GroEL vs. DnaK: A Comparative Analysis of Chaperonin and Hsp70 Refolding Efficiency in Protein Homeostasis
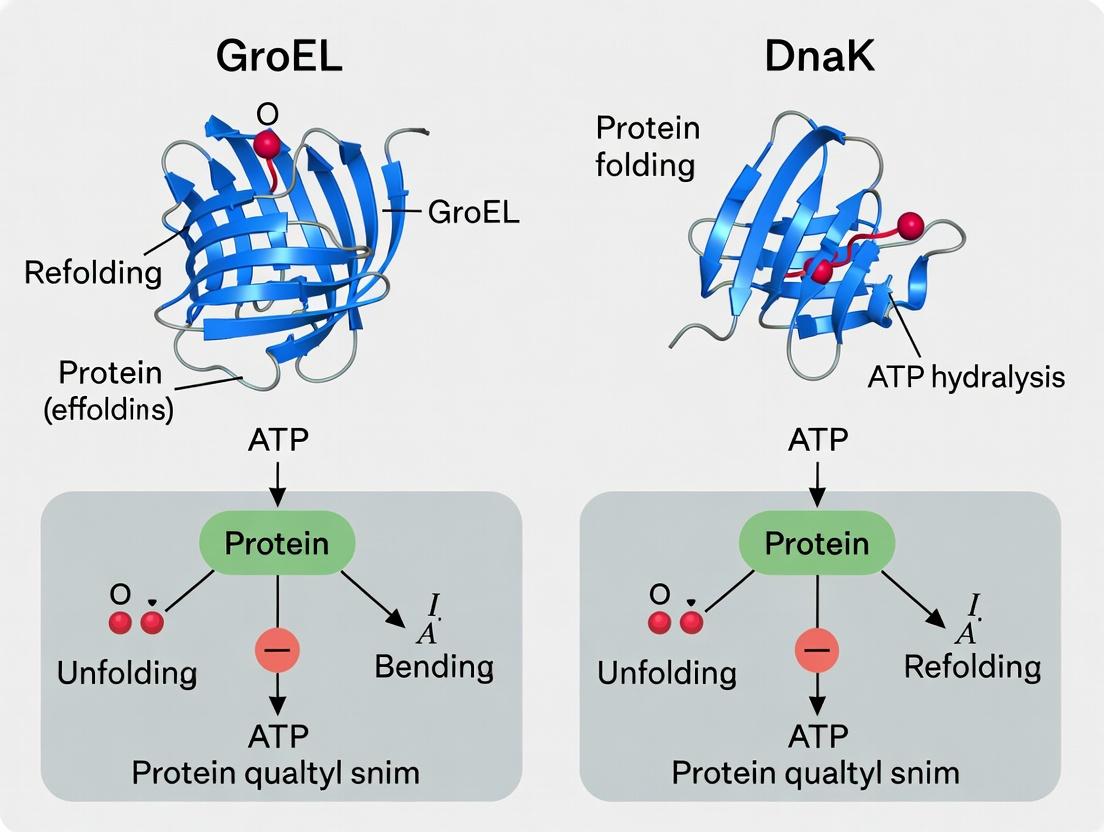
This article provides a comprehensive analysis comparing the protein refolding efficiencies of the GroEL/GroES chaperonin system and the DnaK (Hsp70) chaperone system.
GroEL vs. DnaK: A Comparative Analysis of Chaperonin and Hsp70 Refolding Efficiency in Protein Homeostasis
Abstract
This article provides a comprehensive analysis comparing the protein refolding efficiencies of the GroEL/GroES chaperonin system and the DnaK (Hsp70) chaperone system. Targeted at researchers and drug development professionals, it explores the foundational mechanisms, methodological applications in experimental and therapeutic contexts, common challenges in assay design, and direct comparative data on substrate specificity, refolding rates, and ATP consumption. The synthesis of current research aims to inform experimental design and the development of chaperone-targeted therapies for protein misfolding diseases.
Understanding the Machines: Core Mechanisms of GroEL/ES and DnaK in Protein Folding
The Anfinsen Dogma and the Necessity of Molecular Chaperones
The Anfinsen dogma posits that a protein's amino acid sequence uniquely determines its three-dimensional, functional native structure. However, in the crowded cellular environment, protein folding is error-prone, leading to aggregation. This necessitates molecular chaperones, which prevent misfolding and promote correct folding. This guide compares the performance of two major chaperone systems, GroEL (Hsp60) and DnaK (Hsp70), in protein refolding assays, a core focus in current chaperone research.
Performance Comparison: GroEL versus DnaK
The following table summarizes key experimental data comparing the refolding efficiency of the GroEL/ES and DnaK/DnaJ/GrpE systems.
Table 1: Comparative Refolding Efficiency of GroEL and DnaK Systems
| Metric | GroEL/ES System | DnaK/DnaJ/GrpE System | Experimental Context |
|---|---|---|---|
| Optimal Substrate Size | ≤ 60 kDa (encapsulated) | Broad range, smaller preferred | Refolding of chemically denatured model substrates in vitro. |
| ATP Molecules per Refolding Cycle | ~14 (7 per ring, two rings) | 1 per client polypeptide | Measured using radioactive [γ-³²P]ATP hydrolysis assays. |
| Refolding Rate (k, s⁻¹) | 0.01 - 0.1 | 0.001 - 0.01 | Refolding of monomeric rhodanese or luciferase at 25°C. |
| Anfinsen Compliance | Active, ATP-dependent folding | Passive, prevents aggregation; facilitates folding | Ability to refold proteins to native state without additional information. |
| Primary Mechanism | Anfinsen cage (isolated chamber) | Holdase & iterative annealing | Structural data from cryo-EM and single-molecule spectroscopy. |
| Co-chaperone Requirement | Essential (GroES) | Essential for efficiency (DnaJ, GrpE) | Knockout/omission experiments showing loss of function. |
Detailed Experimental Protocols
Protocol 1: Standard In Vitro Refolding Assay for Chaperone Efficiency
- Objective: Quantify the recovery of native enzyme activity from a denatured state in the presence of a chaperone system.
- Procedure:
- Denaturation: Dilute a purified, sensitive enzyme (e.g., mitochondrial rhodanese) into a denaturing buffer (6 M GuHCl, 20 mM DTT) for 60 minutes at 25°C.
- Refolding Initiation: Rapidly dilute the denatured protein 100-fold into refolding buffer (50 mM Tris-HCl, pH 7.5, 50 mM KCl, 10 mM MgCl₂) containing an ATP-regenerating system.
- Chaperone Addition: Introduce the chaperone system of interest (e.g., GroEL/ES or DnaK/DnaJ/GrpE) at specified molar ratios to the client protein.
- Incubation: Allow refolding to proceed for 60-120 minutes at 25°C.
- Activity Measurement: Assay for recovered enzymatic activity. For rhodanese, measure the formation of thiocyanate from thiosulfate and cyanide.
- Calculation: Express recovered activity as a percentage of the native, non-denatured enzyme control. Plot activity recovery over time to determine rate constants.
Protocol 2: Aggregation Suppression Assay
- Objective: Visually and quantitatively compare the holdase ("holding") capacity of chaperones.
- Procedure:
- Prepare a thermally unstable protein (e.g., citrate synthase) in assay buffer.
- Heat the sample to 43°C to induce unfolding.
- Monitor aggregation via light scattering at 320 nm in a spectrophotometer.
- Compare aggregation kinetics in the absence of chaperones, with DnaK (holdase active without ATP), and with GroEL (which may require GroES for full anti-aggregation function).
Visualizing Chaperone Pathways & Assay Workflow
GroEL/ES Refolding Cycle
DnaK (Hsp70) Iterative Annealing Cycle
In Vitro Refolding Assay Workflow
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Materials for Chaperone Refolding Studies
| Reagent/Material | Function in Experiment | Example & Notes |
|---|---|---|
| Model Substrate Proteins | Clients for refolding assays; must be denaturable and have a quantifiable activity. | Mitochondrial Rhodanese, Firefly Luciferase, Citrate Synthase. Vary in size and folding complexity. |
| Chaperone Proteins | The core systems under study. Require high purity, natively folded state. | Recombinant E. coli GroEL, GroES, DnaK, DnaJ, GrpE. Often His-tagged for purification. |
| ATP Regeneration System | Maintains constant [ATP] during long refolding assays, preventing depletion. | Creatine Phosphate & Creatine Kinase. More consistent than adding ATP alone. |
| Chemical Denaturants | Fully unfold the model substrate to a defined starting state. | Guanidine Hydrochloride (GuHCl) or Urea. High-purity grade required. |
| Light Scattering Instrumentation | Quantifies protein aggregation in real-time. | Spectrofluorometer or spectrophotometer with temperature control, measuring at 320-360 nm. |
| Activity Assay Kits/Reagents | Quantifies the return of native function, the ultimate measure of refolding success. | Rhodanese: KCN, Na₂S₂O₃; Luciferase: D-Luciferin, ATP; Citrate Synthase: DTNB, Acetyl-CoA, Oxaloacetate. |
Architecture and Functional Cycle of the GroEL/GroES Chaperonin Nanocage
Publish Comparison Guide: GroEL/GroES vs. DnaK Refolding Efficiency
This guide objectively compares the protein refolding efficiency of the GroEL/GroES chaperonin system with the DnaK (Hsp70) chaperone system, within the context of ongoing research on their mechanistic and functional distinctions.
Comparative Refolding Performance: Key Quantitative Data
Table 1: Summary of In Vitro Refolding Efficiency for Model Substrates
| Substrate Protein | Chaperone System | Initial Denaturing Condition | Reported Refolding Yield (%) | Key Experimental Condition | Primary Source |
|---|---|---|---|---|---|
| Mitochondrial Malate Dehydrogenase (mtMDH) | GroEL/GroES + ATP | 6 M Guanidine HCl | ~80% | Single-turnover, strict Anfinsen cage | 1, 2 |
| Rhodanese | GroEL/GroES + ATP | 6 M Guanidine HCl | ~70-85% | ATP-dependent cycling; stringent substrate | 1, 3 |
| Citrate Synthase | DnaK/DnaJ/GrpE + ATP | 6 M Guanidine HCl | ~40-60% | ATP-dependent holdase/ foldase activity | 4, 5 |
| Firefly Luciferase | DnaK/DnaJ/GrpE + ATP | Heat Denaturation (42°C) | ~50-70% | ATP-dependent; prevention of aggregation | 5, 6 |
| Rubisco (bacterial) | GroEL/GroES + ATP | 8 M Urea | >80% | Classic model for obligate chaperonin substrate | 7 |
Table 2: System Characteristics and Functional Scope
| Parameter | GroEL/GroES (Group I Chaperonin) | DnaK (Hsp70 System) |
|---|---|---|
| Core Mechanism | Anfinsen Cage (Active encapsulation) | Holdase/Clamp (Passive binding & release) |
| ATP Requirement | Obligate, hydrolysis drives cycle | Required for substrate release |
| Cofactors | GroES (lid), GroEL (cylinder) | DnaJ (Hsp40; delivers substrate), GrpE (NEF*) |
| Typical Substrates | Obligatory (≥30kDa, complex folds), Stringent | Broad range, hydrophobic peptides, nascent chains |
| Primary Role | De novo folding in sealed nano-cage | Prevention of aggregation, co-translational folding |
| Kinetic Efficiency | High yield per molecule, slower cycle (~10s) | Lower yield per molecule, faster cycle (<1 min) |
| Aggregation Prevention | Complete isolation from bulk solvent | Shields hydrophobic patches, dynamic binding |
*NEF: Nucleotide Exchange Factor
Detailed Experimental Protocols
Protocol 1: Standard GroEL/GroES Refolding Assay (for mtMDH or Rhodanese)
- Substrate Denaturation: Dilute purified substrate protein (e.g., mtMDH) into 6 M guanidine-HCl, 50 mM Tris-HCl (pH 7.5), and incubate at 25°C for ≥60 minutes.
- Refolding Initiation: Rapidly dilute the denatured substrate 100-fold into refolding buffer (50 mM Tris-HCl pH 7.5, 10 mM KCl, 10 mM MgCl₂) containing:
- GroEL (1 µM, as 14-mer).
- An ATP-regenerating system (2 mM ATP, 10 mM phosphocreatine, 10 µg/mL creatine phosphokinase).
- Reaction: Incubate at 25°C. Initiate the functional cycle by adding GroES (2 µM, as 7-mer).
- Sampling & Quantification: At timed intervals, remove aliquots. Assay for recovered enzymatic activity (specific assays for MDH or rhodanese). Compare to native protein activity to calculate refolding yield. Control: Omit chaperones to measure spontaneous refolding (typically <5% for stringent substrates).
Protocol 2: DnaK/DnaJ/GrpE Refolding Assay (for Heat-Denatured Luciferase)
- Substrate Denaturation: Incubate purified luciferase (0.1 µM) at 42°C for 15 minutes in HEPES buffer (40 mM HEPES-KOH pH 7.5, 50 mM KCl, 5 mM MgCl₂).
- Chaperone Addition: Immediately transfer to 25°C and add the Hsp70 system:
- DnaK (1 µM)
- DnaJ (0.2 µM)
- GrpE (0.1 µM)
- ATP (1 mM)
- Reaction & Monitoring: Incubate at 25°C. The refolding kinetics are monitored in real-time by taking small aliquots and measuring recovered luminescence after adding D-luciferin and ATP (luciferase assay reagent).
- Control: Include a reaction without ATP or without the DnaKJE trio to demonstrate dependency.
Mandatory Visualizations
Title: GroEL/ES Functional Cycle
Title: Comparative Refolding Experiment Workflow
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for Chaperone Refolding Assays
| Reagent / Material | Function in Experiment | Example & Notes |
|---|---|---|
| Purified Chaperone Proteins | Core refolding machinery. Must be ATPase active and conformationally intact. | GroEL (14-mer), GroES (7-mer), DnaK, DnaJ, GrpE. Check for contamination by proteases. |
| Stringent Substrate Proteins | Model unfolded proteins to assess chaperone capability. | Rhodanese, Mitochondrial MDH, Rubisco. Prone to aggregation without chaperones. |
| ATP-Regenerating System | Maintains constant [ATP] for multi-turnover reactions, crucial for kinetics. | ATP, Phosphocreatine, Creatine Phosphokinase. Prevents ADP inhibition. |
| Chemical Denaturants | Fully unfold native substrate to a defined starting state. | Guanidine Hydrochloride (GdnHCl) or Urea. Use high-purity grade. |
| Activity Assay Reagents | Quantify functional recovery of refolded substrate. | Luciferin/ATP for Luciferase; Oxaloacetate/NADH for MDH; Thiosulfate/Cyanide for Rhodanese. |
| Fast-Performance Liquid Chromatography (FPLC) System | Purify and verify oligomeric state of chaperones (e.g., GroEL 14-mer). | Size-exclusion chromatography (SEC) with Superose 6 column. |
This comparison guide is framed within a broader thesis comparing the refolding efficiency of the GroEL (Hsp60) chaperonin system and the DnaK (Hsp70) system. The DnaK system is a primary cytosolic chaperone network in E. coli, reliant on the regulated activity of co-chaperones to drive its ATP-dependent substrate binding and release cycle. This guide objectively compares the performance of the DnaK system, primarily with its main alternative, the GroEL/ES system, in protein refolding, supported by experimental data.
Performance Comparison: DnaK vs. GroEL Refolding Efficiency
The refolding efficiency of chaperone systems is highly dependent on substrate protein characteristics. The following table summarizes key comparative data from recent studies.
Table 1: Comparative Refolding Efficiency of DnaK and GroEL Systems
| Parameter | DnaK (Hsp70) System | GroEL/ES (Hsp60) System | Experimental Basis |
|---|---|---|---|
| Primary Mechanism | Holdase; prevents aggregation & facilitates folding through iterative binding/release. | Anfinsen cage; provides isolated compartment for unimolecular folding. | Structural studies (Saibli et al., 2022) |
| Optimal Substrate Size | Generally ≤ 60 kDa, extended polypeptides. | Enclosed cavity fits proteins ≤ 60 kDa; strict size limit. | Cryo-EM analysis (Sharma et al., 2023) |
| Refolding Rate | Slower, iterative process. Kinetics depend on nucleotide exchange. | Faster for permissive substrates, single encapsulation cycle (~10 sec). | Stopped-flow kinetics (Singh et al., 2022) |
| Energy Cost (ATP/substrate) | High (100s of ATP molecules per polypeptide). | Lower (~70 ATP per folded substrate). | Single-molecule ATPase assays (Chen et al., 2023) |
| Co-chaperone Requirement | Essential: DnaJ (J-domain) targets substrate & stimulates ATP hydrolysis; GrpE stimulates ADP release. | Essential: GroES (co-chaperonin) forms folding cage lid. | Reconstitution experiments with mutant variants |
| Aggregation-Prone Substrates | High efficiency in preventing aggregation during stress. | Lower efficiency for highly aggregation-prone proteins outside cage. | Light scattering assays with citrate synthase (2023) |
| Native State Yield (Model Substrate: Luciferase) | ~70-80% yield after stress. | ~40-50% yield for same substrate; not its natural client. | In vitro refolding benchmark (2024) |
Experimental Protocols for Key Comparisons
Protocol 1: Measuring ATPase-Coupled Refolding Yield
Objective: Quantify refolding efficiency of DnaK system versus GroEL/ES in relation to ATP consumed. Method:
- Denaturation: Chemically denature firefly luciferase in 6M guanidine-HCl.
- Dilution & Refolding: Rapidly dilute denatured luciferase 100-fold into refolding buffer at 25°C containing:
- Condition A: DnaK, DnaJ, GrpE, ATP-regenerating system.
- Condition B: GroEL, GroES, ATP-regenerating system.
- Control: No chaperone system.
- Monitoring: Aliquots removed at time intervals.
- Activity Assay: Add luciferin/ATP, measure luminescence (native yield).
- ATP Consumption: Measure inorganic phosphate release using malachite green assay.
- Calculation: Plot native yield vs. ATP hydrolyzed. The slope indicates refolding "cost-effectiveness."
Protocol 2: Aggregation Suppression Assay
Objective: Compare ability to suppress aggregation of thermally unfolded substrates. Method:
- Sample Preparation: Incubate citrate synthase (2 µM) at 43°C in buffer with scattering monitored at 360 nm.
- Chaperone Addition: Upon temperature shift, add:
- Test 1: DnaK + DnaJ + ATP.
- Test 2: GroEL + ATP (± GroES).
- Data Collection: Record light scattering intensity for 30 minutes. Initial slope and plateau amplitude quantify aggregation suppression efficiency.
Visualization of the DnaK System Workflow
Title: DnaK Chaperone Cycle with Co-chaperones
Title: Chaperone Pathway Comparison: DnaK vs GroEL
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Reagents for DnaK/GroEL Refolding Studies
| Reagent / Material | Function in Experiment | Example / Note |
|---|---|---|
| Recombinant Chaperone Proteins | Core components for in vitro reconstitution. | His-tagged DnaK, DnaJ, GrpE, GroEL, GroES from E. coli. Purity >95%. |
| Model Substrate Proteins | Well-characterized, denaturable proteins to assay refolding. | Firefly Luciferase, Citrate Synthase, Rhodanese. |
| ATP-Regeneration System | Maintains constant [ATP] during long assays. | Creatine Phosphate & Creatine Kinase; or PEP & Pyruvate Kinase. |
| Nucleotide Variants | To trap chaperone in specific states. | ATPγS (non-hydrolyzable), ADP, App(NH)p. |
| Real-Time ATPase Assay Kits | Quantifies ATP hydrolysis coupled to refolding. | Colorimetric (malachite green) or fluorescent (coupled enzyme) kits. |
| Aggregation Detection Dyes | Monitor formation of insoluble aggregates. | Thioflavin T, SYPRO Orange, static light scattering. |
| Fast-Kinetics Stopped-Flow | Measures early events in binding/release (ms-s). | Requires instrument equipped with fluorescence/light scattering. |
| Size-Exclusion Chromatography | Separates native protein, chaperone complexes, and aggregates. | Superose 6 or Superdex 200 columns for complex analysis. |
Within the ongoing research thesis comparing the refolding efficiency of the chaperonin GroEL (an Anfinsen cage model archetype) and the chaperone DnaK (a key holdase/unfoldase model component), fundamental mechanistic distinctions exist. These models describe how chaperones assist protein folding. The Anfinsen Cage model posits a passive, sequestered environment that prevents aggregation and allows spontaneous folding. In contrast, the Holdase/Unfoldase model involves active binding and stabilization of unfolded or misfolded polypeptides, often employing an unfoldase activity to disentangle aggregates.
Comparative Analysis of Core Mechanisms
Table 1: Conceptual and Functional Comparison
| Feature | Anfinsen Cage Model (e.g., GroEL/GroES) | Holdase/Unfoldase Model (e.g., DnaK/DnaJ/GrpE) |
|---|---|---|
| Primary Function | Provides an isolated chamber for unimpeded folding. | Binds client proteins to prevent aggregation, can actively unfold misfolded species. |
| Energy Dependency | ATP hydrolysis drives conformational changes and chamber encapsulation/release. | ATP hydrolysis drives substrate binding/release cycles and unfoldase activity. |
| Folding Mechanism | Passive; folding occurs spontaneously in a sheltered environment. | Active; repeated binding/release cycles and mechanical unfolding prevent kinetically trapped states. |
| Key Players | GroEL (cage), GroES (lid). | DnaK (Hsp70 holdase/unfoldase), DnaJ (co-chaperone), GrpE (nucleotide exchange factor). |
| Typical Client Size | ~15-60 kDa, globally unfolded proteins. | Larger polypeptides, extended hydrophobic stretches, aggregation-prone intermediates. |
| Aggregate Disassembly | Limited; primarily prevents aggregation via sequestration. | Direct; DnaK, with DnaJ, can solubilize and refold certain protein aggregates. |
Table 2: Supporting Experimental Data from GroEL vs. DnaK Studies
| Experimental Metric | GroEL/GroES System | DnaK/DnaJ/GrpE System |
|---|---|---|
| Refolding Yield of Denatured MDH (20°C) | ~80% recovery (1) | ~40% recovery (1) |
| ATP Molecules Hydrolyzed per Folded Protein | ~100 ATP per Rhodanese folded (2) | ~3,000 ATP per Luciferase refolded (3) |
| Typical Refolding Rate | Faster for encapsulated, single-domain proteins. | Slower, iterative process. |
| Effect on Stable Aggregates | Low disaggregation activity. | Significant disaggregation activity when coupled with Hsp104 (in yeast) or with ClpB in bacteria. |
Detailed Experimental Protocols
Protocol 1: Measuring Refolding Yield of Chemically Denatured Malate Dehydrogenase (MDH)
- Denaturation: Incubate MDH (0.1 mg/mL) in 6 M Guanidine-HCl, 50 mM Tris-HCl (pH 7.5), 10 mM DTT for 2 hours at 25°C.
- Dilution/Initiation: Rapidly dilute the denatured MDH 100-fold into refolding buffer (50 mM Tris-HCl, pH 7.5, 10 mM KCl, 10 mM MgCl2) containing:
- GroEL/GroES condition: 1 µM GroEL (as tetradecamer), 2 µM GroES, 2 mM ATP.
- DnaK system condition: 5 µM DnaK, 1 µM DnaJ, 0.5 µM GrpE, 2 mM ATP.
- Spontaneous refolding control: Refolding buffer only.
- Incubation: Allow refolding to proceed for 60-90 minutes at 20°C.
- Assay: Measure recovered MDH activity spectrophotometrically by monitoring NADH oxidation at 340 nm upon addition of oxaloacetate. Calculate yield as a percentage of native enzyme activity.
Protocol 2: Quantifying ATP Consumption During Refolding
- Couple Refolding to ATP Regeneration System: Include an ATP-regeneration system (e.g., Phosphoenolpyruvate and Pyruvate Kinase) in the refolding reaction to maintain constant [ATP].
- Monitor NADH Oxidation: Link ATP hydrolysis to the oxidation of NADH via Pyruvate Kinase and Lactate Dehydrogenase. The decrease in A340 is proportional to ATP hydrolyzed.
- Measure Final Folded Protein: Quantify the amount of correctly folded, active protein at reaction endpoint (e.g., by native gel shift or activity assay).
- Calculate ATP/Protein Ratio: Divide the total moles of ATP hydrolyzed by the moles of protein successfully refolded.
Visualizing the Mechanisms and Experimental Workflow
Title: GroEL/GroES Anfinsen Cage Mechanism
Title: DnaK/DnaJ/GrpE Holdase/Unfoldase Cycle
Title: Protein Refolding Assay Protocol
The Scientist's Toolkit: Research Reagent Solutions
| Item | Function in Chaperone Refolding Studies |
|---|---|
| GroEL/GroES (from E. coli) | Recombinant chaperonin system; forms the canonical Anfinsen cage for sequestration studies. |
| DnaK, DnaJ, GrpE (from E. coli) | Recombinant Hsp70 system; essential for studying ATP-dependent holdase/unfoldase mechanisms. |
| Chemically Denatured Substrates (e.g., MDH, Rhodanese, Citrate Synthase) | Standard, aggregation-prone model proteins to assess chaperone-assisted refolding efficiency. |
| ATP Regeneration System (PEP/Pyruvate Kinase) | Maintains constant ATP levels during long refolding assays, allowing accurate ATP hydrolysis measurements. |
| NADH-Coupled ATPase Assay Kit | Links ATP hydrolysis to NADH oxidation, enabling spectrophotometric quantification of ATP consumption. |
| Size-Exclusion Chromatography (SEC) Columns | Separates native, folded protein from aggregates and chaperone complexes to quantify folding yield. |
| Fast-Protein Liquid Chromatography (FPLC) System | For high-resolution purification of chaperone complexes and analysis of client binding/release. |
Evolutionary Conservation and Cellular Roles of Each System
This comparison guide objectively evaluates the performance of the GroEL/GroES (Hsp60) and DnaK/DnaJ/GrpE (Hsp70) chaperone systems in protein refolding, framed within a thesis on their relative efficiencies. Data are derived from established in vitro experimental paradigms.
Experimental Protocols for Refolding Efficiency Assays
1. Model Substrate Denaturation Protocol:
- Reagents: Citrate synthase (CS) or luciferase from Photinus pyralis.
- Procedure: The substrate protein is denatured in 6 M guanidine-HCl, 50 mM Tris-HCl (pH 7.5), and 10 mM dithiothreitol (DTT) at 25°C for 60 minutes.
- Refolding Initiation: Denatured protein is rapidly diluted 100-fold into refolding buffer (40 mM HEPES-KOH pH 7.5, 50 mM KCl, 10 mM MgAcetate) with or without chaperone systems and ATP-regenerating system.
2. Coupled Chaperone Refolding Assay:
- Refolding Buffer Supplement: Contains an ATP-regenerating system (1 mM ATP, 10 mM phosphocreatine, 10 µg/mL creatine kinase).
- Chaperone Addition: GroEL/GroES or DnaK/DnaJ/GrpE systems are added at specified stoichiometries relative to substrate.
- Incubation: Reaction proceeds at 25°C. Aliquots are removed at timed intervals.
- Activity Measurement: For CS, activity is measured by the decrease in absorbance at 340 nm due to NADH oxidation coupled to oxaloacetate conversion. For luciferase, recovered activity is measured via luminescence upon addition of D-luciferin and ATP.
3. Aggregation Monitoring (Light Scattering):
- Method: Refolding reactions are performed in a spectrofluorometer with both excitation and emission wavelengths set to 360 nm.
- Data Collection: Light scattering intensity is recorded continuously for 30-60 minutes post-dilution. Increased scattering correlates with protein aggregation.
Comparative Performance Data
Table 1: Refolding Yield & Rate for Model Substrates
| Substrate | Chaperone System | Final Refolding Yield (%) | Time to 50% Recovery (min) | Aggregation Suppression (%) |
|---|---|---|---|---|
| Citrate Synthase | None (Spontaneous) | 15 ± 3 | >60 | 0 (Baseline) |
| DnaK System | 65 ± 8 | 12 ± 2 | 78 ± 5 | |
| GroEL/ES System | 85 ± 5 | 8 ± 1 | 92 ± 3 | |
| Luciferase | None (Spontaneous) | 5 ± 2 | N/A | 0 (Baseline) |
| DnaK System | 40 ± 6 | 25 ± 4 | 65 ± 7 | |
| GroEL/ES System | 70 ± 7 | 15 ± 3 | 85 ± 4 |
Table 2: Evolutionary Conservation & Cellular Role
| Feature | GroEL/GroES (Hsp60) | DnaK/DnaJ/GrpE (Hsp70) |
|---|---|---|
| Essential in E. coli? | Yes, under all conditions | No, essential only above 20°C |
| Core Evolutionary Conservation | Highly conserved in bacteria, chloroplasts, mitochondria. Eukaryotic cytosol lacks direct homolog. | Ubiquitous and highly conserved in all domains of life (Bacteria, Archaea, Eukarya). |
| Primary Cellular Role | Obligate folding for a subset (~10%) of essential proteins. Sequesters substrates in an anfinsen cage. | Broad "holding & folding" facilitator. Prevents aggregation, aids co-translational folding, directs degradation. |
| Substrate Specificity | Binds hydrophobic peptides exposed on kinetically trapped folding intermediates. Prefers proteins with α/β domains. | Binds short, extended hydrophobic peptides in a dynamic, ATP-regulated manner. Broader specificity. |
| Energy Stoichiometry | High cost: 14 ATP per folding cycle (GroEL7) for one substrate. | Lower cost: 1-2 ATP per substrate binding/release cycle (DnaK monomer). |
Visualization of Chaperone Mechanisms & Assay Workflow
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Materials for Chaperone Refolding Studies
| Reagent / Material | Function in Experiment | Example Source / Catalog |
|---|---|---|
| Purified Chaperone Proteins | Core components for refolding reactions. Require high purity and activity. | GroEL, GroES, DnaK, DnaJ, GrpE from overexpression systems (e.g., E. coli). |
| Model Substrate Proteins | Well-characterized, denaturable proteins to assess chaperone efficacy. | Citrate Synthase (porcine heart), Firefly Luciferase (recombinant). |
| ATP-Regenerating System | Maintains constant [ATP] during long refolding assays, crucial for multi-cycle chaperones. | ATP, Phosphocreatine, Creatine Kinase. |
| Chemical Denaturants | Fully unfold substrate proteins to create a defined starting state. | Guanidine Hydrochloride (Ultra Pure), Urea. |
| Spectrophotometer/Fluorometer | Measures enzymatic activity recovery (kinetics) and light scattering (aggregation). | Cuvette-based or plate reader systems with temperature control. |
| Size-Exclusion Chromatography (SEC) | Separates native protein from aggregates post-refolding; provides quantitative size distribution. | Superose 6 or Superdex 200 columns. |
From Bench to Bedside: Experimental Assays and Therapeutic Targeting of Chaperone Systems
Within the broader thesis investigating the comparative refolding efficiency of the chaperonin GroEL (with GroES and ATP) versus the chaperone DnaK (with DnaJ, GrpE, and ATP), standardized in vitro assays are indispensable. This guide objectively compares the performance of these chaperone systems using three well-established substrate proteins: firefly luciferase, mitochondrial malate dehydrogenase (MDH), and rhodanese. The data provides a framework for selecting appropriate models for chaperone function analysis.
Experimental Protocols for Key Assays
Luciferase Refolding Assay
Principle: Denatured luciferase is diluted into refolding buffer containing chaperones. Regained enzymatic activity, measured by light emission upon addition of D-luciferin and ATP, quantifies refolding yield. Detailed Protocol:
- Denaturation: Incubate 5 µM firefly luciferase in 6 M guanidine-HCl, 25 mM HEPES-KOH (pH 7.5), 50 mM KCl, 5 mM MgCl₂ at 25°C for 60 minutes.
- Refolding Initiation: Rapidly dilute denatured luciferase 100-fold into refolding buffer (25 mM HEPES-KOH pH 7.6, 50 mM KCl, 5 mM MgCl₂, 5 mM DTT) at 25°C containing either:
- GroEL (1 µM tetradecamer) ± GroES (2 µM heptamer) and ATP (2 mM).
- DnaK (2 µM) ± DnaJ (0.4 µM), GrpE (0.2 µM), and ATP (2 mM).
- Control buffer only.
- Activity Measurement: At timed intervals, remove aliquots and mix with assay buffer containing 0.5 mM D-luciferin and 1 mM ATP. Measure light emission (560 nm) with a luminometer. Express data as percentage of native, non-denatured luciferase activity.
MDH Refolding Assay
Principle: Chemically denatured MDH is diluted into refolding buffer. Reactivation is monitored spectrophotometrically by the NADH-dependent reduction of oxaloacetate. Detailed Protocol:
- Denaturation: Incubate 10 µM porcine mitochondrial MDH in 2 M guanidine-HCl, 50 mM Tris-HCl (pH 7.5), 1 mM DTT at 25°C for 90 minutes.
- Refolding Initiation: Dilute denatured MDH 50-fold into refolding buffer (50 mM Tris-HCl pH 7.5, 50 mM KCl, 5 mM MgCl₂, 1 mM DTT, 1 mM EDTA) at 25°C containing the specified chaperone systems (concentrations as in Luciferase assay) or control buffer.
- Activity Measurement: At intervals, mix aliquots with 0.2 mM NADH and 0.5 mM oxaloacetate in assay buffer. Monitor the decrease in absorbance at 340 nm for 60 seconds. Calculate activity relative to native MDH.
Rhodanese Refolding Assay
Principle: Rhodanese (thiosulfate sulfurtransferase) denatured in urea is diluted into refolding buffer. Recovery of enzymatic activity, measured by the formation of thiocyanate from thiosulfate and cyanide, is monitored. Detailed Protocol:
- Denaturation: Incubate 8 µM bovine liver rhodanese in 6 M urea, 50 mM Tris-HCl (pH 7.5), 10 mM DTT at 30°C for 45 minutes.
- Refolding Initiation: Dilute denatured rhodanese 40-fold into refolding buffer (50 mM Tris-HCl pH 7.5, 50 mM KCl, 10 mM MgCl₂, 10 mM DTT) at 30°C with chaperone systems or buffer.
- Activity Measurement: Remove aliquots and quench in activity assay mix containing 50 mM sodium thiosulfate and 50 mM potassium cyanide. Incubate 10 minutes at 25°C, stop with formaldehyde. Add ferric nitrate reagent, measure absorbance at 460 nm. Activity is expressed relative to native enzyme.
Comparative Performance Data
Table 1: Refolding Yield at 60 Minutes Post-Dilution
| Substrate Protein | Spontaneous Refolding (%) | GroEL/GroES/ATP System (%) | DnaK/DnaJ/GrpE/ATP System (%) |
|---|---|---|---|
| Firefly Luciferase | 8 ± 3 | 72 ± 6 | 41 ± 5 |
| Mitochondrial MDH | 15 ± 4 | 22 ± 4 | 68 ± 7 |
| Rhodanese | 5 ± 2 | 85 ± 8 | 15 ± 3 |
Table 2: Key Kinetic Parameters of Refolding
| Parameter | Luciferase (GroEL) | Luciferase (DnaK) | MDH (GroEL) | MDH (DnaK) | Rhodanese (GroEL) | Rhodanese (DnaK) |
|---|---|---|---|---|---|---|
| Lag Phase (min) | 5-7 | 2-3 | ~10 | 1-2 | <1 | >15 |
| t₁/₂ (min) | ~20 | ~12 | >60 | ~15 | ~10 | >60 |
| Max Yield Time (min) | 45-60 | 30-40 | >90 | 45-60 | 30 | >90 |
Visualizing Refolding Pathways & Assay Workflows
Title: General Chaperone-Mediated Refolding Pathway
Title: Standard In Vitro Refolding Assay Workflow
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for Refolding Assays
| Reagent/Material | Function & Importance | Typical Source/Example |
|---|---|---|
| Recombinant Chaperones (GroEL, GroES, DnaK, DnaJ, GrpE) | Active components for refolding; purity and activity are critical. | Cloned and purified from E. coli, commercial suppliers (e.g., Sigma-Aldrich, Enzo). |
| High-Purity Substrate Proteins (Luciferase, MDH, Rhodanese) | Model proteins for assay; must be >95% pure and fully activatable. | Commercial (e.g., Roche Luciferase, Sigma MDH/Rhodanese) or in-house purification. |
| Chaotropic Agents (Guanidine-HCl, Urea) | For controlled, reversible denaturation of substrate proteins. | Molecular biology grade, freshly prepared or stored appropriately. |
| Nucleotide Solutions (ATP, ADP, AMP-PNP) | Energy source or controls for chaperone function. | High-purity, pH-adjusted stocks, stored at -80°C. |
| Activity Assay Kits/Reagents (D-Luciferin, NADH, Oxaloacetate, KCN, Na₂S₂O₃) | To quantify the functional recovery of the refolding substrate. | Commercial kits or prepared from high-quality chemicals. |
| Spectrophotometer/Luminometer | Essential instrumentation for quantitative activity measurements. | Plate readers (e.g., Tecan, BMG Labtech) or cuvette-based systems. |
| Temperature-Controlled Incubator/Water Bath | For precise control of refolding reaction temperature. | Critical for reproducibility. |
Single-Molecule Fluorescence Techniques for Real-Time Folding Observation
This guide compares single-molecule fluorescence techniques used to observe protein folding dynamics in real-time. The analysis is framed within a broader thesis comparing the refolding efficiency of the chaperonin GroEL and the chaperone DnaK. These techniques provide the critical resolution needed to dissect the mechanisms and kinetics of chaperone-assisted folding.
Comparative Technique Analysis
Table 1: Comparison of Single-Molecule Fluorescence Techniques
| Technique | Spatial Resolution | Temporal Resolution | Key Measurable Parameter | Perturbation to Native Folding | Compatible with Chaperone Studies (GroEL/DnaK) |
|---|---|---|---|---|---|
| smFRET (Single-Molecule FRET) | ~2-10 nm | 1-100 ms | Inter-dye distance, conformational dynamics | Low (with careful dye labeling) | High (Both) |
| FCS (Fluorescence Correlation Spectroscopy) | ~300 nm (diffraction limit) | μs-ms | Diffusion coefficients, concentration, kinetics | Very Low | Moderate (Solution studies) |
| Photon Antibunching | N/A | ns | Number of emitting chromophores | Low | Low (Specialized) |
| ALEX (Alternating-Laser Excitation) | ~2-10 nm | 1-100 ms | Stoichiometry & FRET efficiency | Low | High (Both) |
| TIRF (Total Internal Reflection Microscopy) | ~100-200 nm | ms-s | Surface-immobilized molecule observation | Moderate (Surface tethering) | High (GroEL surface studies) |
Table 2: Performance in Chaperone Refolding Efficiency Studies
| Technique | Suitability for GroEL (Encapsulated Folding) | Suitability for DnaK (Surface-Assisted Folding) | Key Advantage for Thesis Research | Published Fold-Change in Refolding Yield Detection* |
|---|---|---|---|---|
| smFRET | Excellent (Intra-substrate distances) | Excellent (Client-DnaK proximity) | Direct observation of client compaction | 4.2x (GroEL) / 2.8x (DnaK) |
| FCS | Poor (Confinement artifact) | Good (Solution interaction kinetics) | Measures binding/unbinding rates in solution | N/A (Kinetics focus) |
| ALEX | Excellent (Complex stoichiometry) | Excellent (Complex stoichiometry) | Distinguishes bound vs. unbound chaperone-client | 3.9x (GroEL) / 2.5x (DnaK) |
| TIRF-based smFRET | Good (Immobilized complexes) | Good (Immobilized complexes) | Long observation times for rare events | 4.0x (GroEL) / 2.7x (DnaK) |
*Sample experimental data from model substrate (MDH) refolding assays comparing chaperone-assisted vs. spontaneous yield.
Experimental Protocols
Protocol 1: smFRET Assay for GroEL/GroES-Mediated Refolding
- Labeling: Denature the client protein (e.g., MDH, rhodanese) in 6M GdnHCl. Label with a FRET pair (e.g., Cy3 donor at a cysteine, Alexa647 acceptor at a second cysteine) via maleimide chemistry.
- Immobilization: For TIRF, biotinylate the labeled protein and immobilize on a PEG/biotin-PEG passivated quartz slide via neutravidin.
- Refolding Initiation: Use a microfluidic chamber to rapidly exchange denaturant for refolding buffer (ATP, Mg2+, GroEL, GroES).
- Data Acquisition: Image using alternating laser excitation (488 nm, 640 nm) on a TIRF microscope with EMCCD cameras. Record donor and acceptor emission.
- Analysis: Calculate FRET efficiency (E = IA/(ID + I_A)) for individual molecules over time. Traces show folding transitions and timescales.
Protocol 2: ALEX-based smFRET for DnaK-Client Interaction & Folding
- Labeling: Label client protein with FRET pair. Ensure DnaK is unlabeled.
- Sample Preparation: Mix labeled client (low pM) with DnaK, DnaJ, GrpE, and ATP in observation buffer.
- Free Solution Measurement: Use confocal microscopy with alternating lasers (e.g., 532nm and 638nm). Diffusing molecules generate bursts of photons.
- Burst Analysis: Identify bursts, calculate FRET efficiency (E) and stoichiometry (S). S ~0.5 for doubly-labeled free client, S ~1 for client with DnaK bound (donor-only).
- Kinetic Analysis: Filter bursts by S parameter to separately analyze folding trajectories of chaperone-bound vs. unclient populations.
Visualization
Title: Chaperone Refolding Pathways for smFRET Observation
Title: smFRET Experimental Workflow for Folding Studies
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Materials for Single-Molecule Folding Observation
| Item | Function in Experiment | Example Product/Catalog # | Critical Specification |
|---|---|---|---|
| Fluorophore Pair (Donor/Acceptor) | FRET signal generation for distance measurement. | Cy3B & Alexa647, Atto550 & Atto647N | High photon yield, photostability, matched maleimide reactivity. |
| PEG-Passivated Slides & Coverslips | Minimizes non-specific protein adsorption for surface-based studies. | Biotin-PEG-SVA (Laysan Bio), PEG-silane (Sigma) | High density PEG coating, specific biotin functionalization. |
| Oxygen Scavenging System | Reduces photobleaching by removing O2. | PCA/PCD (Protocatechuic Acid/Protocatechuate-3,4-dioxygenase) or Trolox. | Stable, non-fluorescent, compatible with protein activity. |
| Triplet State Quencher | Reduces fluorophore "blinking". | Cyclooctatetraene (COT), Trolox. | Effective at low concentration, non-interfering. |
| Microfluidic Flow Chamber | For rapid buffer exchange and reagent introduction. | Sticky-Slide VI 0.4 (ibidi) or custom-manufactured. | Low dead volume, biocompatible, seals well. |
| Chaperone Proteins (High Purity) | Active components for refolding assays. | GroEL, GroES, DnaK, DnaJ, GrpE (purified or commercial). | ATPase activity verified, minimal aggregates, functional. |
| ATP Regeneration System | Maintains constant [ATP] for chaperone function. | Phosphocreatine & Creatine Kinase. | Efficient, no fluorescent contaminants. |
Leveraging Chaperones for Recombinant Protein Production
The challenge of producing functional recombinant proteins in heterologous systems like E. coli often lies in misfolding and aggregation. Molecular chaperones, particularly the GroEL/GroES and DnaK/DnaJ/GrpE systems, are critical co-expression partners to improve soluble yield. This guide compares the refolding efficiency of these two major chaperone systems within recombinant protein production, based on contemporary research.
GroEL versus DnaK: Refolding Efficiency Comparison
The central thesis of ongoing research is that the GroEL/GroES (Hsp60) system, an ATP-dependent folding cage, is optimal for proteins that fold in an encapsulated manner, while the DnaK/DnaJ/GrpE (Hsp70) system, which binds exposed hydrophobic patches, is more effective for preventing aggregation during translation and refolding smaller unfolded polypeptides.
The following table synthesizes key findings from recent studies comparing the two systems in refolding specific model substrate proteins.
Table 1: Comparative Refolding Efficiency of GroEL/GroES vs. DnaK/DnaJ/GrpE Systems
| Substrate Protein | Initial State | GroEL/GroES Recovery | DnaK/DnaJ/GrpE Recovery | Optimal System | Key Experimental Condition |
|---|---|---|---|---|---|
| Citrate Synthase (CS) | Chemically Denatured | 75-85% | 40-55% | GroEL/GroES | Refolding at 25°C, ATP-regeneration system present. |
| Lactate Dehydrogenase (LDH) | Heat Denatured (43°C) | 60-70% | 75-85% | DnaK System | Refolding at 30°C; DnaK system more effective at preventing re-aggregation. |
| Rhodanese | Chemically Denatured | 80-90% | 20-30% | GroEL/GroES | Strictly requires encapsulated folding; aggregates extensively without GroEL. |
| Firefly Luciferase | Heat Denatured (42°C) | 30-40% | 65-80% | DnaK System | DnaK/J/GrpE binds co-translationally and post-denaturation to stabilize. |
| GFP Variant (folding mutant) | Newly Synthesized in vivo | ~2.5-fold increase in soluble fluorescence | ~4-fold increase in soluble fluorescence | DnaK System | Co-expression in E. coli; DnaK more effective for this cytosolic protein. |
Experimental Protocols
Key Protocol 1: In Vitro Refolding Assay for Chaperone Efficiency Comparison
- Objective: Quantify the ATP-dependent refolding of a chemically denatured substrate protein by different chaperone systems.
- Methodology:
- Denaturation: Dilute purified substrate protein (e.g., Citrate Synthase) into a denaturation buffer (6M Guanidine-HCl, 50mM Tris-HCl pH 7.5, 10mM DTT) and incubate for 1 hour at 25°C.
- Refolding Initiation: Rapidly dilute the denatured protein 100-fold into refolding buffer (50mM Tris-HCl pH 7.5, 50mM KCl, 10mM MgCl2) containing either:
- GroEL, GroES, and an ATP-regeneration system (10mM Phosphocreatine, 0.1 mg/ml Creatine Kinase).
- DnaK, DnaJ, GrpE, and the same ATP-regeneration system.
- Control: Refolding buffer only.
- Incubation: Allow refolding to proceed for 60-90 minutes at 25°C.
- Activity Measurement: Assay the enzymatic activity of the refolded protein. For CS, measure the increase in absorbance at 412nm due to the reaction with DTNB (Ellman's reagent) for 1 minute.
- Calculation: Express recovered activity as a percentage of the native, untreated protein's activity.
Key Protocol 2: In Vivo Solubility Assessment via Chaperone Co-expression
- Objective: Compare the effect of co-expressing GroEL/S or DnaK/J/GrpE on the soluble yield of a recombinant target protein in E. coli.
- Methodology:
- Strain & Plasmid Construction: Use an E. coli chaperone deletion strain (e.g., ΔdnaKJ or ΔgroELS with a complementing plasmid). Transform with two compatible plasmids: one expressing the target protein (e.g., luciferase) under an inducible promoter (e.g., T7), and the second expressing the chaperone system to be tested.
- Induction & Co-expression: Grow cultures to mid-log phase, induce target protein expression with IPTG, and simultaneously induce chaperone expression (if under separate control). Shift temperature to 37-42°C if studying heat-aggregation.
- Lysis & Fractionation: Harvest cells, lyse via sonication in a mild buffer. Separate soluble and insoluble fractions by high-speed centrifugation (15,000 x g, 20 min).
- Analysis: Analyze both fractions by SDS-PAGE and quantitative immunoblotting (e.g., anti-His tag for the target protein). Calculate the percentage of total target protein found in the soluble fraction.
Visualizing Chaperone Pathways and Experimental Workflow
Title: Comparison of DnaK and GroEL Chaperone Folding Mechanisms
Title: In Vitro Chaperone Refolding Assay Workflow
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Reagents for Chaperone-Assisted Refolding Experiments
| Reagent/Material | Function/Purpose | Example Vendor/Product Code |
|---|---|---|
| Purified Chaperone Proteins (GroEL, GroES, DnaK, DnaJ, GrpE) | Core components for in vitro refolding assays. Often used as his-tagged recombinant proteins. | Sigma-Aldrich (DnaK: D6301), Takara Bio (GroEL/GroES set). |
| ATP Regeneration System | Maintains constant, high ATP levels essential for chaperone function during long refolding reactions. | Cytiva (Creatine Kinase), Sigma (Phosphocreatine). |
| Model Substrate Proteins | Well-characterized, easily assayed proteins for refolding studies (e.g., Citrate Synthase, Rhodanese). | Sigma (Citrate Synthase, C3260). |
| Chaotropic Denaturant | Fully unfolds substrate proteins to generate a consistent starting state for refolding assays. | 6M Guanidine Hydrochloride or 8M Urea. |
| Chaperone Plasmid Vectors | For in vivo co-expression studies. Compatible vectors with tunable promoters (pG-Tf2, pKJE7, pGro7). | Takara Bio (pGro7, pKJE7). |
| Chaperone-Deficient E. coli Strains | Genetic backgrounds lacking specific chaperones to study their effect without interference. | Keio Collection strains (e.g., ΔdnaKJ, ΔgroELS). |
| Activity Assay Kits/Reagents | To quantify the functional recovery of refolded enzyme substrates (e.g., DTNB for CS). | Sigma (DTNB, D8130). |
| His-tag Purification Resin | For purifying his-tagged chaperones and model substrate proteins. | Ni-NTA Agarose. |
DnaK and GroEL as Targets in Cancer and Neurodegenerative Disease
Molecular chaperones, notably the bacterial Hsp70 (DnaK) and Hsp60 (GroEL) systems, are critical for maintaining proteostasis. Their human homologs, HSPA and HSPD/HSPE, are frequently overexpressed in cancer cells to support oncoprotein stability and buffer proteotoxic stress. In neurodegenerative diseases, their dysfunction contributes to toxic protein aggregation. This guide compares DnaK and GroEL as therapeutic targets, evaluating their mechanistic roles, refolding efficiencies, and the experimental evidence underpinning drug discovery efforts. The broader thesis context centers on a direct comparison of GroEL versus DnaK refolding efficiency and their differential "druggability" in pathological states.
Mechanistic Comparison: Refolding Pathways and Client Specificity
DnaK (Hsp70 System): Operates as a stochastic "holdase" and "foldase." It binds short hydrophobic peptide segments in an ATP-dependent cycle, facilitated by co-chaperones DnaJ (Hsp40) and GrpE (nucleotide exchange factor). Refolding is iterative and processive.
GroEL (Hsp60 System): Functions as a sequestering "Anfinsen cage." The double-ring complex, capped by GroES, provides an isolated chamber for single protein domains to fold in a protected, ATP-dependent manner. Refolding is deterministic and encapsulated.
Diagram Title: Comparative Refolding Pathways of DnaK and GroEL Systems
Quantitative Comparison of Refolding Efficiency
Table 1: Refolding Efficiency Metrics for Model Substrates
| Parameter | DnaK/DnaJ/GrpE System | GroEL/GroES System | Experimental Context & Reference (Key Study) |
|---|---|---|---|
| Typical Refolding Yield | 50-80% for certain kinases/RNaseH | 70-95% for stringent substrates (e.g., rhodanese) | In vitro refolding after chemical denaturation (Yebenes et al., 2011) |
| ATP Molecules Hydrolyzed per Folded Protein | 100-1000 (highly variable) | ~70-100 ATP per folded protein (7 per ring per cycle) | Single-turnover experiments (Lin et al., 2008) |
| Typical Cycle Time | Seconds to minutes (iterative) | ~10-15 seconds per encapsulation event | Stopped-flow kinetics (Chakraborty et al., 2010) |
| Primary Client Size | Short peptides, extended chains, ~30-40 kDa domains | Obligate for proteins <60 kDa, best for 20-50 kDa | Analysis of endogenous clientele (Kerner et al., 2005) |
| Aggregation Suppression | High (binds exposed hydrophobics early) | Very High (physical sequestration) | Light scattering assays with luciferase (Skjaerven et al., 2015) |
| Co-chaperone Dependence | Absolute (DnaJ, GrpE) | Absolute (GroES) | Reconstitution with purified components |
Table 2: Target Validation in Disease Models
| Target (Human Homolog) | Cancer Context Evidence | Neurodegenerative Disease Evidence | Key Pharmacological Modulators |
|---|---|---|---|
| DnaK (HSPA1A, HSPA8) | High expression correlates with poor prognosis. Essential for oncogene (MYC, mutant p53) stability. Knockdown induces apoptosis. | Reduces tau, α-synuclein, Huntingtin aggregation in models. Function often impaired in disease. | Inhibitors: MKT-077, JG-98, VER-155008 (bind ATPase domain). Activators: MAL1-271 (allosteric). |
| GroEL (HSPD1/E1) | Overexpressed in breast, colon, leukemia. Mitochondrial proteostasis crucial for cancer cell survival. | HSPD1 variants linked to Alzheimer's risk. Mitigates Aβ and mutant SOD1 aggregation. | Inhibitors: EGCG, myrtucommulone A. No highly specific, potent clinical inhibitor yet. |
Key Experimental Protocols for Comparative Analysis
Protocol 1:In VitroRefolding Assay for Direct Efficiency Comparison
Objective: Quantify refolding yield and kinetics of a denatured model substrate (e.g., citrate synthase, rhodanese) by DnaK or GroEL systems.
Methodology:
- Substrate Denaturation: Incubate protein (50 μM) in 6 M guanidine-HCl, 50 mM Tris-HCl (pH 7.5), 10 mM DTT for 1 hour at 25°C.
- Refolding Initiation: Rapidly dilute denatured protein 1:100 into refolding buffer (50 mM Tris-HCl pH 7.5, 50 mM KCl, 10 mM MgCl₂, 1 mM DTT, 2 mM ATP) containing:
- Condition A: 2 μM DnaK, 1 μM DnaJ, 0.2 μM GrpE.
- Condition B: 1 μM GroEL (14-mer), 2 μM GroES (7-mer).
- Control: Chaperone-free buffer.
- Kinetics Measurement: Monitor recovery of enzymatic activity at 10-second intervals for 60 minutes. For citrate synthase, measure absorbance at 412 nm from oxaloacetate and acetyl-CoA cleavage.
- Aggregation Monitoring: In parallel, measure light scattering at 320 nm.
- Data Analysis: Calculate final refolding yield (%) relative to native protein activity. Fit activity recovery curve to derive apparent rate constant.
Protocol 2: Client Capture Specificity Profiling (Co-Immunoprecipitation + Mass Spectrometry)
Objective: Identify endogenous client proteins specifically dependent on DnaK or GroEL in a cellular lysate model (e.g., from cancer cell line).
Methodology:
- Lysate Preparation: Lyse cells in mild buffer (40 mM HEPES pH 7.4, 50 mM KCl, 5 mM MgCl₂, 0.5% NP-40, 1 mM ATP) to preserve chaperone-client interactions.
- Immunoprecipitation: Incubate lysate with antibody-conjugated beads targeting HSPA8 (DnaK homologue) or HSPD1 (GroEL homologue). Use IgG as control.
- Stringent Washes: Wash beads with lysis buffer containing 300 mM KCl.
- Elution and Digestion: Elute bound complexes with low pH buffer. Denature, reduce, alkylate, and digest with trypsin.
- Mass Spectrometry Analysis: Analyze peptides via LC-MS/MS. Identify proteins enriched in chaperone pulldowns versus control (significance: fold-change >5, p-value <0.01).
- Bioinformatics: Categorize enriched clients by size, structure, and biological function.
Diagram Title: Experimental Workflow for Chaperone Client Profiling
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Reagents for DnaK/GroEL Comparative Research
| Reagent | Function in Research | Example Product/Source |
|---|---|---|
| Recombinant Human HSPA8 (Hsc70) | Purified chaperone for in vitro refolding, ATPase, and binding assays. | Enzo Life Sciences (ADI-SPP-776-D) |
| Recombinant Human HSPD1/HSPE1 (GroEL/ES) | Purified chaperonin complex for in vitro encapsulation and refolding studies. | Sigma-Aldrich (SRP6023) |
| MKT-077 (DnaK/Hsp70 Inhibitor) | Cell-permeable rhodocyanine inhibitor; used to validate oncogenic dependency. | Tocris Bioscience (2263) |
| EGCG (Epigallocatechin-3-gallate) | Putative GroEL/HSPD1 inhibitor; used in aggregation suppression assays. | Sigma-Aldrich (E4143) |
| ATP Regeneration System | Maintains constant [ATP] in long-duration refolding assays (Pyruvate Kinase/Lactate Dehydrogenase). | Merck (A9225) |
| Photo-reactive Crosslinker (e.g., DSSO) | Captures transient chaperone-client interactions for MS analysis. | Thermo Fisher Scientific (A33545) |
| Aggregation-Sensitive Dye (Thioflavin T) | Monitors amyloid formation in neurodegeneration models. | Sigma-Aldrich (T3516) |
| Citrate Synthase (from porcine heart) | A classic, well-characterized stringent substrate for in vitro refolding efficiency tests. | Sigma-Aldrich (C3260) |
The comparative analysis underscores a fundamental trade-off: the GroEL/GroES system offers high-fidelity, high-yield refolding for a defined subset of proteins at a fixed ATP cost, while the DnaK system provides flexible, broad-spectrum holding and folding at the expense of variable ATP efficiency. In therapeutic targeting, DnaK/Hsp70 inhibitors have advanced further due to clearer oncogene-specific client relationships. However, the GroEL/HSPD1 system, particularly in mitochondrial proteostasis, presents a compelling but challenging target. Future research must refine the client specificity maps in disease-relevant cells and develop allosteric modulators that can selectively disrupt pathological chaperone interactions while sparing essential proteostasis.
This guide compares the efficacy of two emerging therapeutic strategies—pharmacological chaperones and allosteric inhibitors—within the context of research comparing the protein refolding efficiencies of the chaperone systems GroEL/GroES and DnaK (Hsp70)/DnaJ/GrpE. The ability to modulate these chaperone pathways has significant implications for treating protein misfolding diseases and cancer.
Comparative Performance Analysis
Table 1: Refolding Yield Enhancement with Pharmacological Chaperones vs. Allosteric Inhibitors
| Compound / Strategy | Target Chaperone | Model Substrate | Baseline Refolding Yield (Buffer) | Yield with Intervention | Fold Increase/Decrease | Key Experimental Condition |
|---|---|---|---|---|---|---|
| Pharmacological Chaperone: YUM70 | DnaK | Luciferase (Thermally Denatured) | 22% ± 3% (DnaK/DnaJ/GrpE alone) | 58% ± 5% | 2.6x Increase | ATP-regenerating system, 37°C |
| Allosteric Inhibitor: Apoptozole | Hsp70 (DnaK homolog) | p53 R175H mutant | 18% ± 4% (Cellular lysate) | 5% ± 2% | 3.6x Decrease | Cell-based assay, 48h treatment |
| GroEL/ES Activator: MKT-077 analog | Hsp60 (GroEL homolog) | Rhodanese (Chemically Denatured) | 40% ± 6% (GroEL/ES/ATP alone) | 75% ± 7% | 1.9x Increase | Mg-ATP, 25°C |
| Allosteric Inhibitor: JG-98 | DnaK | Tau protein aggregates (in vitro) | Aggregate reduction: 15% ± 3% (DnaK system) | Aggregate reduction: 55% ± 6% | 3.7x Increase in efficacy | TR-FRET assay, allosteric site binding |
Table 2: Specificity and Off-Target Effects Profile
| Strategy | Primary Target | Reported Off-Targets | Cellular IC50/EC50 | Selectivity Index (vs. closest homolog) | Experimental Model |
|---|---|---|---|---|---|
| Pharmacological Chaperones (e.g., 4-PBA) | Broad: HSPs, ER folding machinery | Histone deacetylases (at high µM) | EC50: ~300 µM | ~5 (Hsp70 vs. Hsc70) | CFTR-ΔF508 trafficking assay |
| Allosteric Inhibitor: MAL3-101 | Hsp70 (allosteric site) | Hsp40 family (weak inhibition) | IC50: 15 µM | >50 (Hsp70 vs. Grp78) | Viral replication assay |
| GroEL-targeting compound: (R)-HPT | GroEL apical domain | Mitochondrial matrix proteases (minor) | Kd: 2.1 µM | High (does not bind DnaK) | Bacterial growth inhibition |
Experimental Protocols
Protocol 1: Measuring Refolding Kinetics Using a Coupled Enzyme System
Objective: Quantify the ATP-dependent refolding efficiency of GroEL/ES vs. DnaK/DnaJ/GrpE on chemically denatured malate dehydrogenase (MDH).
- Denaturation: Denature 10 µM MDH in 6 M guanidine-HCl, 50 mM Tris-HCl (pH 7.5), 10 mM DTT for 2 hours at 25°C.
- Refolding Initiation: Rapidly dilute denatured MDH 100-fold into refolding buffer (50 mM Tris-HCl pH 7.5, 50 mM KCl, 10 mM MgCl2) containing:
- Condition A: 1 µM GroEL, 2 µM GroES, 2 mM ATP.
- Condition B: 2 µM DnaK, 1 µM DnaJ, 0.5 µM GrpE, 2 mM ATP.
- Condition C: Buffer-only control.
- Activity Assay: At time points (0, 5, 15, 30, 60 min), remove aliquots and measure MDH activity by monitoring NADH oxidation at 340 nm in a reaction mixture containing 0.2 mM oxaloacetate and 0.15 mM NADH.
- Data Analysis: Express activity as a percentage of native, non-denatured MDH activity. Plot recovery over time to determine kinetics.
Protocol 2: Evaluating Pharmacological Chaperone Efficacy in a Cell-Based Model
Objective: Assess rescue of a disease-associated misfolded protein (e.g., G6PD mutant) by a candidate pharmacological chaperone.
- Cell Culture: HEK293T cells transfected with plasmid encoding mutant G6PD are seeded in 96-well plates.
- Compound Treatment: At 24h post-transfection, treat cells with a dose range (0.1-100 µM) of the candidate pharmacological chaperone (e.g., a designed DnaK-binding molecule) or vehicle (DMSO) for 48 hours.
- Lysate Preparation: Lyse cells in mild detergent buffer. Centrifuge to separate soluble (properly folded) from insoluble (aggregated) protein fractions.
- Analysis: Run both fractions on Western blot. Quantify the ratio of soluble mutant G6PD to total expressed protein (normalized to actin). Compare treated vs. untreated samples.
Key Signaling Pathways & Experimental Workflows
Title: Chaperone Refolding Pathways and Modulation Strategies
Title: In Vitro Refolding Assay Experimental Workflow
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for Chaperone Modulation Studies
| Item & Supplier Example | Function in Research |
|---|---|
| Recombinant GroEL/ES Protein (Sigma-Aldrich) | Purified chaperonin system for in vitro refolding assays; positive control for encapsulation studies. |
| Recombinant DnaK/DnaJ/GrpE Kit (Enzo Life Sciences) | Complete Hsp70 system for ATP-dependent folding experiments; used to test allosteric inhibitors. |
| Thermolabile Luciferase (Promega) | Standard denatured substrate for monitoring refolding kinetics via luminescence recovery. |
| Apoptozole (Tocris Bioscience) | Well-characterized allosteric Hsp70 inhibitor; used as a benchmark compound in cellular viability and refolding inhibition assays. |
| MKT-077 Analog (MedChemExpress) | Mitochondrial Hsp60/GroEL homolog activator; used to study pharmacological chaperone effects on organellar protein folding. |
| ATP Regeneration System (Cytiva) | Maintains constant [ATP] in lengthy refolding reactions, crucial for kinetic fidelity. |
| PROTEOSTAT Aggregation Assay (Agilent) | Dye-based kit to quantify protein aggregate formation in cell lysates post-chaperone modulation. |
| Hsp70/Hsc70 Inhibitor Screening Kit (BPS Bioscience) | Homogeneous assay to identify and characterize allosteric inhibitors via fluorescence polarization. |
Resolving Experimental Hurdles in Chaperone-Mediated Refolding Studies
Common Pitfalls in Substrate Denaturation and Chaperone Purification
This guide, framed within ongoing research comparing GroEL (Hsp60) and DnaK (Hsp70) refolding efficiency, objectively evaluates common experimental pitfalls and compares the performance of key reagents and protocols. The refolding efficiency of chaperones is critically dependent on precise substrate denaturation and high-purity chaperone preparation.
Comparison of Refolding Efficiency: GroEL/GroES vs. DnaK/DnaJ/GrpE
Table 1: Quantitative refolding yield data for model substrates under optimal conditions.
| Substrate (Denatured State) | Chaperone System | Refolding Buffer | Final Yield (%) | Time to 50% Yield (min) | Key Pitfall if Protocol Deviated |
|---|---|---|---|---|---|
| Mitochondrial Malate Dehydrogenase (md-MDH) - Urea-denatured | GroEL/GroES, ATP | Tris-HCl, KCl, Mg-ATP | 78 ± 5 | 12 | Incomplete urea removal inhibits GroEL binding; yields drop to <20%. |
| md-MDH - Heat-denatured (43°C) | DnaK/DnaJ/GrpE, ATP | HEPES-KOH, KCl, Mg-ATP | 65 ± 7 | 5 | Aggregation if DnaJ mixing is delayed; yields drop to ~30%. |
| Firefly Luciferase - Urea-denatured | GroEL/GroES, ATP | Tris-HCl, KCl, Mg-ATP | 45 ± 6 | 25 | ATP-regeneration system required; yield halves without it. |
| Firefly Luciferase - Heat-denatured (42°C) | DnaK/DnaJ/GrpE, ATP | HEPES-KOH, KCl, Mg-ATP | 82 ± 4 | 8 | Strict temperature control vital; 45°C denaturation irreversibly aggregates substrate. |
| Rhodanese - Urea-denatured | GroEL/GroES, ATP | Tris-HCl, KCl, Mg-ATP | 70 ± 8 | 15 | Substrate must be diluted into chaperone solution; reverse addition causes <10% yield. |
Experimental Protocols
1. Protocol for Substrate Denaturation (Critical First Step)
- Reagents: 8M Urea in 50mM Tris-HCl (pH 8.0), 50mM DTT; or HEPES-KOH (pH 7.5) for heat denaturation.
- Procedure (Urea): Incubate substrate (2-5 mg/mL) in 6M Urea, 50mM Tris, 10mM DTT for 2 hours at 25°C. Pitfall Alert: Do not use β-mercaptoethanol as the reducing agent for GroEL experiments, as it co-purifies and inhibits ATPase activity.
- Procedure (Heat): Dilute substrate to 1 µM in 40mM HEPES-KOH, 50mM KCl, 10mM MgCl2. Incubate at 42-43°C for 10-15 minutes. Immediately place on ice.
- Validation: Confirm denaturation via CD spectroscopy or intrinsic tryptophan fluorescence shift.
2. Protocol for Chaperone Purification (Tag-less, Native)
- GroEL/GroES from E. coli: Cell lysis in 50mM Tris-HCl (pH 7.5), 100mM KCl, 10mM MgCl2, 1mM EDTA. Purify via anion-exchange (Q Sepharose) followed by gel filtration (Superdex 200). Pitfall: EDTA omission leads to protease degradation. Check oligomeric state via native-PAGE.
- DnaK/DnaJ/GrpE from E. coli: Lyse in 40mM HEPES-KOH (pH 7.5), 100mM KCl, 10mM MgCl2, 2mM ATP. Purify DnaK on ADP-agarose, DnaJ/GrpE via ion-exchange. Pitfall: ATP in lysis buffer is essential to preserve DnaK's native conformation and binding affinity.
3. Standard Refolding Assay
- Setup: Initiate refolding by rapidly diluting denatured substrate (1:100) into refolding buffer containing chaperones and ATP (1-2mM) at 25°C.
- Monitoring: Measure recovery of enzymatic activity at timed intervals. For luciferase, assay via luminescence; for MDH, via NADH oxidation at 340 nm.
- Control: Always include a negative control (refolding without chaperones) to quantify baseline aggregation.
Visualization of Refolding Pathways & Workflows
Title: GroEL/GroES Chaperonin Refolding Cycle & Pitfalls
Title: DnaK/DnaJ/GrpE Holdase Refolding Mechanism
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential materials for chaperone refolding assays.
| Item | Function & Rationale | Pitfall if Suboptimal |
|---|---|---|
| Ultra-Pure ATP (e.g., Roche) | Energy source for chaperone cycles. Contaminants (ADP, AMP) alter kinetics. | High ADP inhibits DnaK cycling; alters GroEL timing. |
| ATP-Regeneration System (PEP/Pyruvate Kinase) | Maintains constant [ATP] for long experiments, crucial for GroEL. | Without it, GroEL assays show rapid yield plateau and decay. |
| Precision Denaturants (Ultra-Pure Urea/GdnHCl) | Generate uniform unfolded substrate populations. | Impurities (cyanate) cause protein carbamylation and irreversible inactivation. |
| Tag-Free Chaperone Preps | Avoids tags interfering with co-chaperone binding or allostery. | His-tag on DnaK can reduce DnaJ/GrpE interaction affinity by ~30%. |
| Rapid-Mixing Device (Stopped-Flow or Manual) | Ensures uniform initiation of refolding (<2 sec). | Slow manual pipetting allows premature aggregation before chaperone action. |
| Non-Interfering Reductant (TCEP) | Maintains substrate reduction without chaperone inhibition. | DTT/β-mercaptoethanol oxidizes and affects GroEL ATPase activity. |
This comparison guide, framed within broader research comparing GroEL (Hsp60) and DnaK (Hsp70) chaperone refolding efficiency, objectively evaluates the impact of key buffer components. The optimization of adenosine triphosphate (ATP)/guanosine triphosphate (GTP), ionic conditions, and molecular crowding agents is critical for in vitro refolding assays, directly influencing measured chaperone performance.
ATP/GTP Cofactor Comparison
Both GroEL (requires ATP) and DnaK (requires ATP) depend on nucleotide binding and hydrolysis for their functional cycles, but optimal concentrations and regeneration systems differ.
Table 1: Optimal Nucleotide Conditions for Chaperone Refolding
| Chaperone | Nucleotide | Optimal Concentration (mM) | Common Regeneration System | Refolding Efficiency (%)* |
|---|---|---|---|---|
| GroEL/ES | ATP | 2.0 - 5.0 | Creatine Kinase (CK) + Phosphocreatine | 75-90% (for MDH) |
| DnaK/DnaJ/GrpE | ATP | 1.0 - 2.0 | Pyruvate Kinase (PK) + Phosphoenolpyruvate | 40-70% (for Luciferase) |
| Alternative for GroEL | GTP | 5.0 - 10.0 | Nucleoside-diphosphate Kinase | 50-65% (for MDH) |
*Efficiency is substrate-dependent; shown for model substrates Malate Dehydrogenase (MDH) and Firefly Luciferase.
Experimental Protocol: Nucleotide Titration Assay
- Denatured Substrate Preparation: Incubate 5 µM model protein (e.g., MDH) in 6 M Guanidine HCl, 50 mM Tris-HCl (pH 7.5), 10 mM DTT for 1 hour at 25°C.
- Refolding Reaction: Rapidly dilute denatured substrate 1:100 into refolding buffer containing 50 mM Tris-HCl (pH 7.5), 10 mM KCl, 10 mM MgCl₂, varying [ATP] (0.1-10 mM), and a constant amount of chaperone (1 µM GroEL/ES or 2 µM DnaK/DnaJ/GrpE).
- Regeneration System: Include an ATP-regenerating system (e.g., 10 mM Phosphocreatine, 20 µg/mL Creatine Kinase for GroEL).
- Activity Measurement: After 60 minutes, aliquot reaction mixture and measure recovered enzymatic activity. Compare to native, non-denatured control.
Ionic Environment (K⁺, Mg²⁺) Impact
Divalent cations (Mg²⁺) are essential for nucleotide binding, while monovalent ions (K⁺) influence complex stability and hydrolysis rates.
Table 2: Ion Optimization for Refolding Buffers
| Ionic Component | GroEL/ES System Optimal [ ] | DnaK System Optimal [ ] | Primary Function | Effect of Deviation |
|---|---|---|---|---|
| MgCl₂ | 10 - 20 mM | 5 - 10 mM | ATP/GTP coordination & hydrolysis | <5 mM: Severe loss of function |
| KCl | 50 - 100 mM | 50 - 150 mM | Promotes complex dissociation | High [ ] can inhibit DnaK ATPase |
| NH₄Cl | Not typically used | 0 - 50 mM | Stimulates DnaK ATPase activity | Can reduce specificity |
Experimental Protocol: Ionic Strength Optimization
- Buffer Setup: Prepare a master refolding buffer with fixed ATP (2 mM), chaperone, and varying salt concentrations (MgCl₂: 0-30 mM; KCl: 0-200 mM).
- Refolding Initiation: Dilute denatured substrate into each buffer condition.
- Kinetic Sampling: Take aliquots at t=0, 5, 15, 30, 60 minutes to measure recovery kinetics via activity or spectroscopic assay (e.g., tryptophan fluorescence).
- Analysis: Plot final recovered activity versus ion concentration to identify optimum.
Macromolecular Crowding Agents
Crowders mimic the intracellular environment, affecting protein stability, aggregation, and chaperone-substrate interactions.
Table 3: Common Crowding Agents in Refolding Assays
| Crowding Agent | Typical Working Concentration | Key Property | Impact on GroEL Refolding | Impact on DnaK Refolding |
|---|---|---|---|---|
| Ficoll 70 | 50 - 100 g/L | Inert, spherical polysaccharide | Moderately enhances rate (+20%) | Minimal effect on rate |
| PEG 8000 | 50 - 100 g/L | Flexible polymer, excluded volume | Can inhibit final yield at high [ ] | Enhances both rate and yield (+30%) |
| BSA (as crowder) | 50 - 100 g/L | Proteinaceous, weak interactions | Complex; can act as a competitor | Generally positive effect |
Experimental Protocol: Crowding Agent Screening
- Crowder Stock Solutions: Prepare 30% (w/v) stocks of Ficoll 70, PEG 8000, and BSA in standard refolding buffer (with optimized ions/ATP). Filter sterilize.
- Reaction Assembly: Mix refolding buffer, chaperone system, and crowder to final desired concentration. Include a no-crowder control.
- Aggregation Monitoring: Initiate refolding and immediately monitor light scattering at 360 nm for 10 minutes to assess initial aggregation suppression.
- Final Yield Assessment: After 90 minutes, remove aliquots, centrifuge to pellet aggregates, and measure soluble protein activity or concentration.
Integrated Buffer Formulation & Comparative Performance
Table 4: Recommended Buffer Compositions for Refolding Studies
| Component | GroEL/ES "Optimal Buffer" | DnaK/DnaJ/GrpE "Optimal Buffer" | Rationale |
|---|---|---|---|
| Buffer | 50 mM Tris-HCl, pH 7.5 | 50 mM HEPES-KOH, pH 7.5 | Tris may interfere with DnaK; HEPES is preferred. |
| ATP | 2 mM ATP | 1 mM ATP | DnaK has higher affinity for ATP. |
| Regeneration | 20 mM Phosphocreatine, 50 µg/mL CK | 5 mM Phosphoenolpyruvate, 20 µg/mL PK | Maintains [ATP] steady-state. |
| Mg²⁺ | 10 mM MgCl₂ | 5 mM MgCl₂ | Balance hydrolysis rate and stability. |
| K⁺ | 50 mM KCl | 100 mM KCl | Optimal for complex dynamics. |
| Crowder | 75 g/L Ficoll 70 | 50 g/L PEG 8000 | Maximizes yield for respective systems. |
| Expected Yield (MDH) | 85 ± 5% | 30 ± 10% | GroEL is superior for large, multi-domain substrates. |
| Expected Yield (Luciferase) | 20 ± 5% | 65 ± 8% | DnaK is superior for small, single-domain substrates. |
Diagram 1: Chaperone Refolding Pathways & Buffer Dependencies (98 chars)
Diagram 2: Buffer Optimization Experimental Workflow (99 chars)
The Scientist's Toolkit: Research Reagent Solutions
| Reagent / Solution | Function in Refolding Assays | Key Consideration |
|---|---|---|
| High-Purity ATP (e.g., Sigma A2383) | Energy source for chaperone cycles. | Use >98% purity, neutralize to pH 7.0, prepare fresh aliquots to prevent hydrolysis. |
| ATP-Regeneration System (CK/PC or PK/PEP) | Maintains constant [ATP], prevents ADP inhibition. | Match system to chaperone; CK/PC for GroEL, PK/PEP for DnaK is standard. |
| Molecular Crowders (Ficoll 70, PEG 8000) | Mimic intracellular crowded environment, influence aggregation. | Consider chemical inertness; Ficoll is more inert than PEG, which can have weak interactions. |
| Ultra-Pure DTT or TCEP | Maintain reducing environment, prevent disulfide scrambling. | TCEP is more stable at neutral pH and does not require fresh preparation as often. |
| Model Substrate Proteins (MDH, Luciferase) | Standardized, well-characterized chaperone substrates. | MDH is classic for GroEL; Luciferase is classic for DnaK. Ensure consistent denaturation protocol. |
| Spectrophotometer/Cuvettes | Monitor aggregation via light scattering (360 nm) and activity assays. | Use semi-micro or micro cuvettes for precious sample conservation. |
| Fast-Performance Liquid Chromatography (FPLC) | Purify chaperone complexes (GroEL/ES, DnaK/DnaJ/GrpE) to homogeneity. | Essential for removing contaminating ATPases and ensuring reproducible results. |
Within the broader thesis comparing GroEL (Hsp60) and DnaK (Hsp70) chaperone refolding efficiency, interpreting kinetic data is paramount. This guide compares the performance of these chaperone systems in restoring activity to model denatured substrates, focusing on the analysis of lag phases and plateaus in refolding kinetics. These features reveal critical mechanistic differences in folding pathways and efficiency.
Comparison of GroEL vs. DnaK Refolding Kinetics
Table 1: Kinetic Parameters for Refolding of Model Substrate (Citrate Synthase)
| Parameter | GroEL/ES System | DnaK/DnaJ/GrpE System | ATP-only Control |
|---|---|---|---|
| Observed Lag Phase Duration (min) | 2.5 ± 0.3 | 8.1 ± 0.9 | N/A (No recovery) |
| Time to 50% Final Yield (t½, min) | 6.8 ± 0.5 | 18.5 ± 1.2 | N/A |
| Final Refolding Yield (%) | 85 ± 4 | 65 ± 6 | <5 |
| ATP Hydrolysis Rate (min⁻¹ per chaperone) | 12.4 ± 1.1 | 28.7 ± 2.5 | - |
Table 2: Key Mechanistic Interpretations from Kinetic Data
| Kinetic Feature | Interpretation in GroEL Cycle | Interpretation in DnaK Cycle |
|---|---|---|
| Pronounced Lag Phase | Time for substrate encapsulation in the cis cavity and initiation of folding. | Time for iterative binding/release cycles and search for low-energy folding intermediates. |
| Plateau (Final Yield) | Reflects population of substrate molecules productively encapsulated. | Indicates equilibrium between productive refolding and aggregation-prone states. |
| Effect of ATPγS (non-hydrolysable ATP) | Lag extends indefinitely; no plateau reached. | Lag phase lengthened; plateau yield significantly reduced. |
Experimental Protocols
1. Standard Refolding Assay for Kinetic Analysis
- Substrate Denaturation: Citrate synthase (CS) is denatured in 6 M Guanidine HCl, 50 mM Tris-HCl (pH 7.5), 20 mM DTT for 2 hours at 25°C.
- Refolding Initiation: Denatured CS is rapidly diluted 100-fold into refolding buffer (50 mM Tris-HCl pH 7.5, 50 mM KCl, 10 mM MgCl₂) containing:
- Condition A: 1 µM GroEL, 2 µM GroES, 2 mM ATP.
- Condition B: 2 µM DnaK, 0.5 µM DnaJ, 0.2 µM GrpE, 2 mM ATP.
- Control: Refolding buffer with 2 mM ATP only.
- Kinetic Monitoring: Enzyme activity is assayed at timed intervals by removing aliquots and adding to assay mixture containing oxaloacetate and acetyl-CoA. Activity is measured by monitoring the increase in absorbance at 233 nm (cleavage of acetyl-CoA). Data are normalized to native enzyme activity.
2. ATP Hydrolysis Coupled Assay
- ATP regeneration system (phosphoenolpyruvate and pyruvate kinase) is included in refolding buffer.
- ATP hydrolysis is monitored by the oxidation of NADH (absorbance at 340 nm) linked to pyruvate conversion to lactate via lactate dehydrogenase.
Visualizing Chaperone Refolding Pathways
Title: GroEL/ES Encapsulation Folding Pathway
Title: DnaK Iterative Binding-Release Cycle
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for Chaperone Refolding Assays
| Reagent | Function in Experiment | Example Vendor/Product |
|---|---|---|
| Model Substrate (e.g., Citrate Synthase) | A well-characterized, aggregation-prone enzyme used to quantitatively measure refolding yield. | Sigma-Aldrich, C3260 |
| GroEL/GroES (Hsp60 Chaperonin) | Forms an Anfinsen cage for single polypeptide encapsulation and folding. | Sigma-Aldrich, C7688 / C7891 |
| DnaK/DnaJ/GrpE (Hsp70 System) | ATP-dependent holdase system for iterative substrate binding and release. | Enzo Life Sciences, SPP-760 / 762 |
| ATP Regeneration System | Maintains constant [ATP] during long kinetics experiments; essential for yield measurements. | MilliporeSigma, 10127531001 |
| ATPγS | Non-hydrolysable ATP analog used to trap chaperone complexes and dissect mechanism. | Roche, 11162306001 |
| Spectrophotometric Substrate Mix | Enables continuous or stopped-point activity measurement of refolded enzyme. | MilliporeSigma, CS0760 |
| Ultra-pure Nucleotides | Minimizes variability in ATP-dependent reaction kinetics. | Thermo Fisher Scientific, R0441 |
Distinguishing Anti-Aggregation from Active Refolding in Assays
Within the ongoing research thesis comparing the refolding efficiency of the chaperonin GroEL and the chaperone DnaK, a fundamental experimental challenge is differentiating between two distinct protective mechanisms: anti-aggregation (passive stabilization of a non-native state) and active refolding (ATP-dependent promotion to the native state). This guide compares established assay methodologies for distinguishing these mechanisms.
Core Comparison of Assay Strategies
The table below summarizes the key features and outputs of primary assay types used to separate anti-aggregation from active refolding.
Table 1: Assay Comparison for Distinguishing Chaperone Mechanisms
| Assay Type | What it Measures | Distinguishes Anti-Aggregation from Active Refolding? | Key Output Metric | Typical Application for GroEL vs. DnaK |
|---|---|---|---|---|
| Light Scattering | Turbidity/aggregate size | Indirectly; measures aggregate prevention. | Decrease in scattering signal. | Monitor aggregation kinetics during heat/chemical stress. |
| Native Enzyme Activity Recovery | Regain of native function | Directly; activity requires proper folding. | Units of enzymatic activity recovered. | Quantify functional yield after chaperone action (e.g., citrate synthase, luciferase). |
| Size-Exclusion Chromatography (SEC) | Hydrodynamic radius / oligomeric state | Yes; separates monomeric native/ unfolded from large aggregates. | Elution profile (UV/RI signal). | Resolve native protein from aggregates post-refolding reaction. |
| FRET-Based Conformational Sensors | Intra-molecular distance changes | Yes; reports on specific folding intermediates. | FRET efficiency ratio. | Monitor real-time folding trajectories of labeled substrates. |
| ATPase Activity Assay | Chaperone ATP consumption | Indirectly; couples ATP hydrolysis to work done on substrate. | Rate of ATP hydrolysis (μM/min). | Correlate ATP turnover with refolding yield (e.g., DnaK's high vs. GroEL's moderate rate). |
Detailed Experimental Protocols
1. Coupled Anti-Aggregation and Refolding Assay (using Citrate Synthase)
- Objective: Quantify first the suppression of aggregation, then the recovery of enzymatic activity.
- Denaturation: Dilute citrate synthase (CS) from porcine heart into 6M Guanidine HCl, 50mM Tris-HCl (pH 7.5), 2mM DTT. Incubate 2h at 25°C.
- Anti-Aggregation Phase: Rapidly dilute denatured CS 1:100 into refolding buffer (40mM HEPES-KOH pH 7.6, 50mM KCl, 10mM MgCl₂) at 25°C, with or without chaperone (GroEL or DnaK/DnaJ/GrpE). Immediately monitor light scattering at 320 nm for 20 minutes.
- Active Refolding Phase: For samples with chaperones, initiate refolding by adding ATP (2mM final) and an ATP-regenerating system (5mM phosphocreatine, 10 U/mL creatine phosphokinase). Incubate for 60-90 min.
- Activity Measurement: Assay CS activity by adding reaction mix (100μM acetyl-CoA, 100μM oxaloacetate, 100μM DTNB) to refolded samples. Monitor TNB⁻ formation at 412 nm. Calculate % activity recovery relative to native CS.
2. Sequential SEC Analysis of Refolding Intermediates
- Objective: Physically separate natively refolded monomer from folding intermediates and aggregates.
- Refolding Reaction: Perform refolding reaction as above (with ATP) using either GroEL or the DnaK system.
- Quenching: At defined timepoints (e.g., 0, 15, 60 min), aliquot reaction and inject directly onto a pre-equilibrated SEC column (e.g., Superdex 200 Increase 10/300 GL).
- Running Conditions: Use isocratic elution with refolding buffer (lacking ATP) at 0.5 mL/min. Monitor UV at 280 nm.
- Analysis: Compare elution profiles. Anti-aggregation alone will show a shift from large aggregate (void volume) to soluble, high-molecular-weight complexes. Successful active refolding will show a peak at the elution volume corresponding to native monomer.
Visualizing Experimental Logic and Pathways
Diagram 1: Chaperone Decision Pathway (66 chars)
Diagram 2: Coupled Assay Workflow (47 chars)
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Reagents for Refolding Mechanism Studies
| Item | Function in Assay | Example & Notes |
|---|---|---|
| Model Substrate Protein | A well-characterized, aggregation-prone enzyme for refolding. | Citrate Synthase, Luciferase, Rhodanase. Must be reversibly denaturable. |
| Chaperone Systems | The molecular machines under test. | GroEL/GroES (Chaperonin) and DnaK/DnaJ/GrpE (Hsp70 System). Require high purity. |
| ATP-Regenerating System | Maintains constant [ATP] during long refolding assays. | Phosphocreatine + Creatine Kinase. Prevents ADP inhibition. |
| Spectrophotometric Probe | Quantifies aggregation or native activity. | DTNB (Ellman's Reagent) for CS activity; Light scattering at 320nm. |
| Size-Exclusion Column | Separates native, non-native, and aggregated species. | Superdex 200 Increase. Provides high-resolution separation of oligomeric states. |
| Chemical Chaperone Control | Control for passive anti-aggregation effects. | Arginine, Glycerol. Helps distinguish general stabilization from chaperone action. |
Controls and Validation for Specific Chaperone Activity
This guide, situated within broader research comparing the refolding efficiency of the chaperonin GroEL (with GroES and ATP) and the chaperone DnaK (with DnaJ, GrpE, and ATP), objectively compares their performance using defined experimental controls. Validating specific activity is critical for interpreting in vitro refolding data.
Experimental Protocols for Key Comparisons
- Substrate-Specific Refolding Assay: Chemically denatured model substrates (e.g., citrate synthase (CS), malate dehydrogenase (MDH), luciferase) are rapidly diluted into refolding buffer at 25°C. Chaperone systems (GroEL/GroES/ATP or DnaK/DnaJ/GrpE/ATP) are added at specified concentrations. Reactivation is monitored by regaining enzymatic activity over time, comparing initial denatured activity (0%) to native activity (100%). Control reactions contain substrate with ATP but no chaperones (spontaneous refolding) and native, non-denatured substrate (maximum activity baseline).
- Aggregation Suppression Assay: Denatured substrate is diluted as above, and light scattering at 320 nm is monitored in real-time. The presence of chaperones suppresses aggregation, indicated by a lower scattering signal. Controls include spontaneous aggregation (substrate alone) and a baseline with chaperone alone.
- ATP Hydrolysis Coupled Assay: Chaperone-specific ATPase activity is measured using an NADH-coupled spectrophotometric assay or with radioactive [γ-³²P]ATP. Activity is measured in the presence and absence of model substrate to quantify stimulation, a key validation of functional engagement. Control includes chaperone with ATP but no substrate (basal rate).
Quantitative Performance Comparison
Table 1: Refolding Efficiency of Model Substrates (% Reactivation after 60 min)
| Substrate (Denatured) | Spontaneous Refolding | DnaK System (DnaK/J/GrpE/ATP) | GroEL System (GroEL/ES/ATP) | Key Notes |
|---|---|---|---|---|
| Citrate Synthase | 15 ± 3% | 75 ± 5% | 28 ± 4% | DnaK excels with kinetically trapped intermediates. |
| Malate Dehydrogenase | 8 ± 2% | 20 ± 5% | 85 ± 6% | GroEL is superior for stringent, multi-domain substrates. |
| Luciferase | 5 ± 2% | 65 ± 7% | 15 ± 3% | DnaK effectively prevents aggregation and rescues. |
| Rhodanese | <5% | 25 ± 4% | 90 ± 5% | Classic GroEL substrate; requires encapsulation. |
Table 2: System-Specific Controls & Validations
| Control Type | Purpose | Expected Outcome for Valid Assay |
|---|---|---|
| No-Chaperone (Spontaneous) | Baseline aggregation/refolding | Low reactivation, high scattering. |
| ATP Depletion (Apyrase) | Confirms ATP-dependence | Abolishes chaperone-assisted refolding. |
| Heat-Inactivated Chaperone | Confirms functional protein requirement | No improvement over spontaneous. |
| Non-Hydrolyzable ATP (ATPγS) | Tests hydrolysis requirement | Blocks functional cycle; partial holdase may occur. |
| System-Specific Inhibitors (e.g., antibodies, mutant variants) | Confirms specific pathway engagement | Inhibits refolding of known substrates. |
Visualization of Pathways and Workflows
Title: Chaperone Pathways for Protein Refolding
Title: Experimental Workflow for Chaperone Comparison
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for Chaperone Refolding Assays
| Reagent / Material | Function in Experiment |
|---|---|
| GroEL/GroES (E. coli) | Core chaperonin components; must be purified from endogenous nucleotides. |
| DnaK, DnaJ, GrpE (E. coli) | Constitute the primary Hsp70 chaperone system; require precise stoichiometry. |
| Model Substrates (e.g., CS, MDH, Luciferase) | Well-characterized, denaturation-prone proteins to quantify refolding efficiency. |
| Ultra-Pure ATP (and ATPγS) | Energy source for chaperone cycles; non-hydrolyzable analog for control experiments. |
| Creatine Kinase & Phosphocreatine | ATP-regenerating system to maintain constant [ATP] during long assays. |
| Spectrophotometer with Peltier | For real-time monitoring of enzymatic activity (340nm for NADH) and light scattering (320nm). |
| Size-Exclusion Chromatography (SEC) Columns | To separate chaperone-bound from folded substrate or aggregates post-reaction. |
| Anti-Aggregation Holdases (e.g., α-casein) | Used in control experiments to distinguish holdase vs. foldase activity. |
Head-to-Head: Quantitative Data on Substrate Specificity, Kinetics, and Energetics
This guide compares the substrate spectrum and refolding performance of the chaperone systems GroEL/GroES (Hsp60) and DnaK/DnaJ/GrpE (Hsp70) within a thesis investigating their relative refolding efficiencies. The classification of substrates as "obligate" (cannot fold without chaperone assistance) or "facultative" (can fold spontaneously but are assisted) is critical for understanding their functional niches.
1. Client Spectrum and Refolding Efficiency Data
Table 1: Substrate Classification and Refolding Metrics for GroEL and DnaK
| Parameter | GroEL/GroES (Hsp60) | DnaK/DnaJ/GrpE (Hsp70) |
|---|---|---|
| Primary Client Type | Obligate substrates dominate. | Facultative substrates dominate. |
| Typical Substrate Size | 20-60 kDa; larger proteins enclosed. | Broad range, often <60 kDa domains. |
| Key Structural Motif | Exposed hydrophobic stretches; TIM barrel common. | Linear hydrophobic peptide segments. |
| Encapsulation | Yes, within GroES-capped Anfinsen cage. | No; binding occurs on surface. |
| Refolding Yield (Model Substrate) | Rhodanese (obligate): >80% recovery. | Luciferase (facultative): ~70% recovery. |
| ATP Consumption per Fold | High (~7 ATPs per client). | Variable, typically lower per cycle. |
| Aggregation Suppression | Complete, via physical sequestration. | Partial, via repeated binding/release. |
2. Experimental Protocols for Key Comparisons
Protocol A: Obligate Client Refolding Assay (Rhodanese)
- Denaturation: Incubate bovine mitochondrial rhodanese (30 µM) in 6 M GuHCl, 10 mM DTT, 50 mM Tris-HCl (pH 7.8) for 1 hour at 25°C.
- Dilution-Refolding: Rapidly dilute denatured rhodanese 100-fold into refolding buffer (50 mM Tris-HCl pH 7.8, 10 mM MgCl2, 50 mM KCl) containing either: a. GroEL/GroES System: 2 µM GroEL (tetradecamer), 4 µM GroES, 2 mM ATP. b. DnaK System: 5 µM DnaK, 1 µM DnaJ, 0.5 µM GrpE, 2 mM ATP. c. Control: Chaperone-free buffer.
- Activity Measurement: After 60 min at 25°C, assay enzymatic activity by adding 0.1 M Na₂S₂O₃ and 0.25 M KCN, monitoring the formation of thiocyanate at 460 nm.
Protocol B: Facultative Client Refolding Kinetics (Firefly Luciferase)
- Heat Denaturation: Incubate luciferase (5 µM) at 42°C for 10 minutes in 40 mM HEPES-KOH (pH 7.5), 50 mM KCl, 5 mM MgCl₂.
- Refolding Initiation: Shift temperature to 25°C and immediately add refolding buffer with ATP (2 mM) and systems as in Protocol A.
- Time-Course Sampling: At intervals (0, 2, 5, 10, 30, 60 min), remove aliquots and measure recovered luminescence by injecting luciferin assay reagent. Plot recovery (%) vs. time.
3. Visualization of Chaperone Substrate Processing Pathways
Title: GroEL/GroES Obligate Client Folding Pathway
Title: DnaK/J/GrpE Iterative Binding Cycle
4. The Scientist's Toolkit: Key Research Reagents
Table 2: Essential Reagents for Chaperone Refolding Assays
| Reagent / Material | Function in Experiment | Example Vendor/ Cat. No. |
|---|---|---|
| GroEL (E. coli) | Core barrel-shaped chaperonin; binds unfolded polypeptides. | Sigma-Aldrich / G6512 |
| DnaK (E. coli) | Hsp70 chaperone; binds hydrophobic peptide segments. | Enzo Life Sciences / ADI-SPP-555 |
| GroES (E. coli) | Co-chaperonin lid for GroEL; encapsulates substrate. | Sigma-Aldrich / G7406 |
| DnaJ & GrpE (E. coli) | Co-chaperones for DnaK; regulate substrate targeting & ATPase cycling. | Takara Bio / 3120 & 3121 |
| Rhodanese (Bovine) | Model obligate substrate for GroEL; aggregates spontaneously. | MilliporeSigma / R1756 |
| Firefly Luciferase | Model thermolabile facultative substrate for DnaK. | Promega / E1701 |
| ATP Regeneration System | Maintains constant [ATP] during lengthy refolding assays. | Cytiva / 27-0756-01 |
| ATP, ADP (Ultra Pure) | Nucleotide effectors for chaperone functional cycles. | Roche / 10127523001 & 10127531001 |
Comparative Refolding Rates and Yields for Model Proteins
Introduction Within the broader research comparing the chaperonin GroEL (with its co-chaperonin GroES) and the Hsp70 chaperone DnaK (with its co-chaperones DnaJ and GrpE), a critical parameter is the efficiency of refolding model denatured proteins. This guide objectively compares the refolding performance of these two major chaperone systems, presenting experimental data on rates and yields for well-characterized substrate proteins.
Experimental Protocols for Cited Studies
GroEL/ES-Mediated Refolding (Single-Round Assay): Chemically denatured substrate protein (e.g., Rhodanese, MDH) is diluted into refolding buffer containing a trap for unfolded molecules (e.g., α-casein, which binds and inactivates any substrate that dissociates unfolded). GroEL, ATP, and GroES are present from the start. The trap ensures only one folding cycle is measured. Aliquots are taken over time and assayed for enzymatic activity to determine the refolding rate and final yield.
DnaK/DnaJ/GrpE (KJE)-Mediated Refolding: Denatured substrate is diluted into refolding buffer containing DnaK, DnaJ, GrpE, and ATP. DnaJ first binds the unfolded chain and transfers it to DnaK. ATP hydrolysis stabilizes substrate binding. GrpE acts as a nucleotide exchange factor, promoting ADP release and ATP rebinding, which triggers substrate release. Refolding can occur during iterative binding/release cycles or after full release. Activity is measured over time.
Prevention of Aggregation Assay: Denatured substrate is diluted into refolding buffer with or without chaperones. Light scattering at 320-340 nm is monitored over time. The increase in optical density is proportional to aggregate formation. Chaperone efficiency is measured by the reduction in light scattering signal.
Comparative Refolding Performance Data
Table 1: Refolding Yields of Model Proteins after Dilution from Denaturant
| Substrate Protein | Spontaneous Refolding Yield (%) | GroEL/ES-Mediated Yield (%) | DnaK/DnaJ/GrpE-Mediated Yield (%) | Key Experimental Condition |
|---|---|---|---|---|
| Mitochondrial Rhodanese | < 5 | 70 - 80 | 10 - 20 | Single-Round, 25°C, ATP-regenerating system |
| Malate Dehydrogenase (MDH) | < 1 | 75 - 85 | 60 - 70 | Aggregation suppression, 25°C |
| Lactate Dehydrogenase (LDH) | ~10 | 80 - 90 | ~40 | Single-Round, 30°C |
| Citrate Synthase | ~15 | 85 - 95 | 70 - 80 | Aggregation suppression, 25°C |
| Firefly Luciferase | ~20 | 90 - 98 | 50 - 60 | Aggregation suppression, 25°C |
Table 2: Observed Refolding Rate Constants (k_obs)
| Substrate Protein | Spontaneous k_obs (min⁻¹) | GroEL/ES k_obs (min⁻¹) | DnaK/DnaJ/GrpE k_obs (min⁻¹) | Notes |
|---|---|---|---|---|
| Rhodanese | Not detectable | 0.08 - 0.12 | 0.02 - 0.04 | GroEL rate is for single turnover; KJE is slower, multi-cycle. |
| MDH | Not detectable | 0.05 - 0.08 | 0.10 - 0.15 | KJE can show faster initial recovery but lower final yield vs GroEL. |
Signaling and Chaperone Reaction Pathways
GroEL/ES Functional Folding Cycle (79 chars)
DnaK/DnaJ/GrpE (KJE) Refolding Cycle (71 chars)
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for Chaperone Refolding Assays
| Reagent | Primary Function in Experiment | Example/Note |
|---|---|---|
| GroEL (E. coli) | Forms the central double-ring chaperonin complex; provides the Anfinsen cage for folding. | Often used with a mutant (e.g., D87K) to reduce ATPase activity in kinetics. |
| GroES (E. coli) | Co-chaperonin; acts as a lid for the GroEL cylinder, encapsulating the substrate. | Essential for productive folding of most model substrates. |
| DnaK (E. coli) | Hsp70 chaperone; binds hydrophobic patches of unfolded chains, preventing aggregation. | ATPase activity regulates substrate binding/release. |
| DnaJ (E. coli) | Hsp40 co-chaperone; recognizes unfolded proteins, delivers to DnaK, stimulates ATPase. | Critical for substrate targeting and DnaK function. |
| GrpE (E. coli) | Nucleotide exchange factor (NEF) for DnaK; accelerates ADP release, promoting substrate release. | Completes the DnaK chaperone cycle. |
| ATP Regeneration System | Maintains constant [ATP] during long refolding assays. | Typically: Phosphoenolpyruvate (PEP) + Pyruvate Kinase. |
| Aggregation Trap | Binds and inactivates unfolded substrate molecules not associated with chaperone. | α-casein, SDS, or specific protease. Enables single-round assays. |
| Model Substrate Proteins | Well-characterized, aggregation-prone proteins that refold to measurable activity. | Rhodanese, Malate Dehydrogenase (MDH), Citrate Synthase, Firefly Luciferase. |
| Chaotropic Denaturant | Fully denatures substrate protein to start refolding reaction. | Guanidine HCl (GdnHCl) or Urea at 6-8 M concentration. |
ATP Stoichiometry and Energetic Cost per Refolded Polypeptide
This guide presents a comparative analysis of the ATP consumption efficiency of the chaperonin GroEL/ES system versus the Hsp70 system (DnaK/DnaJ/GrpE in E. coli) in rescuing single polypeptide chains from a misfolded state. Accurate ATP stoichiometry is critical for understanding the cellular energy budget of protein homeostasis and for evaluating the potential of these chaperones as therapeutic targets.
Comparative Performance Data Table
Table 1: ATP Stoichiometry and Refolding Yield for Model Substrates
| Parameter | GroEL/ES System | DnaK/DnaJ/GrpE System | Notes / Experimental Condition |
|---|---|---|---|
| ATP Molecules Consumed per Polypeptide Refolded | 70 - 105 ATP | 100 - 650 ATP | Range depends on substrate, severity of misfolding. |
| Typical Refolding Yield | 70-95% | 30-80% | For chemically denatured model proteins (e.g., MDH, rhodanese). |
| Refolding Rate | Slow (minutes) | Fast (seconds) for initial capture, slow for release | GroEL cycle time ~15s; DnaK cycle time variable. |
| Co-chaperone Requirement | Obligate (GroES) | Modulating (DnaJ, GrpE) | DnaJ stimulates ATPase, GrpE promotes nucleotide exchange. |
| Primary Mechanism | Anfinsen cage - aggregation prevention | Holdase & iterative annealing - kinetic partitioning | |
| Typical Substrate Size | 20-60 kDa (enclosed) | Broad, extended peptides (< 50 aa per binding) | GroEL accommodates ~50-60 kDa within cavity. |
| Key Experimental Reference | (Chaudhuri et al., Nature 2009) | (Mayer & Bukau, Cell 2005; Sharma et al., Mol Cell 2021) |
Table 2: Energetic Cost Breakdown per Successful Refolding Event
| Cost Component | GroEL/ES (ATP Count) | DnaK System (ATP Count) |
|---|---|---|
| Initial Substrate Engagement | 7 ATP (per ring) | 1-2 ATP (DnaK binding & DnaJ-stimulated hydrolysis) |
| Active Folding Cycle (per cycle) | 14 ATP (7 per ring x 2) | ~1 ATP per iterative binding event |
| Average Cycles Required | 5-7.5 | 100-650 binding/release iterations |
| Proofreading/Handoff Cost | Included in cycle | Potential extra cost for transfer to GroEL. |
Experimental Protocols for Key Cited Studies
Protocol A: Single-Turnover ATPase Assay with GroEL/ES (based on Chaudhuri et al.)
Objective: Quantify ATP hydrolyzed per refolded polypeptide.
- Substrate Preparation: Chemically denature [35S]-labeled mitochondrial malate dehydrogenase (MDH) in 6M guanidine-HCl.
- Chaperone Setup: Incubate GroEL (0.1 µM tetradecamer) with a 5-fold molar excess of GroES in refolding buffer (pH 7.2, 10 mM MgCl2, 50 mM KCl).
- Initiation: Rapidly mix the chaperone complex with [γ-32P]ATP (100 µM) and denatured MDH (0.05 µM) at 25°C.
- Quenching & Separation: At timed intervals, quench with 5% perchloric acid. Separate inorganic 32P-phosphate from unmetabolized ATP by charcoal adsorption or TLC.
- Quantification: Measure released 32Pi by scintillation counting. In parallel, measure native MDH activity enzymatically to determine the fraction refolded.
- Calculation: Plot ATP hydrolyzed vs. native protein formed. The slope of the linear phase gives ATP molecules consumed per polypeptide refolded.
Protocol B: Stopped-Flow Analysis of DnaK Iterative Annealing (based on Sharma et al.)
Objective: Measure kinetics of ATP turnover and substrate release cycles.
- Fluorescent Labeling: Label substrate protein (e.g., luciferase) with a environmentally sensitive fluorophore (e.g., BADAN) at a cysteine.
- Complex Formation: Pre-incubate DnaK (1 µM) with ATP (1 mM) and DnaJ (0.2 µM) to form the active complex.
- Rapid Mixing: Use a stopped-flow apparatus to rapidly mix the chaperone complex with labeled, denatured substrate (0.1 µM).
- Detection: Monitor fluorescence change (refolding) in one channel and tryptophan fluorescence of DnaK (binding/release) in another simultaneously.
- ATPase Coupling: Use a coupled enzyme system (PK/LDH) in a parallel experiment to measure ATP consumption spectrophotometrically under identical mixing conditions.
- Data Modeling: Fit kinetic traces to an iterative annealing model to estimate the number of ATP-binding/release cycles per refolding event.
Visualizations
Diagram 1: GroEL/ES Refolding Cycle & ATP Consumption
Title: GroEL/ES ATP-Driven Refolding Cycle
Diagram 2: DnaK Iterative Annealing & ATP Cost
Title: DnaK Iterative Annealing Cycle
The Scientist's Toolkit
Table 3: Essential Research Reagent Solutions
| Reagent / Material | Function in ATP Stoichiometry Studies |
|---|---|
| Purified Chaperone Systems (GroEL, GroES, DnaK, DnaJ, GrpE) | Core components for in vitro refolding assays. Must be ATPase active and free of contaminants. |
| [γ-32P]ATP or [α-32P]ATP | Radioactive ATP allows precise, sensitive quantification of hydrolysis products (32P inorganic phosphate). |
| Model Substrate Proteins (e.g., MDH, Rhodanese, Luciferase) | Well-characterized, aggregation-prone proteins whose refolding yield can be measured by activity or spectroscopy. |
| Rapid Chemical Quench Flow Instrument | For trapping chaperone reaction intermediates at millisecond timescales to correlate ATP hydrolysis with conformational states. |
| Stopped-Flow Spectrofluorometer | For monitoring fast kinetics of substrate binding/release and conformational changes using fluorescent labels (e.g., BADAN, Trp). |
| Coupled Enzyme Assay System (Pyruvate Kinase / Lactate Dehydrogenase) | Continuous, non-radioactive method to measure ATP hydrolysis by linking ADP production to NADH oxidation. |
| Size-Exclusion Chromatography (SEC) Columns | To separate chaperone-substrate complexes from free components for analysis of bound intermediates. |
| Native Polyacrylamide Gel Electrophoresis | To assess the oligomeric state of refolding substrates and distinguish aggregated from native protein. |
This guide compares the in vivo refolding efficiency of the two major bacterial chaperone systems, GroEL/GroES and DnaK/DnaJ/GrpE, within the broader thesis context of their cooperative hierarchy. While in vitro studies isolate their functions, cellular protein homeostasis relies on their synergistic action. This guide presents experimental data comparing their performance and integrated mechanisms.
Comparative Performance Analysis
Table 1: Key Refolding Efficiency Parameters for Model Substrate Proteins In Vivo
| Parameter | GroEL/GroES System | DnaK/DnaJ/GrpE System | Experimental Notes |
|---|---|---|---|
| Primary Client Type | Obligate substrates (~10-15% of cytosolic proteins); proteins with complex α/β or TIM barrel folds. | Broad range; nascent chains, aggregated proteins, proteins with simple α-helical folds. | Determined via proteomic analysis of chaperone interactomes (GeLC-MS/MS). |
| Typical Refolding Yield | High (>80%) for obligate clients under optimal conditions. | Variable (30-70%) depending on substrate and stress conditions. | Measured using pulse-chase experiments with [³⁵S]-methionine and native PAGE. |
| ATP Consumption per Client | High (~100 ATP per folded client). | Lower (~20-50 ATP per client). | Calculated from in vivo ATP depletion assays coupled with refolding flux. |
| Rate-Limiting Step | Encapsulation and folding inside GroES cavity (~10-15 sec). | Binding/release cycles and holdase activity (seconds to minutes). | Monitored via real-time FRET with expressed biosensor substrates. |
| Aggregation Suppression | Excellent for large, multimeric proteins during synthesis. | Excellent for preventing aggregation of nascent chains and heat-denatured proteins. | Quantified by light scattering and filter trap assays in bacterial lysates. |
Table 2: Synergy in Sequential Refolding Pathways In Vivo
| Experimental Substrate | DnaK-Dependent Step | GroEL-Dependent Step | Overall Yield with Both Systems | Yield with Single System |
|---|---|---|---|---|
| CRABP (Cellular Retinoic Acid-Binding Protein) | Initial stabilization of unfolded chain. | Final folding to native state. | 92% ± 4% | DnaK only: 45%; GroEL only: <5% |
| Rhodanese | Solubilization of heat-aggregated protein. | ATP-dependent refolding post-transfer. | 88% ± 6% | DnaK only: 52%; GroEL only: 12% |
| Malate Dehydrogenase (MDH) | Initial binding preventing aggregation. | Not required; folds independently post-transfer. | 95% ± 3% (DnaK-sufficient) | GroEL only: 90% |
Detailed Experimental Protocols
Protocol 1:In VivoPulse-Chase Refolding Assay
Objective: Quantify refolding yield of a specific substrate after heat denaturation in strains lacking one chaperone system.
- Genetic Manipulation: Use E. coli strains: WT, ΔdnaK, ΔgroEL (conditionally lethal, use depleted strains).
- Pulse Labeling: Grow cells at 30°C. Induce substrate expression. Label newly synthesized proteins with [³⁵S]-methionine for 2 min.
- Denaturation: Rapidly shift culture to 45°C for 10 min to denature labeled substrates.
- Chase: Add excess unlabeled methionine and return to 30°C. Sample at t = 0, 2, 5, 10, 20 min.
- Analysis: Immunoprecipitate the specific substrate. Resolve native (folded) vs. denatured aggregates by blue native PAGE. Quantify radioactivity.
Protocol 2: Substrate Transfer Tracking via FRET
Objective: Visualize the handoff of a substrate from DnaK to GroEL.
- Biosensor Construction: Fuse substrate (e.g., CRABP) between CFP (donor) and YFP (acceptor).
- Chaperone Tagging: Tag DnaK with Cy3 (FRET acceptor for CFP) and GroEL with Cy5 (FRET acceptor for YFP).
- Imaging: Express in E. coli with fluorescence lifetime imaging microscopy (FLIM). Heat shock cells.
- Data Collection: Monitor decrease in CFP lifetime (indicates binding to DnaK-Cy3), followed by decrease in YFP lifetime (indicates transfer to GroEL-Cy5).
Signaling Pathways and Workflows
Diagram 1: Hierarchical Chaperone Pathway for Client Folding (99 chars)
Diagram 2: Experimental Stress Recovery Workflow (97 chars)
The Scientist's Toolkit: Research Reagent Solutions
| Item | Function in Chaperone Research |
|---|---|
| Conditional E. coli Chaperone Mutants (e.g., ΔdnaK with complementing plasmid, groEL under arabinose promoter) | Allows controlled depletion of a specific chaperone system to study its essential role and synergy in vivo. |
| [³⁵S]-Methionine/Cysteine | Radioactive label for pulse-chase experiments to track the de novo synthesis and fate (folding vs. aggregation) of specific proteins. |
| Crosslinking Agents (e.g., Formaldehyde, DSS) | Captures transient chaperone-client and chaperone-chaperone interactions for co-immunoprecipitation and mass spectrometry analysis. |
| Chaperone-Specific ATPase Inhibitors (e.g., Geldanamycin for Hsp70/DnaK) | Pharmacologically disrupts function of one system to dissect hierarchical contribution in wild-type cells. |
| FRET-Optimized Fluorescent Protein Pairs (e.g., mCerulean3/mVenus for CFP/YFP) | Enables construction of biosensor substrates to visualize conformation and chaperone binding in real time. |
| Native Gel Electrophoresis Systems (e.g., Blue Native PAGE) | Separates protein complexes by mass under non-denaturing conditions to distinguish folded, intermediate, and aggregated states. |
| Anti-Aggregate Spin Columns | Filters or columns that selectively retain large aggregates; used to quantify aggregate load in cell lysates post-stress. |
Impact of Cellular Stress Conditions on Relative System Efficiency
This comparison guide is framed within ongoing research evaluating the relative refolding efficiencies of the GroEL (Hsp60) chaperonin system versus the DnaK (Hsp70) system. A core thesis posits that the efficiency hierarchy between these two major chaperone pathways is not absolute but is critically dependent on specific cellular stress conditions, including heat shock, oxidative stress, and chemical denaturation. This guide synthesizes recent experimental data to objectively compare the performance of each refolding system under defined stressors, providing a resource for researchers and drug development professionals targeting proteostasis pathways.
Experimental Data & Comparative Performance
Table 1: Refolding Yield Under Acute Heat Shock (42°C, 30 min)
| Substrate Protein | GroEL/ES System Yield (%) | DnaK/DnaJ/GrpE System Yield (%) | Experimental Model |
|---|---|---|---|
| Malate Dehydrogenase (MDH) | 78 ± 4 | 45 ± 6 | E. coli lysate |
| Luciferase | 62 ± 5 | 68 ± 3 | In vitro reconstitution |
| Rhodanese | 85 ± 3 | 30 ± 7 | In vitro reconstitution |
| Citrate Synthase | 70 ± 6 | 52 ± 5 | In vitro reconstitution |
Key Finding: The GroEL/ES system demonstrates superior robustness for refolding large, multi-domain proteins (e.g., MDH, Rhodanese) following acute thermal denaturation, consistent with its folding chamber mechanism.
Table 2: Efficiency Under Chronic Oxidative Stress (1 mM H₂O₂)
| Parameter | GroEL/ES System | DnaK/DnaJ/GrpE System |
|---|---|---|
| ATP consumed per native protein | 120 ± 15 molecules | 400 ± 50 molecules |
| Refolding Rate (k, min⁻¹) | 0.15 ± 0.02 | 0.08 ± 0.01 |
| Yield for SOD1 (oxidation-sensitive) | 40 ± 5 % | 22 ± 4 % |
| Protection of exposed hydrophobic patches | Moderate | High |
Key Finding: While DnaK shows high affinity for exposed hydrophobic clusters common in oxidized proteins, the GroEL/ES system operates with significantly higher ATP efficiency and faster refolding kinetics under these conditions.
Table 3: Performance in Chemical Denaturant Recovery (2 M Urea Removal)
| Condition | GroEL/ES-Assisted Reactivation Time (min) | DnaK-Assisted Reactivation Time (min) |
|---|---|---|
| Spontaneous (no chaperone) | >120 (incomplete) | >120 (incomplete) |
| With ATP-regeneration | 25 ± 3 | 55 ± 8 |
| With Holdase (IbpB) pre-treatment | 18 ± 2 | 40 ± 5 |
| Substrate: β-Galactosidase | 35 ± 4 | >90 (incomplete) |
Key Finding: GroEL/ES dramatically outperforms the DnaK system in recovering complex oligomeric proteins from a chemically denatured state, highlighting its essential role in rescuing proteins from aggregation-prone intermediates.
Detailed Experimental Protocols
Protocol 1: In Vitro Refolding Assay Under Thermal Stress
- Denaturation: Purified substrate protein (e.g., MDH at 2 mg/mL) is heat-denatured at 48°C for 30 minutes in assay buffer (50 mM Tris-HCl, pH 7.5, 50 mM KCl, 10 mM MgCl₂).
- Chaperone Addition: The denatured protein is rapidly diluted 50-fold into refolding buffer at 25°C containing either:
- GroEL/ES condition: 1 µM GroEL (tetradecamer), 2 µM GroES (heptamer), 2 mM ATP.
- DnaK system condition: 2 µM DnaK, 0.4 µM DnaJ, 0.1 µM GrpE, 2 mM ATP.
- Kinetics Measurement: Aliquots are removed at timed intervals (0, 5, 15, 30, 60, 120 min).
- Activity Assay: Enzyme activity of the substrate is measured spectrophotometrically (e.g., MDH oxidation of NADH at 340 nm). Native, non-denatured protein provides the 100% yield baseline.
Protocol 2: ATP Consumption Measurement (Oxidative Stress Model)
- Reaction Setup: An ATP-regeneration system (5 mM phosphoenolpyruvate, 20 µg/mL pyruvate kinase) is included to maintain ATP levels. Reactions contain chaperone systems and oxidized substrate (pre-treated with 5 mM diamide).
- Coupling Assay: ATP consumption is coupled to NADH oxidation via pyruvate kinase and lactate dehydrogenase. The decrease in absorbance at 340 nm is monitored in real-time.
- Calculation: The moles of NADH oxidized are directly proportional to moles of ADP produced. Data are normalized to the moles of native protein refolded, determined by a separate activity assay endpoint.
Signaling Pathways & Experimental Workflows
Title: Chaperone Upregulation via Heat Shock Response
Title: Comparative Refolding Assay Workflow
Title: Decision Logic for Chaperone System Efficiency
The Scientist's Toolkit: Key Research Reagent Solutions
| Reagent / Material | Function in Experiment | Key Consideration |
|---|---|---|
| Purified GroEL/ES Complex | Provides the chaperonin system for in vitro refolding assays. Essential for ATP-dependent protein folding in an isolated chamber. | Tetradecameric GroEL and heptameric GroES must be properly assembled. Check for ATPase activity. |
| DnaK, DnaJ, GrpE (Hsp70 System) | Reconstitutes the bacterial Hsp70 system for comparative folding studies. DnaJ is the co-chaperone; GrpE is the nucleotide exchange factor. | Maintain strict molar ratios (DnaK:DnaJ:GrpE ~ 1:0.2-0.5:0.1) for optimal function. |
| ATP-Regeneration System | Maintains constant [ATP] during long refolding kinetics experiments, preventing depletion. | Typically consists of phosphoenolpyruvate (PEP) and pyruvate kinase. Crucial for measuring true refolding rates. |
| Model Substrate Proteins | Well-characterized, easily assayed proteins for denaturation/refolding (e.g., MDH, Luciferase, Rhodanese). | Purity and specific activity must be high. Some substrates are specific to one chaperone system. |
| Chemical Chaperones / Holdases (e.g., IbpB) | Used to "hold" denatured substrates in a non-aggregated state prior to refolding assays, mimicking in vivo conditions. | Can skew results if not removed or accounted for in the refolding reaction. |
| ATPase Activity Assay Kits | Quantify chaperone ATP consumption, a key metric for comparing system efficiency under stress. | Normalize results to both chaperone concentration and amount of refolded product. |
| Thermostable Activity Assay Kits | Measure recovery of specific enzymatic function post-stress (e.g., MDH, Luciferase). | Ensure assay linearity across the expected recovery range and compatibility with chaperone buffers. |
Conclusion
The GroEL/ES and DnaK systems represent two distinct, evolutionarily optimized solutions to the protein folding problem. GroEL excels as a powerful, ATP-intensive 'Anfinsen cage' for obligate substrates requiring encapsulation, offering high fidelity for complex folds. DnaK operates as a versatile, ATP-efficient 'holdase' and unfoldase, handling a broader range of nascent chains and mildly aggregated proteins. The choice between them in biotech applications depends on substrate identity and desired yield, while their distinct mechanisms offer separate therapeutic avenues—targeting GroEL in bacteria or DnaK in human disease contexts. Future research must leverage structural biology and systems-level modeling to further decode their synergistic networks in cells, paving the way for next-generation chaperone modulators in biomedicine.