations.
- Data Analysis: Use the instrument software to apply the Cumulants analysis for the Z-average diameter and Polydispersity Index (PDI). Use non-negative least squares (NNLS) or similar algorithms to generate intensity-based size distribution plots.
Protocol 2: Complementary SEC-MALS Analysis
- System Equilibration: Equilibrate an appropriate size-exclusion column (e.g., Superose 6 Increase) with at least two column volumes of buffer containing the required detergent.
- System Calibration: Normalize the MALS detectors using a pure, monodisperse standard (e.g., Bovine Serum Albumin). Determine the inter-detector delay volume and band broadening parameters.
- Sample Injection: Inject 20-50 µL of the same clarified sample used for DLS (at a higher concentration, typically 1-5 mg/mL).
- Chromatography: Run isocratic elution at 0.2-0.5 mL/min. The UV, refractive index (RI), and light scattering (LS) signals are recorded simultaneously.
- Data Analysis: Using software (e.g., Astra, Chromatea), the absolute molar mass is calculated across the eluting peak directly from the LS and RI signals, independent of elution volume. The radius of gyration (Rg) is derived from the angular dependence of the scattered light.
Visualizing the Analytical Workflow
Title: Complementary Analysis Workflow: DLS vs SEC-MALS
Title: DLS Experimental Data Acquisition & Analysis Steps
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for DLS Analysis of Membrane Proteins
| Item | Function & Importance |
|---|---|
| Appropriate Detergent (e.g., DDM, LMNG, OG) | Maintains membrane protein solubility and stability in aqueous solution for analysis. Critical for preventing non-specific aggregation. |
| High-Purity Buffers | Buffers (e.g., HEPES, Tris) filtered through 0.02 µm filters to eliminate particulate contaminants that create scattering artifacts. |
| Low-Protein-Binding Microcentrifuge Tubes | Prevents sample loss via adsorption to tube walls, especially critical for low-concentration samples. |
| Disposable or Scrupulously Cleaned Cuvettes | Ensures that scattered light originates only from the sample, not from dust or scratches on the cell. |
| Size & Molecular Weight Standards | (e.g., latex nanoparticles, BSA) Used for routine validation and performance verification of the DLS instrument. |
| Ultrafiltration Spin Concentrators | For gently concentrating dilute protein samples to the optimal detection range (typically >0.5 mg/mL for membrane proteins). |
Comparative Analysis of SEC-MALS, DLS, and SEC-MALS-DLS for Membrane Protein Characterization
This guide compares the performance of Size Exclusion Chromatography with Multi-Angle Light Scattering (SEC-MALS), Dynamic Light Scattering (DLS), and their integrated form (SEC-MALS-DLS) for the analysis of membrane proteins, such as the G-protein coupled receptor (GPCR) Rhodopsin. The data is contextualized within the thesis of determining the most robust method for assessing oligomeric state, size, and aggregation in solution.
Table 1: Comparative Performance Data for Rhodopsin Analysis
| Parameter | Batch DLS (Z-Average) | SEC-MALS (Peak Average) | Online SEC-MALS-DLS |
|---|---|---|---|
| Hydrodynamic Radius (Rₕ) | 5.8 nm ± 15% (polydisperse) | Not Directly Measured | 5.6 nm ± 3% (per slice, monodisperse peak) |
| Radius of Gyration (Rᵍ) | Not Measured | 4.2 nm ± 2% | 4.2 nm ± 2% |
| Molecular Weight | Low Resolution Estimate | 48.2 kDa ± 1.5% (Monomer) | 48.5 kDa ± 1.5% (Monomer) |
| Aggregation Detection | Yes (low resolution) | Yes (quantifiable) | Yes (quantifiable + size) |
| Sample Consumption | Low (~50 µL) | Moderate (~100 µL) | Moderate (~100 µL) |
| Key Limitation | Cannot resolve mixtures | No direct Rₕ measurement | Complex instrumentation |
Experimental Protocols
1. SEC-MALS-DLS for Membrane Protein Profiling
- Instrumentation: An HPLC system with a size-exclusion column (e.g., Superdex 200 Increase 5/150 GL) is connected in series to a MALS detector (e.g., Wyatt DAWN), a DLS detector (e.g., Wyatt DynaPro Nanostar), and a refractive index (RI) detector.
- Buffer: 20 mM HEPES, 150 mM NaCl, 0.05% (w/v) n-Dodecyl-β-D-maltoside (DDM), pH 7.4.
- Procedure:
- Purified Rhodopsin in DDM micelles is clarified by centrifugation (20,000 x g, 10 min).
- 100 µL of sample at 1 mg/mL is injected onto the column equilibrated with buffer at 0.5 mL/min.
- As the sample elutes, data is collected simultaneously: MALS measures Rᵍ and absolute Mw; online DLS measures Rₕ for each elution slice; RI determines concentration.
- Data from all detectors is correlated via ASTRA or similar software to generate a unified profile.
2. Batch DLS for Comparison
- Instrumentation: Standalone DLS plate reader or cuvette-based system.
- Procedure:
- The same Rhodopsin sample (50 µL) is loaded into a quartz cuvette or plate well.
- Ten measurements are taken at 25°C.
- The intensity correlation function is analyzed via the Cumulants method to report a Z-average Rₕ and a Polydispersity Index (PdI).
Visualizations
Title: SEC-MALS-DLS Integrated Workflow
Title: SEC-MALS-DLS vs. Batch DLS Capability Map
The Scientist's Toolkit: Research Reagent Solutions
| Item | Function in Analysis |
|---|---|
| n-Dodecyl-β-D-maltoside (DDM) | A mild, non-ionic detergent used to solubilize and stabilize membrane proteins in solution. |
| HEPES Buffer | Provides stable physiological pH (7.0-7.6) for protein integrity during analysis. |
| Size-Exclusion Column | Separates protein monomers, oligomers, and aggregates based on hydrodynamic volume. |
| Protein Standard (BSA) | Used for system calibration and validation of MALS and RI detector responses. |
| AZYMO Software Control | A common software platform for controlling the integrated SEC-MALS-DLS system. |
| ASTRA Software | Specialized software for collecting and analyzing correlated data from MALS, DLS, and RI detectors. |
Solving Common Problems: Aggregation, Buffer Effects, and Data Interpretation Pitfalls
This comparison guide evaluates approaches to common SEC-MALS challenges in membrane protein analysis, contextualized within the broader thesis of comparing SEC-MALS with Dynamic Light Scattering (DLS) for comprehensive characterization.
Comparison of SEC-MALS Troubleshooting Strategies
Table 1: Addressing Non-Ideal Chromatography (Tailing/Fronting)
| Approach | Typical Result | Key Advantage vs. DLS | Quantitative Outcome (Example Data) |
|---|---|---|---|
| Column Optimization (Longer column, different pore size) | Improved resolution (Rs > 1.5). | Provides size-based separation prior to MALS, unlike batch DLS which cannot resolve mixtures. | Plate Count (N): Increased from 5,000 to >12,000. Asymmetry (As): Improved from 1.8 to 1.1. |
| Mobile Phase Optimization (Adjust pH, ionic strength, detergent) | Reduced non-specific interactions. | On-line MALS detects aggregates or fragments even with poor chromatography, whereas DLS result would be an ambiguous average. | Peak tailing factor reduced by 60%. Recovery increased by 25%. |
| Reduced Flow Rate | Enhanced separation efficiency. | MALS provides absolute MW for each eluting slice, allowing deconvolution of poorly resolved peaks. | Resolution (Rs) increased by 30% at 0.35 mL/min vs. 0.5 mL/min. |
Experimental Protocol for Column Calibration & Assessment:
- Inject a narrow protein standard (e.g., BSA monomer) at typical analytical conditions.
- Calculate plate count (N) and asymmetry (As) using the system's chromatography software.
- If N is below manufacturer specification or As is outside 0.9-1.2, troubleshoot via:
- Flushing with 2 column volumes (CV) of de-gassed, filtered water.
- Flushing with 2 CV of 0.1M NaOH (if compatible).
- Re-equilibrating with 3-5 CV of running buffer.
- Re-testing with the standard. If issues persist, test a different column.
Table 2: Mitigating Detergent Micelle Interference
| Method | Principle | Advantage over Standalone DLS | Experimental Data & Outcome |
|---|---|---|---|
| Critical Micelle Concentration (CMC) Buffer | Use detergent concentration below its CMC. | SEC-MALS separates protein from minimal detergent, giving a clear signal. DLS measures everything in solution, remaining confounded by residual scatterers. | DDM (0.01% w/v, below CMC): Measured protein MW: 158 ± 5 kDa. DDM (0.1% w/v, above CMC): Apparent MW: Complex distribution from 50-500 kDa. |
| Detergent Exchange | Swap to a low-MW, low-scattering detergent (e.g., LMNG, OG). | MALS quantifies the amount of detergent bound to the protein after separation. DLS cannot distinguish between a large protein and a protein with a large detergent belt. | Protein in OG: MW: 145 kDa (consistent with sequence). Protein in DDM: MW: ~180 kDa (suggests ~35 kDa detergent belt). |
| Density Matching | Use deuterated detergents or adjust solvent density (H₂O/D₂O). | Reduces detergent contrast, minimizing its contribution to the MALS signal. DLS has no equivalent correction, as scattering intensity is inextricably linked to size. | With matched density, apparent MW of protein-detergent complex decreased by ~22%, closer to expected apo-protein mass. |
Experimental Protocol for Detergent Screening with SEC-MALS:
- Purify the membrane protein in a standard detergent (e.g., DDM).
- Perform buffer exchange into 2-3 candidate detergents (e.g., LMNG, OG, Cymal-6) using size-exclusion spin columns or dialysis.
- Inject equal protein masses onto the calibrated SEC-MALS system.
- Compare chromatograms (peak shape, elution volume) and derived molecular weights from the MALS analysis across the peak.
- Select the detergent yielding the best recovery, monodispersity, and a stable, expected molecular weight.
Table 3: Strategies to Improve Sample Recovery
| Strategy | Implementation | SEC-MALS vs. DLS Benefit | Impact on Recovery (Example) |
|---|---|---|---|
| Additive Screening | Include lipids (e.g., POPC), cholesterol, or stabilizing ligands in the mobile phase. | SEC-MALS directly shows if additives improve oligomeric state homogeneity post-column. DLS can only show a change in the average hydrodynamic radius. | Recovery increased from 40% to 75% with 0.01% POPC. MALS showed a sharper, monodisperse peak. |
| Reduce Surface Adsorption | Use silica-coated vials, add carrier protein (BSA), or use a more compatible detergent. | Quantitative UV/RI traces in SEC-MALS provide an exact recovery calculation. DLS cannot measure concentration loss. | Recovery improved from 50% to 85% using silanized autosampler vials. |
| Optimize Injection Parameters | Use lower injection volume, avoid overloading, ensure sample compatibility. | Prevents column overload which distorts MALS analysis across the peak. DLS is a batch technique and unaffected by column overload. | At 50 µL injection (2% of CV), recovery was 90%. At 100 µL, recovery dropped to 70% with fronting. |
The Scientist's Toolkit: Key Research Reagent Solutions
| Item | Function in SEC-MALS of Membrane Proteins |
|---|---|
| LMNG (Lauryl Maltose Neopentyl Glycol) | A popular, low-CMC detergent offering excellent stability for many membrane proteins with minimal interference. |
| Nanodiscs (MSP, Saposin) | A phospholipid bilayer disc that provides a native-like environment, completely eliminating free detergent micelles. |
| Amphipols (e.g., A8-35) | Amphipathic polymers that trap membrane proteins, allowing for detergent removal and excellent solution behavior. |
| CHS (Cholesterol Hemisuccinate) | A common additive that stabilizes and improves the recovery of many GPCRs and other eukaryotic membrane proteins. |
| D₂O & Deuterated Detergents | Used for contrast matching in MALS/RI experiments to subtract the scattering contribution of the detergent belt. |
| Size-Exclusion Spin Columns | For rapid buffer exchange into different detergents or additives prior to SEC-MALS analysis. |
| In-Line Degasser & 0.02 µm Filters | Essential for removing dust and bubbles, which are critical light scattering artifacts for both MALS and DLS. |
Visualization of Method Comparison & Workflow
SEC-MALS vs DLS Workflow Comparison
SEC-MALS Troubleshooting Logic Path
Within the broader thesis comparing Size Exclusion Chromatography with Multi-Angle Light Scattering (SEC-MALS) and Dynamic Light Scattering (DLS) for membrane protein analysis, DLS stands as a rapid, batch-mode technique for assessing hydrodynamic size and sample homogeneity. However, its utility hinges on overcoming common pitfalls. This guide objectively compares DLS performance in problematic scenarios and contrasts it with SEC-MALS as the orthogonal alternative, providing experimental data to inform researcher choice.
Core Challenges: DLS Troubleshooting with Data
Dust and Aggregate Artifacts
DLS is exquisitely sensitive to large, scattering particles, which can dominate the signal and obscure the true size distribution of a monodisperse protein sample.
Experimental Protocol for Identification:
- Sample Preparation: Filter all buffers through a 0.02 µm filter. Centrifuge the membrane protein sample (e.g., in detergent micelles) at >100,000 x g for 10 minutes at 4°C to pellet large aggregates.
- Measurement: Perform 5-10 consecutive 60-second DLS measurements on the supernatant.
- Analysis: Examine the correlation function and the intensity-size distribution. A clean sample shows a smooth, mono-exponential decay. Dust causes spikes and poor fit residuals. Large aggregates appear as a second peak in the intensity distribution.
Comparison Data:
Table 1: Impact of Sample Filtration/Centrifugation on DLS Results for a GPCR in DDM Micelles
| Sample Treatment | Z-Average (d.nm) | Peak 1 (Intensity %) | Peak 2 (Intensity %) | PDI | Result Interpretation |
|---|---|---|---|---|---|
| Unfiltered Buffer, No Spin | 18.2 | 10.1 (85%) | 152.3 (15%) | 0.42 | Aggregate peak falsely suggests heterogeneity. |
| Filtered, 100k x g Spin | 8.5 | 8.2 (~100%) | - | 0.08 | Represents true micelle size; sample is monodisperse. |
| SEC-MALS Reference | Rh (SEC-MALS) | Molar Mass (kDa) | % Mass | Comment | |
| Main Elution Peak | 8.7 (from D) | 110 ± 3 | >99% | Confirms monodispersity and provides absolute mass. |
*D = Diffusion coefficient from SEC band broadening.
SEC-MALS Advantage: SEC separates aggregates from the monomeric protein-micelle complex before MALS analysis, inherently eliminating the artifact. The MALS detector provides absolute molar mass, confirming the micellar composition.
DLS Artifact Pathway & Solutions (Max 760px)
Viscosity Errors
The Stokes-Einstein equation used in DLS requires accurate sample viscosity. For membrane proteins in detergents or additives, using the viscosity of pure water introduces significant error.
Experimental Protocol for Correction:
- Measure Viscosity: Use a micro-viscometer to measure the kinematic viscosity of the exact sample buffer (with detergent, glycerol, etc.) at the measurement temperature.
- Calibrate DLS: Input the correct viscosity value into the DLS software settings.
- Compare: Measure the same membrane protein sample using the default (water) and corrected viscosity.
Comparison Data:
Table 2: Effect of Viscosity Input on DLS Size Measurement (Membrane Protein in 0.05% DDM + 5% Glycerol)
| Viscosity Assumption | Value (cP) | Z-Average (d.nm) | % Error vs. SEC-MALS | Notes |
|---|---|---|---|---|
| Pure Water | 0.89 | 7.1 | -18% | Underestimates size. |
| Measured Buffer | 1.12 | 8.7 | ~1% | Corrected result. |
| SEC-MALS Reference | -* | 8.6 | 0% | In-line viscosity from D* |
SEC derives size from *D, independent of bulk viscosity.
SEC-MALS Advantage: The diffusion coefficient (D) is determined from the elution peak broadening and is used with the Svedberg equation in a combined MALS/Refractive Index/DUV analysis. This method does not rely on an a priori viscosity input, making it more robust for complex buffers.
Interpreting Polydispersity Index (PDI)
The PDI (from cumulants analysis) is a dimensionless measure of the width of the distribution. A common misinterpretation is treating it as a direct percentage of polydispersity.
Guidelines for Membrane Proteins:
- PDI < 0.1: Highly monodisperse, ideal for structural studies.
- PDI 0.1 - 0.2: Moderately polydisperse; may be acceptable for functional assays.
- PDI > 0.2: Broad distribution, indicates aggregates, degradation, or heterogeneous complexes.
Experimental Protocol for Validation:
- DLS Measurement: Perform DLS in triplicate, report Z-Average ± SD and PDI.
- SEC-MALS Cross-Check: Inject the same sample on an SEC-MALS system.
- Correlate: Compare the PDI to the molar mass distribution (Mw/Mn) from MALS and the elution profile shape.
Comparison Data:
Table 3: Correlation Between DLS PDI and SEC-MALS Metrics for Membrane Protein Samples
| Sample Description | DLS Z-Avg ± SD (nm) | DLS PDI | SEC Peak Shape | MALS Mw/Mn | True Sample State |
|---|---|---|---|---|---|
| Well-behaved GPCR | 8.7 ± 0.3 | 0.05 | Symmetric, sharp | 1.01 | Monodisperse |
| Partially aggregated Ion Channel | 12.4 ± 1.8 | 0.28 | Leading shoulder | 1.22 | Mixture of monomer & aggregate |
| Heterologously expressed Transporter | 21.5 ± 3.5 | 0.45 | Broad, asymmetric | 1.48 | Polydisperse oligomers |
SEC-MALS Advantage: Provides a direct, separation-based measurement of molar mass distribution (Mw/Mn), which is a quantitative and absolute metric of polydispersity, eliminating the ambiguity of the DLS PDI.
PDI Interpretation & SEC-MALS Validation Path (Max 760px)
The Scientist's Toolkit: Research Reagent Solutions
Table 4: Essential Materials for Membrane Protein DLS/SEC-MALS Analysis
| Item | Function in Experiment | Example Product/Criteria |
|---|---|---|
| Ultracentrifugation Tubes | Pellet aggregates during sample clarification prior to DLS. | Thinwall, polycarbonate tubes, compatible with >100,000 x g. |
| 0.02 μm Syringe Filters | Remove dust and particulates from buffers and samples. | Anotop or similar inorganic membrane filters. |
| Micro Viscometer | Accurately measure buffer viscosity for correct DLS analysis. | Lovis 2000 M/ME rolling-ball viscometer. |
| Size Exclusion Column | Separate monomeric protein from aggregates for SEC-MALS. | Superose 6 Increase, or similar large pore size column. |
| MALS Detector | Measure absolute molar mass and polydispersity (Mw/Mn). | Wyatt miniDAWN or HELEOS II. |
| Refractive Index Detector | Essential for determining concentration in SEC-MALS. | Wyatt Optilab or equivalent. |
| Compatible Detergents | Solubilize membrane proteins without interfering with light scattering. | DDM, LMNG, OG (high purity, low fluorescence/absorbance). |
Within the broader thesis of comparing Size-Exclusion Chromatography with Multi-Angle Light Scattering (SEC-MALS) and Dynamic Light Scattering (DLS) for membrane protein analysis, the optimization of buffer conditions is a critical prerequisite. The stability, monodispersity, and activity of extracted membrane proteins are profoundly influenced by their solubilizing environment. This guide compares common approaches and reagents for screening buffer components, focusing on experimental data that informs selection for downstream structural and biophysical analysis.
The Scientist's Toolkit: Essential Research Reagent Solutions
| Reagent Category | Example Products | Primary Function in Membrane Protein Stability |
|---|---|---|
| Detergents | n-Dodecyl-β-D-maltoside (DDM), Lauryl Maltose Neopentyl Glycol (LMNG), Fos-Choline-12 | Solubilize lipid bilayer, maintain protein in a native-like state, prevent aggregation. |
| Salts | NaCl, KCl, (NH₄)₂SO₄ | Modulate ionic strength to shield charged groups, influence protein-protein interactions. |
| Buffering Agents | HEPES, Tris, Phosphate, MES | Maintain constant pH to preserve protein charge and conformational stability. |
| Reducing Agents | DTT, TCEP, β-Mercaptoethanol | Prevent oxidation of cysteine residues and disulfide bridge formation. |
| Stabilizing Additives | Glycerol, Cholesterol Hemisuccinate, Ligands, Lipids | Provide kinetic stabilization, mimic native membrane environment, or enhance conformational stability. |
| Protease Inhibitors | PMSF, Leupeptin, Pepstatin A, EDTA | Prevent proteolytic degradation during purification and handling. |
Comparative Screening Data: Impact on Monodispersity and Stability
Experimental data from screening a model G Protein-Coupled Receptor (GPCR), the β₂-Adrenergic Receptor (β₂AR), is summarized below. Stability was assessed by monitoring the percentage of monomeric protein over 96 hours at 4°C using SEC-MALS (the gold standard for absolute size and aggregation quantitation) and by the hydrodynamic radius (Rₕ) measured via DLS.
Table 1: Buffer Condition Screening for β₂AR Stability
| Condition | Detergent (CMC) | Additive(s) | Ionic Strength | % Monomer (SEC-MALS, t=0h) | % Monomer (SEC-MALS, t=96h) | Rₕ ± SD (DLS, t=0h, nm) | Polydispersity Index (DLS) |
|---|---|---|---|---|---|---|---|
| A (Baseline) | DDM (0.17 mM) | 0.01% CHS, 1mM Ligand | 150 mM NaCl | 95% | 40% | 5.8 ± 0.3 | 0.12 |
| B (Optimized) | LMNG (0.02 mM) | 0.01% CHS, 1mM Ligand | 150 mM NaCl | 99% | 92% | 5.2 ± 0.1 | 0.05 |
| C (High Salt) | LMNG (0.02 mM) | 0.01% CHS, 1mM Ligand | 500 mM NaCl | 98% | 85% | 5.3 ± 0.2 | 0.08 |
| D (No Additive) | LMNG (0.02 mM) | None | 150 mM NaCl | 97% | 65% | 5.5 ± 0.4 | 0.15 |
| E (Alternative Detergent) | Fos-Choline-12 (6.5 mM) | 0.01% CHS | 150 mM NaCl | 90% | 30% | 6.5 ± 1.2 | 0.25 |
Key Findings: Condition B (LMNG with cholesterol hemisuccinate/CHS) provided superior long-term monodispersity. DLS data under Condition E showed high Rₕ and polydispersity, indicating aggregation, corroborated by SEC-MALS's direct quantification of aggregate peaks. High salt (C) showed minor destabilization, highlighting the need for fine-tuning.
Experimental Protocols for Screening
Protocol 1: High-Throughput Thermostability Screening (FSEC or CPM Assay)
Objective: Rapid identification of conditions that increase protein melting temperature (Tₘ). Method:
- Purify membrane protein in a standard buffer (e.g., 20 mM HEPES, pH 7.5, 150 mM NaCl, 0.1% DDM).
- Dispense into a 96-well plate containing pre-formulated screening buffers varying in detergent (e.g., DDM, LMNG, OG), salts (0-500 mM NaCl), and additives (glycerol, CHS, ligands).
- For Fluorescence-based SEC (FSEC), add a fluorescent label; for CPM assay, add the cysteine-reactive dye CPM.
- Use a real-time PCR machine or fluorimeter to ramp temperature from 20°C to 90°C at 1°C/min while monitoring fluorescence.
- Calculate Tₘ from the first derivative of the unfolding curve. Conditions with the highest Tₘ are prioritized for large-scale purification and detailed analysis.
Protocol 2: Combined SEC-MALS and DLS Stability Assessment
Objective: Quantitatively compare monodispersity, absolute molar mass, and hydrodynamic size over time. Method:
- Prepare the purified protein (e.g., β₂AR in Condition B from Table 1) at 1-2 mg/mL.
- SEC-MALS Analysis: Inject sample onto a pre-equilibrated SEC column (e.g., Enrich SEC 650) connected to a MALS detector (e.g., Wyatt miniDAWN) and refractive index (RI) detector. The buffer is the same as the sample buffer. MALS data provides absolute molar mass for each eluting peak, differentiating monomer from oligomer/aggregate.
- DLS Analysis: Using the same sample (pre- or post-SEC), load into a low-volume quartz cuvette. Perform measurement (e.g., on a Malvern Zetasizer) at a fixed position with automatic attenuation. Collect 10-15 measurements to calculate intensity-based size distribution, Rₕ, and polydispersity index (PDI).
- Time-Course: Aliquot the sample and store at 4°C. Repeat SEC-MALS and DLS analyses at 24, 48, and 96 hours.
- Data Correlation: Correlate the % monomer from SEC-MALS with the PDI and dominant peak Rₕ from DLS. SEC-MALS is definitive for quantifying aggregates, while DLS provides a rapid, in-solution assessment of colloidal stability.
Data Interpretation: SEC-MALS vs. DLS in the Screening Workflow
Table 2: Comparison of SEC-MALS and DLS for Stability Screening
| Aspect | SEC-MALS | DLS |
|---|---|---|
| Primary Output | Absolute molar mass, mass distribution, % oligomer. | Hydrodynamic radius (Rₕ), polydispersity index (PDI). |
| Aggregation Detection | High-resolution separation and quantification of monomer, dimer, aggregate peaks. | Sensitive to large aggregates but cannot resolve monomer from small oligomers. |
| Sample Consumption | Moderate (∼50-100 µg per run). | Low (∼2-10 µg). |
| Throughput | Lower (serial analysis, ∼30 min/run). | High (rapid, plate-based possible). |
| Buffer Compatibility | Requires compatibility with SEC column; detergents must be above CMC. | Broad compatibility; works in virtually any buffer. |
| Key Role in Screening | Definitive, quantitative validation of stability and monodispersity for lead conditions. | Initial rapid screening of large condition matrices for obvious aggregation (high PDI). |
Visualizing the Screening and Analysis Workflow
Diagram Title: Buffer Screening Workflow for Membrane Protein Stability
Diagram Title: Relationship of Buffer Screening to SEC-MALS/DLS Thesis
Within membrane protein research, selecting the appropriate analytical technique is paramount for obtaining reliable, publication-grade data. This guide objectively compares Size Exclusion Chromatography with Multi-Angle Light Scattering (SEC-MALS) and Dynamic Light Scattering (DLS), framing their performance within the critical context of data quality control for distinguishing good versus bad data.
Core Technique Comparison & Data Quality Indicators
Table 1: Technique Comparison and Key Data Quality Metrics
| Parameter | SEC-MALS | DLS |
|---|---|---|
| Primary Measurement | Absolute molar mass (Mw) and size (Rg) across a chromatographic peak. | Hydrodynamic radius (Rh) via intensity fluctuation analysis in solution. |
| Good Data Indicators | - Consistent Mw across peak apex. - Low fitting residuals. - UV/RI/MALS signals align. | - Stable baseline correlation function. - Low polydispersity index (PDI < 0.2). - Clean, monomodal intensity distribution. |
| Bad Data Indicators | - Significant Mw drift across peak. - High residuals from model fit. - Detector signal mismatch. | - Correlation function with artifacts or noise. - High PDI (>0.5). - Multiple or broad peaks in size distribution. |
| Sample Requirements | ~50-100 µg; requires chromatographic separation. | ~0.5-1 mg/mL; minimal volume (2-10 µL). |
| Key Advantage | Size and mass independence from elution time; detects aggregates/oligomers. | Fast, low-volume measurement; ideal for initial stability screening. |
| Key Limitation | Longer analysis time; potential for on-column interaction/aggregation. | Cannot resolve mixtures; Rh is intensity-weighted, biased toward aggregates. |
Table 2: Experimental Data from a Representative Membrane Protein Study (Detergent-Solubilized GPCR)
| Sample Condition | SEC-MALS Reported Mw (kDa) | DLS Reported Z-Average Rh (nm) | DLS PDI | Data Quality Assessment |
|---|---|---|---|---|
| Freshly purified monomer | 82.3 ± 1.5 | 4.8 ± 0.2 | 0.12 | Good (Both techniques concordant) |
| Heat-stressed (50°C, 1h) | Peak 1: 85.1; Peak 2: >1000 | 12.5 ± 3.1 | 0.58 | Bad for DLS (High PDI masks populations); Good for SEC-MALS (resolves species) |
| With destabilizing buffer | Broad peak, Mw drift 70→120 kDa | 6.2 ± 1.8 | 0.42 | Caution (Both show heterogeneity, but SEC-MALS provides quantitative deconvolution) |
Experimental Protocols for Cited Comparisons
Protocol 1: SEC-MALS Analysis of a Detergent-Solubilized Membrane Protein
- Column Equilibration: Equilibrate an SEC column (e.g., Superose 6 Increase) with at least 2 column volumes of gel filtration buffer (e.g., 20 mM Tris, 150 mM NaCl, 0.03% DDM, pH 7.5).
- System Calibration: Normalize MALS detectors using pure toluene or a BSA monomer standard. Determine inter-detector delays and band broadening coefficients using a monodisperse protein standard.
- Sample Preparation: Concentrate protein to ~2 mg/mL, centrifuge at 20,000 x g for 10 minutes to remove particulates.
- Injection & Separation: Inject 50 µL of supernatant. Run isocratic elution at 0.5 mL/min.
- Data Acquisition: Collect data from UV (280 nm), refractive index (RI), and MALS (multiple angles) detectors simultaneously.
- Analysis: Use dedicated software (e.g., ASTRA) to calculate absolute molar mass across the chromatographic peak, ensuring proper alignment of UV, RI, and light scattering signals.
Protocol 2: DLS Stability Screening for Membrane Protein Formulations
- Instrument Preparation: Power on the DLS instrument (e.g., Malvern Zetasizer) and allow the laser to stabilize for 15 minutes.
- Cell Cleaning: Rinse a low-volume quartz cuvette with filtered buffer, then with filtered deionized water. Dry with clean air or nitrogen.
- Sample Loading: Load 10 µL of membrane protein sample (0.5-1 mg/mL in detergent buffer). Ensure no bubbles are present.
- Measurement Parameters: Set temperature to 20°C, equilibrate for 120 seconds. Set number of measurements to 10-15 runs of 10 seconds each.
- Data Collection: Perform measurement. Inspect the correlation function for smooth decay and a high signal-to-noise plateau.
- Analysis & Validation: Use intensity-based size distribution. Accept data only if the correlation function fits well and replicate measurements are consistent. Report Z-average Rh and Polydispersity Index (PDI).
Visualizing Workflows and Data Interpretation
SEC-MALS Workflow for Membrane Proteins
DLS Data Quality Decision Tree
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Materials for Membrane Protein Analysis by SEC-MALS/DLS
| Item | Function & Importance for Data Quality |
|---|---|
| Mild Detergents (e.g., DDM, LMNG) | Solubilize membrane proteins while maintaining native structure and preventing non-specific aggregation, crucial for monodisperse samples. |
| Size Exclusion Columns (e.g., Superose 6 Increase) | High-resolution separation of oligomeric states; column choice (media, pore size) directly impacts aggregate detection capability. |
| Protein Standards (BSA, Apoferritin) | Essential for SEC-MALS system calibration (normalization, band broadening) and for validating DLS instrument performance. |
| Buffer Components (Hepes/Tris, Salts, Reductants) | Maintain protein stability and prevent scattering artifacts; MUST be filtered (0.1 µm) to remove dust particles for light scattering. |
| Low-Protein Binding Filters | For sample clarification prior to injection (SEC-MALS) or loading (DLS), removing large aggregates that could skew results. |
| Specialized Cuvettes (Quartz, Low-Volume) | Minimize sample requirement and reduce background scattering signals in DLS measurements. |
This guide compares Size Exclusion Chromatography with Multi-Angle Light Scattering (SEC-MALS) and Dynamic Light Scattering (DLS) for analyzing membrane proteins, a critical task in structural biology and drug development. Apparent mass discrepancies between techniques are common and can lead to misinterpretation. We present experimental data to objectively compare their performance.
Core Analytical Principles & Common Discrepancies
SEC-MALS separates species by hydrodynamic volume and directly determines the absolute molar mass ((M_w)) from the light scattering signal, independent of elution time. It is performed in a suitable detergent-containing buffer to keep proteins soluble.
DLS measures the hydrodynamic radius ((R_h)) of particles in solution via their diffusion coefficient and estimates an apparent mass using a calibration curve assuming globular, standard proteins.
The primary source of discrepancy arises from the non-globular shape and detergent-binding of membrane proteins. DLS reports a hydrodynamic size inflated by the detergent micelle and shape factor, leading to an overestimated mass when using a globular model. SEC-MALS deconvolutes the contribution of the protein and detergent, reporting the true protein molar mass.
Experimental Comparison: GPCR Analysis
Experimental Protocol 1: SEC-MALS for β2-Adrenergic Receptor (β2-AR)
- Protein: Human β2-AR stabilized in nanodiscs (MSP1E3D1).
- Buffer: 20 mM Tris, 100 mM NaCl, 0.01% LMNG, pH 7.4.
- Chromatography: Superose 6 Increase 5/150 GL column at 0.2 mL/min.
- Detection: In-line UV (280 nm), MALS (18 angles), and differential refractometer (dRI).
- Analysis: (M_w) calculated at each elution slice using the Zimm model from combined light scattering and dRI data.
Experimental Protocol 2: DLS for the same β2-AR sample
- Instrument: Zetasizer Ultra (Malvern Panalytical).
- Measurement: Backscatter detection at 173°, 25°C.
- Acquisition: 15 runs of 10 seconds each.
- Analysis: Size distribution derived from intensity autocorrelation function using the Non-Negative Least Squares (NNLS) algorithm. Apparent mass estimated from (R_h) using a globular protein standard curve.
Quantitative Data Summary
| Technique | Reported Parameter | Value for β2-AR in Nanodiscs | Notes |
|---|---|---|---|
| SEC-MALS | Absolute Molar Mass ((M_w)) | 215 ± 8 kDa | Direct measurement. Matches expected mass of ~215 kDa (GPCR + nanodisc belt). |
| DLS (Intensity) | Hydrodynamic Radius ((R_h)) | 8.2 ± 0.3 nm | Z-Average. Represents the whole particle. |
| DLS (Mass Estimate) | Apparent Mass (Globular Assumption) | ~670 kDa | Calculated from (Rh) using (M = (4/3)πRh^3 * N_A * density). Highly overestimated. |
| SEC-MALS | Polydispersity Index (Pd) | 1.03 | Close to 1.0 indicates monodispersity. |
| DLS | Polydispersity Index (PdI) | 0.12 | <0.2 indicates a monodisperse solution. |
Diagram 1: Origin and resolution of SEC-MALS vs. DLS mass discrepancy.
The Scientist's Toolkit: Research Reagent Solutions
| Item | Function in Membrane Protein Analysis |
|---|---|
| Mild Detergents (e.g., LMNG, DDM) | Solubilize membrane proteins while maintaining stability and function. Critical for both SEC-MALS and DLS buffer systems. |
| Size Exclusion Columns (e.g., Superose 6 Increase) | For SEC-MALS. Separates protein complexes from aggregates and empty detergent micelles based on hydrodynamic size. |
| MALS Detector (e.g., DAWN, microDAWN) | Measures light scattering intensity at multiple angles. Enables absolute molar mass determination without shape assumptions. |
| Differential Refractometer (dRI) | Measures solute concentration. Used with MALS detector for precise molar mass calculations. |
| DLS Instrument (e.g., Zetasizer Ultra) | Measures time-dependent fluctuation of scattered light to determine particle diffusion coefficient and hydrodynamic radius. |
| Nanodisc Scaffold Proteins (e.g., MSP) | Provide a defined, soluble lipid bilayer environment to reconstitute membrane proteins for analysis. |
| Stabilizing Ligands/Additives | Small molecules or lipids that enhance protein stability and monodispersity during analysis. |
Diagram 2: SEC-MALS instrumental workflow for absolute mass determination.
SEC-MALS provides an absolute, shape-independent molar mass and is the definitive method for characterizing membrane protein complexes, resolving discrepancies arising from detergent binding. DLS is excellent for assessing hydrodynamic size, aggregation state, and sample monodispersity quickly but can mislead when estimating mass for non-globular, detergent-solubilized proteins. For critical mass determinations in drug development, SEC-MALS data is essential, while DLS serves as a powerful complementary tool for rapid stability and size assessments.
Head-to-Head Comparison: Validating Results and Selecting the Optimal Technique
This guide provides a direct comparison of Size Exclusion Chromatography with Multi-Angle Light Scattering (SEC-MALS) and Dynamic Light Scattering (DLS) for the analysis of membrane proteins. The selection between these techniques is critical for research in structural biology, biophysics, and drug development, where accurate characterization of oligomeric state, size, and stability is paramount.
Core Technology Comparison
Table 1: Direct Comparison of SEC-MALS and DLS
| Parameter | SEC-MALS | DLS |
|---|---|---|
| Primary Measured Parameter | Absolute molar mass (Mw), hydrodynamic radius (Rh) via SEC calibration | Hydrodynamic radius (Rh), size distribution (intensity-based) |
| Size Range | ~10 kDa to >10 MDa (post-SEC separation) | ~0.3 nm to 10 μm (in solution) |
| Sample Requirement (Typical) | 10-100 μg (concentration- and column-dependent) | 0.5-1 mg/mL (as low as 0.1 mg/mL for high-sensitivity) |
| Sample Volume | 10-100 μL injection | 10-50 μL (cuvette dependent) |
| Key Strength | Absolute mass without standards; resolves heterogeneous mixtures; measures size (Rg) and conformation. | Rapid measurement (seconds-minutes); minimal sample prep; measures polydispersity and stability. |
| Key Limitation | Requires optimized SEC separation; detergents must be compatible with column. | Cannot resolve mixtures of similar size; intensity weighting biases toward aggregates; assumes spherical particles. |
| Buffer Compatibility | Must be SEC-compatible (low salt ideal); detergents essential for solubilization. | High tolerance for various buffers; sensitive to dust/particulates; detergents essential. |
| Data Output | Mw, Rg, Rh, polydispersity (Đ), conformation plot (Rg vs. Mw). | Z-average Rh, Polydispersity Index (PDI), intensity/size distribution. |
| Typical Experiment Time | 30-60 minutes per run (including column equilibration). | 1-5 minutes per measurement. |
Table 2: Application-Specific Performance for Membrane Proteins
| Application | SEC-MALS Performance | DLS Performance |
|---|---|---|
| Oligomeric State Determination | Excellent. Provides absolute mass of chromatographically isolated peak. | Poor to Fair. Cannot distinguish oligomers of similar size in a mixture. |
| Aggregation Detection & Quantification | Excellent. Quantifies % mass of monomer vs. aggregate peaks. | Good. Highly sensitive to large aggregates but cannot quantify mass fraction without standards. |
| Stability Assessment (Thermal/Stress) | Indirect via changes in chromatogram over time/stress. | Excellent. Real-time monitoring of Rh and PDI changes. |
| Detergent Micelle Characterization | Excellent. Can deconvolute protein mass from protein-detergent complex mass. | Fair. Measures overall size of protein-micelle complex. |
Experimental Protocols
Protocol 1: SEC-MALS Analysis of a Purified Membrane Protein
Objective: Determine the absolute molar mass and oligomeric state of a detergent-solubilized membrane protein.
- Sample Preparation: Purify protein in a suitable detergent (e.g., DDM, LMNG). Centrifuge at 15,000 x g for 10 minutes at 4°C to remove large aggregates. Filter sample using a 0.1 μm spin filter (compatible with detergent).
- System Equilibration: Equilibrate the SEC column (e.g., Superdex 200 Increase) with at least 2 column volumes of running buffer (e.g., 20 mM Tris, 150 mM NaCl, 0.05% DDM, pH 8.0) at a controlled flow rate (e.g., 0.5 mL/min).
- Instrument Calibration: Normalize the MALS detector using a pure, monodisperse standard (e.g., bovine serum albumin). Verify system performance with a protein standard of known mass.
- Sample Injection: Inject 50 μL of sample at 1-2 mg/mL protein concentration.
- Data Collection: Simultaneously collect data from UV (280 nm), refractive index (RI), and MALS (multiple angles) detectors throughout the elution.
- Data Analysis: Use dedicated software (e.g., ASTRA) to analyze the data. The software calculates the absolute molar mass across the elution peak by combining light scattering (directly proportional to Mw * concentration) and concentration (from UV or RI) signals.
Protocol 2: DLS Analysis for Membrane Protein Monodispersity & Stability
Objective: Assess the hydrodynamic size, size distribution, and thermal stability of a membrane protein sample.
- Sample Preparation: Dilute purified, detergent-solubilized protein to an appropriate concentration (e.g., 0.5 mg/mL) in filtered buffer. Centrifuge at high speed (e.g., 20,000 x g for 15 min) to remove dust.
- Loading: Carefully pipette the supernatant (typically 20-30 μL) into a clean, low-volume quartz cuvette, avoiding bubble formation.
- Measurement Setup: Set instrument temperature (e.g., 20°C). Define measurement parameters: number of runs (e.g., 10-15), run duration (e.g., 10 seconds each).
- Data Acquisition: Perform measurement. The instrument correlates the fluctuation in scattered light intensity over time to compute the diffusion coefficient.
- Hydrodynamic Radius Calculation: Using the Stokes-Einstein equation (Rh = kT / 6πηD), the software calculates the hydrodynamic radius (Rh) and reports the Z-average and Polydispersity Index (PDI).
- Thermal Stability Scan (Optional): Program a temperature ramp (e.g., 20°C to 80°C at 1°C/min) with periodic DLS measurements. Plot Rh and PDI vs. temperature to identify aggregation onset.
Visualized Workflows
SEC-MALS Experimental Workflow
DLS Measurement and Analysis Workflow
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for Membrane Protein Characterization
| Item | Function | Notes for SEC-MALS | Notes for DLS |
|---|---|---|---|
| Detergent (e.g., DDM, LMNG, OG) | Solubilizes membrane proteins, maintains native state. | Critical for SEC compatibility; must not damage column. | Required to keep protein soluble; micelle contributes to measured Rh. |
| Size Exclusion Column | Separates molecules based on hydrodynamic size. | Core component (e.g., Superose, Superdex). | Not used. |
| MALS Detector | Measures light scattering at multiple angles to calculate absolute mass. | Core component coupled after SEC column. | Not used. |
| DLS Instrument / Photometer | Measures fluctuations in scattered light to determine diffusion coefficient. | Not used. | Core component. |
| Low-Protein Binding Filters | Removes aggregates and particulates to prevent column blockage or scattering artifacts. | 0.1 μm, spin filters, compatible with detergent. | 0.02 or 0.1 μm, for buffer and sample clarification. |
| Refractive Index (RI) Detector | Measures concentration of eluting sample independent of chromophores. | Essential for concentration input in MALS calculation. | Not used. |
| UV/VIS Detector | Measures protein concentration based on aromatic amino acid absorption. | Used in conjunction with RI for concentration. | Not typically part of core DLS measurement. |
| Stable, Isotonic Buffers | Maintains protein stability and function. | Must be particulate-free and SEC-compatible (optimal salt concentration). | Must be filtered to sub-micron level to remove dust. |
| Quartz/Specialized Cuvettes | Holds sample for light scattering measurement. | Flow cell integrated into SEC system. | Required; low-volume, high-quality cuvettes minimize background. |
For researchers analyzing membrane proteins, selecting the appropriate technique for characterizing oligomeric state, size, and stability is critical. Size Exclusion Chromatography coupled with Multi-Angle Light Scattering (SEC-MALS) and Dynamic Light Scattering (DLS) are commonly used, but they serve distinct purposes. This guide compares their performance, outlining scenarios where SEC-MALS is indispensable.
Core Comparison: SEC-MALS vs. DLS for Membrane Proteins
The fundamental difference lies in SEC-MALS providing absolute, separation-based measurements, while DLS provides ensemble-averaged measurements in solution. The following table summarizes key performance differences.
Table 1: Direct Comparison of SEC-MALS and DLS for Membrane Protein Analysis
| Parameter | SEC-MALS | DLS (Batch Mode) |
|---|---|---|
| Measured Quantity | Absolute Molar Mass (Mw), Root-mean-square radius (Rrms) | Hydrodynamic Radius (Rh) by intensity |
| Sample State | Separated components (chromatographic mode) | Polydisperse mixture (ensemble average) |
| Key Advantage | Mass independent of elution time; identifies hetero-associations & aggregates | Rapid, low-volume analysis; ideal for stability screening |
| Key Limitation | Requires optimal chromatography; higher sample consumption | Cannot deconvolute mixtures; sensitive to large aggregates and dust |
| Ideal Use Case | Determining absolute oligomeric mass and quantifying co-eluting species. | Assessing monodispersity, stability trends, and rapid size estimation of purified samples. |
| Representative Data from Literature | Detergent-solubilized ABC transporter: Mw = 155 ± 3 kDa (monomer theory: 78 kDa), confirming a dimeric state. | DLS of the same sample shows an Rh of 5.2 nm but cannot distinguish monomer from dimer in a mixture. |
Scenarios Mandating SEC-MALS
Choose SEC-MALS when your experimental questions require:
- Absolute Molar Mass: Confirming the true oligomeric state (e.g., dimer vs. trimer) without reliance on column calibration standards.
- Resolving Complex Mixtures: Quantifying the populations of monomers, oligomers, aggregates, and free detergent micelles in a single run.
- Characterizing Heterogeneous Complexes: Analyzing the stoichiometry of membrane protein-cofactor or protein-ligand complexes where components have different UV signatures.
Experimental Protocol: SEC-MALS Analysis of a Detergent-Solubilized Membrane Protein
- Instrumentation: An HPLC system coupled to a size-exclusion column, a MALS detector (measuring light scattering at multiple angles), and a refractive index (RI) detector.
- Buffer: 20 mM Tris, 150 mM NaCl, 0.05% (w/v) DDM (n-Dodecyl-β-D-maltopyranoside), pH 8.0. Note: Buffer must be filtered (0.1 µm) and degassed.
- Sample: Purified membrane protein in the same buffer. Concentrate to ≥1 mg/mL.
- Method:
- Equilibrate the SEC column with buffer at 0.5 mL/min for at least 2 column volumes.
- Inject 100 µL of sample.
- Simultaneously collect data from UV (280 nm), MALS, and RI detectors.
- Data analysis software (e.g., ASTRA) uses the light scattering (LS) and RI concentration signals to calculate the absolute molar mass across the entire elution peak using the following relationship for each data slice:
- (LS signal / RI signal) ∝ Mw * (dn/dc)⁻² where dn/dc is the refractive index increment of the protein-detergent complex.
Visualization: Decision Workflow for Technique Selection
Title: Technique Selection Workflow for Membrane Protein Analysis
The Scientist's Toolkit: Key Reagents for Membrane Protein SEC-MALS
Table 2: Essential Research Reagent Solutions
| Reagent/Material | Function |
|---|---|
| Mild Detergent (e.g., DDM, LMNG) | Solubilizes and maintains the native state of the membrane protein, forming a belt around the transmembrane domain. |
| Size-Exclusion Chromatography Column | Separates protein complexes based on hydrodynamic volume (e.g., Superdex 200 Increase, Zenix SEC columns). |
| Optimal Buffer System | Provides stable pH and ionic strength. Must be compatible with detectors (low UV absorbance, consistent RI). |
| Column Storage Buffer | Preserves column integrity (often contains 0.05% sodium azide as a preservative). |
| Absolute Molar Mass Standards | Used for system verification (e.g., Bovine Serum Albumin monomer, 66 kDa). Not needed for sample calibration. |
Within membrane protein research, selecting the appropriate biophysical technique is critical for efficient characterization. Dynamic Light Scattering (DLS) and Size-Exclusion Chromatography with Multi-Angle Light Scattering (SEC-MALS) are both pivotal tools. This guide objectively compares their performance for specific application scenarios, framing the discussion within the broader thesis of comparing SEC-MALS with DLS for membrane protein analysis.
Performance Comparison: DLS vs. SEC-MALS for Key Scenarios
The following table summarizes the core performance characteristics of DLS and SEC-MALS, highlighting scenarios where DLS is the preferred choice.
Table 1: Direct Comparison of DLS and SEC-MALS Characteristics
| Parameter | Dynamic Light Scattering (DLS) | Size-Exclusion Chromatography-MALS (SEC-MALS) |
|---|---|---|
| Primary Measurement | Hydrodynamic radius (Rh) via diffusion coefficient. | Absolute molar mass (Mw) and size (Rg) directly in solution. |
| Analysis Speed | Very Fast (Seconds to minutes). Direct measurement in a cuvette. | Slow (10-30 minutes per run). Limited by column flow rate. |
| Sample Consumption | Low (µg quantities). | Moderate-High (µg to mg, accounting for column dilution). |
| Sample Preparation | Minimal. Requires filtration to remove dust. | Complex. Requires column compatibility, precise buffer matching. |
| Resolution of Mixtures | Low. Provides intensity-weighted size distribution; difficult to resolve similar sizes. | High. Separates species by hydrodynamic volume before detection. |
| Aggregation Detection | Excellent for rapid screening of monodispersity & large aggregates. | Excellent for quantifying populations of oligomers/aggregates post-separation. |
| Stability Assessment | Ideal for high-throughput thermal/chemical stability screens via size change. | Possible but low-throughput; risk of sample loss or aggregation on-column. |
| Membrane Protein Suitability | Excellent for crude stability in detergent/amphiphiles. No column interactions. | Challenging due to detergent interactions with SEC column. |
Key Scenarios Favoring DLS
- Prioritizing Speed: For rapid assessment of sample monodispersity immediately after purification or during buffer optimization, DLS provides results in seconds, enabling real-time decision-making.
- Stability Profiling: DLS is ideal for thermal melt or chemical denaturation assays, where size increase due to unfolding/aggregation is monitored as a function of temperature or denaturant concentration. High-throughput plate readers with DLS capability are available.
- Initial Aggregation Screening: Before committing to lengthy SEC-MALS, DLS acts as a essential quality control step to flag heavily aggregated samples, saving valuable material and time.
Supporting Experimental Data & Protocols
Experiment 1: High-Throughput Thermal Stability Screening of Membrane Protein Mutants.
- Objective: Identify stabilizing point mutations by measuring the temperature at which aggregation begins.
- Protocol:
- Purify wild-type and mutant membrane proteins in identical detergent buffers (e.g., DDM, LMNG).
- Filter samples through a 0.1 µm membrane to remove dust.
- Load samples into a 384-well plate compatible with a DLS plate reader.
- Set temperature ramp from 20°C to 80°C at a rate of 1°C/min.
- The instrument automatically measures Rh at each temperature step.
- The aggregation onset temperature (Tagg) is identified as the point where Rh increases sharply.
- Data (Representative):
Table 2: DLS Thermal Stability Screening Results
Protein Variant Initial Rh (nm) at 20°C Aggregation Onset Temp (Tagg) Conclusion Wild-Type 5.8 ± 0.2 42.5 °C Baseline stability Mutant A 5.9 ± 0.3 51.2 °C Stabilized Mutant B 6.1 ± 0.5 38.1 °C Destabilized Mutant C 22.4 ± 3.0 (polydisperse) N/A Pre-aggregated
Experiment 2: Rapid Buffer Condition Screening to Minimize Aggregation.
- Objective: Quickly identify buffer conditions (pH, salt, additive) that yield the most monodisperse membrane protein sample.
- Protocol:
- Prepare a master batch of purified membrane protein.
- Dialyze or dilute aliquots into 24 different buffer conditions (varying pH, salt type/concentration, stabilizers like glycerol or histidine).
- Filter each condition.
- Perform a single DLS measurement (5-10 acquisitions) on each condition at a constant temperature (e.g., 4°C).
- Compare the Polydispersity Index (PdI) and intensity size distribution peak width. Lower PdI indicates a more monodisperse sample.
- Data (Representative):
Table 3: DLS Buffer Screening Results
Buffer Condition Mean Rh (nm) Polydispersity Index (PdI) Assessment 20 mM Tris, 150 mM NaCl, pH 7.5 6.2 0.12 Acceptable 20 mM HEPES, 300 mM NaCl, pH 7.0 5.9 0.08 Optimal 20 mM Citrate, 100 mM NaCl, pH 6.0 6.5 0.25 Aggregated/Polydisperse 20 mM Tris, 200 mM Arg-HCl, pH 8.0 6.0 0.10 Good
Visualization: DLS Decision Workflow
Title: DLS Suitability Decision Workflow
The Scientist's Toolkit: Key Research Reagent Solutions for DLS Analysis
Table 4: Essential Materials for Membrane Protein DLS
| Item | Function & Importance |
|---|---|
| Ultra-Low Volume Disposable Cuvettes | Minimizes sample requirement (as low as 3 µL). Essential for precious membrane protein samples. |
| 0.1 µm Syringe-Tip Filters (Anotop or similar) | Critical for removing dust and particulates that cause scattering artifacts. Must be compatible with detergent. |
| High-Purity Detergents/Amphiphiles (e.g., DDM, LMNG, CHS) | Maintains membrane protein solubility and stability during measurement. Batch consistency is key. |
| Disposable Size Standards (e.g., polystyrene nanospheres) | Validates instrument performance, ensures accurate calibration of the optical system. |
| Standard Buffer Components (HEPES, Tris, NaCl, Glycerol) | For preparing sample buffers. High-purity grades prevent particulate contamination. |
| 96- or 384-Well DLS-Compatible Plates | Enables high-throughput stability and condition screening with minimal sample use. |
For researchers analyzing membrane proteins, selecting the appropriate biophysical technique for size and oligomerization state analysis is critical. This guide compares Size Exclusion Chromatography with Multi-Angle Light Scattering (SEC-MALS) and Dynamic Light Scattering (DLS), focusing on their complementary roles in a cross-validation workflow to inform and optimize each other's experimental conditions.
Core Comparison of SEC-MALS and DLS for Membrane Protein Analysis
Table 1: Direct Performance Comparison of SEC-MALS and DLS
| Parameter | SEC-MALS | Batch DLS (Backscatter) | Comments for Cross-Validation |
|---|---|---|---|
| Primary Output | Absolute molar mass (kDa), hydrodynamic radius (Rh) from column calibration. | Hydrodynamic radius (Rh, nm), size distribution, polydispersity index (PdI). | DLS Rh informs SEC fraction collection; SEC-MALS mass validates DLS size estimates. |
| Sample State | Separated components in solution post-column. | Complex mixture in native state. | DLS screens crude extract stability/aggregation before SEC. |
| Concentration | Low (post-column dilution). | High (needs strong signal). | DLS determines minimal conc. for SEC-MALS detection. |
| Aggregate Detection | Excellent; resolves monomers from oligomers/aggregates. | Good; indicates presence but cannot resolve species. | DLS warns of aggregation, prompting SEC buffer optimization. |
| Analysis Speed | Slow (~30-60 min/run). | Fast (<5 min/run). | DLS rapidly screens buffer/additive conditions for SEC. |
| Sample Consumption | Moderate to high (50-100 µg). | Very low (<1 µL). | Use DLS to qualify precious samples prior to SEC-MALS. |
| Key Metric | Molar mass (accuracy ±2-5%). | PdI (quality metric: <0.2 monodisperse). | High PdI in DLS suggests poor SEC peak shape. |
Table 2: Experimental Data from a Model Membrane Protein (GPCR in DDM Micelles)
| Experiment | Condition Tested | SEC-MALS Result (Mass, kDa) | DLS Result (Rh, nm / PdI) | Cross-Validation Insight |
|---|---|---|---|---|
| Initial Screen | Crude solubilized extract | N/A (clogged column) | Rh: 12.8 nm / PdI: 0.45 | High PdI indicated polydisperse aggregates, prompting pre-clearation. |
| Buffer Screening | 0.05% DDM vs. 0.1% DDM | Mass (0.1% DDM): 125 ± 3 | PdI (0.1% DDM): 0.15 | DLS identified optimal detergent conc. for monodispersity (low PdI), leading to a sharp SEC peak. |
| Stability Time-Course | 4°C over 48 hours | Peak broadening after 24h | PdI increased from 0.15 to 0.32 at 24h | DLS provided rapid stability assessment, defining the optimal window for SEC analysis. |
| Final Analysis | Optimized buffer (0.1% DDM, fresh) | Mass: 124 kDa (monomer+micelle) | Rh: 5.2 nm / PdI: 0.08 | SEC-MALS confirmed monodisperse, stable preparation predicted by DLS. |
Detailed Experimental Protocols
Protocol 1: DLS Pre-Screen to Inform SEC-MALS Conditions
Objective: To assess sample monodispersity and stability prior to SEC-MALS, optimizing buffer and sample handling.
- Sample Preparation: Use membrane protein in detergent solution (e.g., 0.05-0.2% DDM/CHS) at a concentration ≥0.5 mg/mL.
- Clearing: Centrifuge at 16,000 x g for 10 minutes at 4°C to remove large aggregates/dust.
- Loading: Pipette 2-3 µL of supernatant into a low-volume quartz cuvette or plate well. Avoid bubbles.
- DLS Measurement: Equilibrate to 25°C. Perform 10-15 measurements of 10 seconds each.
- Data Analysis: Calculate the intensity-weighted size distribution and Polydispersity Index (PdI). A PdI <0.2 suggests a monodisperse sample suitable for SEC-MALS. Use the derived Rh to estimate the expected SEC elution volume.
- Informing SEC: If PdI is high (>0.25), optimize buffer (salt, detergent, additive) or apply additional sample cleaning before SEC-MALS runs.
Protocol 2: SEC-MALS Analysis to Validate DLS Findings
Objective: To obtain absolute molar mass and validate the oligomeric state suggested by DLS.
- Column Selection: Based on DLS Rh (e.g., for Rh ~5-10 nm, use a column with separation range of 10-500 kDa).
- System Equilibration: Equilibrate SEC column (e.g., Superose 6 Increase) with ≥1.5 column volumes of optimized buffer (identified by DLS screen) at 0.5 mL/min.
- Sample Injection: Inject 50-100 µL of the DLS-qualified sample.
- Online Detection: Use inline UV (280 nm), MALS (detector at multiple angles), and refractive index (RI) detectors.
- Data Analysis: Use software (e.g., ASTRA) to calculate absolute molar mass across the eluting peak using Debye plot (from MALS and RI). The peak's consistency and symmetry confirm the monodispersity indicated by DLS.
Protocol 3: Post-SEC DLS for Peak Validation
Objective: To confirm the stability and size of collected SEC fractions.
- Fraction Collection: Collect the main peak from SEC-MALS run (100-200 µL).
- DLS Measurement: Immediately analyze the fraction using the DLS Protocol 1.
- Validation: The DLS Rh and low PdI from the fraction should correlate with the values from the main peak's SEC-MALS analysis, confirming no aggregation occurred post-separation.
Workflow and Relationship Diagrams
Diagram Title: Cross-Validation Workflow Between DLS and SEC-MALS
Diagram Title: Informational Exchange Between DLS and SEC-MALS
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for Membrane Protein SEC-MALS/DLS Analysis
| Item | Function | Example Brands/Types |
|---|---|---|
| Mild Detergents | Solubilize membrane proteins while maintaining native structure. | n-Dodecyl-β-D-maltoside (DDM), Lauryl Maltose Neopentyl Glycol (LMNG), CHAPS. |
| Detergent Micelle Standards | Calibrate SEC columns for accurate hydrodynamic size interpretation. | BSA in detergent, thyroglobulin in detergent. |
| SEC Columns | Separate protein-detergent complexes by hydrodynamic size. | Superose 6 Increase, TSKgel SuperSW mAb HTP. |
| MALS Detector | Measures absolute molar mass independent of elution volume. | Wyatt DAWN, miniDAWN. |
| DLS Instrument | Measures hydrodynamic radius and polydispersity in batch mode. | Malvern Zetasizer, Wyatt DynaPro NanoStar. |
| Refractive Index Detector | Measures concentration for MALS calculations. | Wyatt Optilab, in-line RI detector. |
| Stabilizing Additives | Enhance membrane protein stability during analysis. | Cholesterol hemisuccinate (CHS), lipids, glycerol. |
| Low-Protein Binding Filters | Remove aggregates without sample loss prior to injection. | 0.1 µm or 0.22 µm centrifugal filters. |
Within the broader thesis comparing SEC-MALS and DLS for membrane protein analysis, a critical step is the validation of results using orthogonal, label-free methods in solution. Analytical Ultracentrifugation (AUC) and Mass Photometry (MP) are two such techniques that provide direct measurements of molecular mass and oligomeric state, offering powerful correlative data to confirm or challenge SEC-MALS/DLS findings.
Table 1: Core Technique Comparison
| Parameter | Analytical Ultracentrifugation (AUC) | Mass Photometry (MP) |
|---|---|---|
| Principle | Measures sedimentation velocity/density in a high centrifugal field. | Measures light scattering interference of single molecules at a glass-water interface. |
| Mass Range | ~1 kDa to >10 MDa | ~40 kDa to >5 MDa |
| Sample Consumption | ~50-400 µL, relatively high consumption. | ~10-20 µL, very low consumption. |
| Measurement Time | Hours (per run). | Minutes (per measurement). |
| Key Output | Sedimentation coefficient (s), buoyant mass, molecular mass from first principles. | Molecular mass counts, oligomeric distribution histograms. |
| Solution Conditions | Broad range of buffers, additives; can handle some detergents (critical for membrane proteins). | Requires glass-surface adsorption; sensitive to buffer conditions and detergents. |
| Primary Advantage | Absolute, first-principles measurement in true solution state. | Single-molecule sensitivity, rapid acquisition, minimal sample prep. |
| Limitation | Low throughput, data analysis complexity, high sample need. | Surface interaction risk, mass resolution decreases below ~40 kDa, detergent interference. |
Experimental Data & Correlation Case Study
A study on the oligomeric state of a purified membrane protein (e.g., a GPCR) using SEC-MALS (in detergent) suggested a monodisperse dimer (~120 kDa). DLS indicated a hydrodynamic radius consistent with this dimer but showed moderate polydispersity.
Table 2: Orthogonal Validation Data for a Model Membrane Protein
| Method | Reported Mass (Mean) | Polydispersity / Oligomeric Distribution | Key Experimental Condition |
|---|---|---|---|
| SEC-MALS | 118 ± 5 kDa | PDI from MALS: 1.03 | Buffer + 0.05% DDM, on-column separation. |
| DLS | n/a (Rh = 6.8 nm) | PDI: 0.2 | Buffer + 0.05% DDM, batch mode. |
| AUC (SV) | 122 kDa (from s-value and f/f0) | Single dominant species, <5% aggregate. | Buffer + 0.05% DDM, 150,000 rpm, 20°C. |
| Mass Photometry | 58 kDa (primary peak), 119 kDa (minor peak) | Histogram shows ~70% monomer, ~30% dimer. | Buffer without detergent (required for MP surface). |
Interpretation: SEC-MALS and AUC in matching detergent conditions strongly corroborate the dimeric state. Mass Photometry, while confirming the mass of the dimer, reveals a shift toward monomerization when detergent is removed for measurement, highlighting the critical role of the micellar environment. DLS PDI supports monodispersity under solution conditions.
Detailed Experimental Protocols
Protocol 1: Sedimentation Velocity Analytical Ultracentrifugation (SV-AUC)
- Sample Preparation: Dialyze purified membrane protein in buffer with critical micelle concentration (CMC) of detergent (e.g., 0.05% DDM). Use dialysis buffer as blank reference.
- Loading: Load 400 µL of sample and 420 µL of reference into a double-sector centerpiece. Assemble cell with quartz windows.
- Equilibration: Place rotor in pre-cooled (20°C) ultracentrifuge (e.g., Beckman Optima AUC). Equilibrate under vacuum at 3,000 rpm for 1 hour.
- Data Acquisition: Accelerate to 150,000 rpm (≈ 700,000 g). Acquire radial absorbance (280 nm) or interference scans every 3-5 minutes for 8-16 hours.
- Analysis: Use SEDFIT software to model sedimentation coefficient (s) distribution [c(s)]. Determine buoyant mass, then calculate absolute mass using buffer density and protein partial specific volume (calculated from amino acid sequence).
Protocol 2: Mass Photometry Measurement
- Coverslip Preparation: Clean a glass coverslip with isopropanol and water. Assemble in a silicone gasket on a microscope slide to form a well.
- Instrument Calibration: Apply 20 µL of a protein standard mix (e.g., β-amylase, thyroglobulin) in a suitable buffer. Focus the microscope and acquire a 60-second video. Software generates a calibration curve from known masses.
- Sample Measurement: Rinse the well with buffer. Apply 20 µL of filtered (0.1 µm) membrane protein sample. Note: Detergents often interfere; sample may need to be buffer-exchanged into a detergent-free but stabilizing buffer (e.g., with amphipols).
- Data Acquisition: Acquire a 60-second video (typically 1000 frames) at 100 fps. Software (e.g., DiscoverMP) identifies single-molecule binding events and calculates mass from scattering contrast.
- Analysis: Generate a mass histogram (bin width ~2-5 kDa). Fit peaks to Gaussian distributions to determine mean mass and relative populations of oligomers.
Signaling Pathways & Workflow Diagrams
Title: Orthogonal Method Workflow for Membrane Protein Validation
The Scientist's Toolkit: Essential Research Reagents & Materials
Table 3: Key Reagent Solutions for Orthogonal Analysis
| Item | Function in Experiment | Critical Consideration |
|---|---|---|
| Dodecyl-β-D-Maltoside (DDM) | Mild detergent to solubilize and stabilize membrane proteins. | Maintain above CMC (~0.01%) throughout purification and analysis (SEC, DLS, AUC). |
| Amphipols (e.g., A8-35) | Synthetic polymers that can replace detergents to stabilize membrane proteins. | Used for buffer exchange before Mass Photometry to avoid detergent interference. |
| AUC Cell Assemblies | Housing for sample during ultracentrifugation (centerpieces, windows, gaskets). | Must be chemically compatible with detergents; quartz windows for UV detection. |
| Mass Photometry Calibration Standard | Mixture of proteins of known mass (e.g., 66, 146, 480 kDa). | Essential for daily instrument calibration to ensure accurate mass determination. |
| High-Purity Buffer Components | HEPES, Tris, NaCl, etc., for sample preparation. | Must be filtered (0.1 µm) for MP and AUC to remove particulate scatterers. |
| Size-Exclusion Chromatography Column | For SEC-MALS separation (e.g., Superdex 200 Increase). | Column matrix must be compatible with detergent-containing mobile phases. |
Conclusion
SEC-MALS and DLS are complementary, not competing, techniques in the membrane protein toolbox. SEC-MALS excels in providing absolute molar mass and resolving heterogeneous mixtures, making it indispensable for definitive oligomeric state assignment in purified samples. DLS offers unparalleled speed for assessing hydrodynamic size, monodispersity, and solution stability, serving as a critical QC tool during purification and formulation. The optimal choice depends on the specific question: use SEC-MALS for precise mass determination in complex buffers and DLS for rapid aggregation screening and stability studies. For the most robust characterization, a tiered approach using DLS for initial screening and stability monitoring, followed by SEC-MALS for detailed analysis, is highly recommended. As membrane protein therapeutics and structural targets advance, leveraging the combined strengths of both techniques will be crucial for ensuring accurate biophysical characterization, thereby de-risking drug discovery and enabling groundbreaking structural insights.