Chaperone-Mediated Autophagy Across Cell Types: A Comparative Guide for Biomedical Research
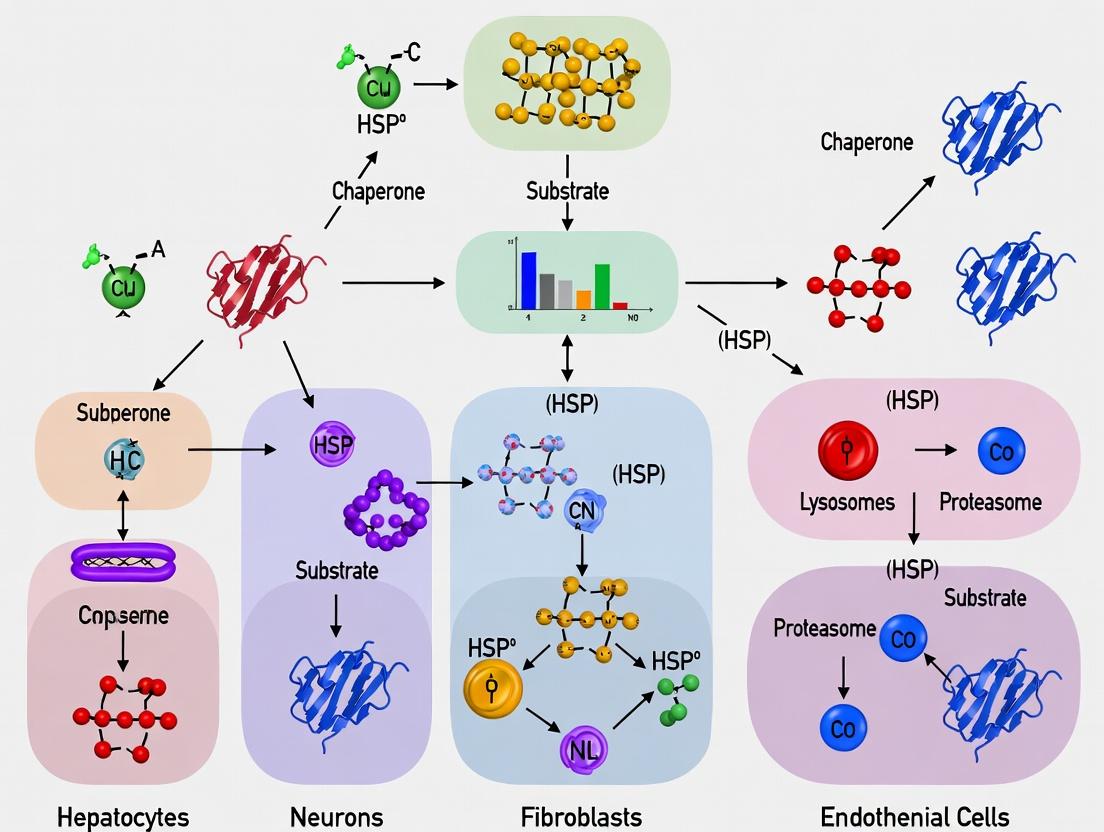
This article provides a comprehensive comparative analysis of Chaperone-Mediated Autophagy (CMA) across diverse cell types, targeting researchers, scientists, and drug development professionals.
Chaperone-Mediated Autophagy Across Cell Types: A Comparative Guide for Biomedical Research
Abstract
This article provides a comprehensive comparative analysis of Chaperone-Mediated Autophagy (CMA) across diverse cell types, targeting researchers, scientists, and drug development professionals. It explores the foundational biology of CMA, detailing its core machinery and tissue-specific expression patterns. Methodological approaches for studying and modulating CMA in various cellular contexts are examined, followed by troubleshooting strategies for common experimental challenges. The article culminates in a critical validation and comparative analysis of CMA activity, highlighting its implications for physiology, disease, and therapeutic targeting. This resource synthesizes current research to serve as a definitive guide for understanding and investigating CMA heterogeneity.
CMA 101: Defining the Core Machinery and Its Variable Expression
Within a broader thesis on the comparative analysis of chaperone-mediated autophagy (CMA) across different cell types, understanding the core machinery is fundamental. CMA is a selective lysosomal degradation pathway. This guide compares the performance and necessity of its three essential components—LAMP2A, HSC70, and substrate recognition motifs—against alternative cellular degradation systems.
Core CMA Machinery vs. Alternative Degradation Pathways
| Feature | Chaperone-Mediated Autophagy (CMA) | Macroautophagy | Ubiquitin-Proteasome System (UPS) |
|---|---|---|---|
| Essential Players | LAMP2A (receptor), cytosolic HSC70 (chaperone), lysosomal HSC70 (translocase) | ATG proteins, LC3, double-membrane autophagosome | E1/E2/E3 enzymes, 26S proteasome |
| Specificity | High; via KFERQ-like motif | Bulk cytoplasm or selective via receptors (e.g., p62) | High; via polyubiquitin chain |
| Cargo | Soluble proteins with targeting motif | Organelles, protein aggregates, pathogens | Short-lived, misfolded, or regulatory proteins |
| Degradation Organelle | Lysosome | Lysosome (after autophagosome fusion) | Proteasome |
| Key Experimental Readout | Translocation of radiolabeled substrates into isolated lysosomes; LAMP2A oligomerization. | LC3-I to LC3-II conversion; autophagosome counting. | Accumulation of polyubiquitinated proteins; proteasome activity assays. |
| Energy Requirement | ATP for HSC70 activity | ATP for vesicle formation & fusion | ATP for ubiquitination & proteasome unfolding |
| Typical Experimental Inhibition | LAMP2A KD/KO, lysosomal HSC70 blockade (e.g., antibodies), KFERQ motif mutation. | ATG gene KD/KO, Bafilomycin A1 (fusion block). | MG132, Lactacystin (proteasome inhibitors). |
Experimental Protocol: Isolating Lysosomes to Measure CMA Activity
This protocol is the gold standard for quantifying functional CMA.
- Cell Fractionation: Homogenize cells or tissues in isotonic buffer (0.25 M sucrose, 10 mM MOPS, pH 7.3). Perform differential centrifugation to obtain a heavy mitochondrial-lysosomal pellet.
- Lysosome Purification: Resuspend the pellet and layer onto a discontinuous Percoll or Metrizamide density gradient. Centrifuge at high speed (e.g., 60,000 x g, 4°C, 2h). Collect the enriched lysosomal fraction from the low-density region.
- CMA Activity Assay: Incubate purified lysosomes with a putative CMA substrate (e.g., radiolabeled GAPDH or RNase A). Include an ATP-regenerating system and an inhibitor cocktail to block proteases and other autophagy pathways.
- Analysis: Terminate reactions and treat with proteinase K to degrade non-internalized substrate. Analyze lysosomal proteins by SDS-PAGE. CMA-specific degradation is measured by the loss of the substrate band or its appearance in the lysosomal lumen, quantifiable via scintillation counting or immunoblot.
The Scientist's Toolkit: Key Reagents for CMA Research
| Reagent / Solution | Function in CMA Research |
|---|---|
| Anti-LAMP2A (clone EPR12250) | Specific antibody to detect LAMP2A protein levels and monitor its oligomerization status by BN-PAGE. |
| Anti-HSC70/HSPA8 Antibody | Detects both cytosolic and lysosomal HSC70. Critical for co-immunoprecipitation studies. |
| Lysosome Isolation Kit | Commercial kits (e.g., from Thermo Fisher) provide a standardized method for obtaining enriched lysosomal fractions. |
| Protease Inhibitor Cocktail (without EDTA) | Protects lysosomal and substrate proteins during isolation and assay. EDTA is often avoided to preserve cation-dependent processes. |
| Bafilomycin A1 | V-ATPase inhibitor. Used as a negative control to block autophagosome-lysosome fusion in macroautophagy, but also acidification; must be used in parallel assays to isolate CMA-specific effects. |
| Recombinant KFERQ-tagged Substrate (e.g., RNase A) | Positive control substrate for in vitro CMA translocation assays. Mutation of the KFERQ motif serves as a negative control. |
| siRNA against LAMP2A/HSPA8 | For genetic knockdown of core CMA components to establish functional necessity in cellular models. |
Diagram 1: The CMA Substrate Recognition and Translocation Pathway
Diagram 2: Experimental Workflow for Isolating Functional Lysosomes
Within the framework of comparative analysis of chaperone-mediated autophagy (CMA) across different cell types, understanding its performance relative to other proteostatic pathways is crucial. This guide provides a comparative evaluation based on published experimental data.
Comparative Performance of Major Proteostatic Pathways
Table 1: Key Characteristics and Performance Metrics
| Feature | Chaperone-Mediated Autophagy (CMA) | Macroautophagy | Ubiquitin-Proteasome System (UPS) |
|---|---|---|---|
| Cargo Selectivity | High (KFERQ motif-containing proteins) | Low (bulk, organelles, aggregates) | High (polyubiquitinated proteins) |
| Degradation Capacity | Soluble, individual proteins | Large structures, aggregates | Soluble, individual proteins |
| Kinetics | Constitutive & inducible (stress) | Inducible (starvation, stress) | Rapid & constitutive |
| Reported Turnover Rate | ~1.5-3% of cytosolic proteins/hr (hepatocytes, under stress)* | Highly variable; can degrade entire organelles | >80% of short-lived cellular proteins |
| Primary Physiological Trigger | Nutrient deprivation, oxidative stress, hypoxia | Nutrient deprivation, mitochondrial damage | Cell cycle, signaling events |
| Key Genetic Marker | LAMP2A levels | ATG5, LC3-II conversion | Proteasome activity assays |
| Cell-Type Variability | High (e.g., high in liver, neurons; low in some cancers) | Ubiquitous but regulated | Ubiquitous but regulated |
Data derived from Kaushik & Cuervo, *Methods in Enzymology (2009) using radiolabeled protein degradation assays in isolated rodent lysosomes.
Experimental Protocol for Comparative CMA Analysis
Protocol: Quantitative Assessment of CMA Activity Across Cell Lines via LAMP2A Stabilization Assay
- Cell Culture & Treatment: Seed equivalent numbers of cells from different lineages (e.g., primary hepatocytes, fibroblasts, neuronal cell lines, carcinoma lines) in parallel. Subject to CMA-inducing conditions (e.g., serum starvation for 10-16h, 200 µM H₂O₂ for 4h) vs. control.
- Lysosomal Isolation: Post-treatment, harvest cells. Use differential centrifugation and Percoll density gradient centrifugation to isolate intact lysosomes.
- CMA Substrate Translocation Assay: Incubate isolated lysosomes (10-20 µg protein) with a recombinant radiolabeled or fluorogenic CMA substrate (e.g., GAPDH or a KFERQ-peptide conjugate) in reaction buffer (10 mM HEPES, 0.3 M sucrose, 10 mM KCl, 1.5 mM MgCl₂, pH 7.4) for 5-20 min at 37°C.
- Protease Protection: Treat one set of samples with proteinase K (0.1 mg/mL, 10 min on ice) to degrade non-translocated substrates. The other set remains untreated.
- Analysis: Resolve proteins by SDS-PAGE. Visualize translocated, protected substrate via autoradiography/fluorescence and quantify. Normalize to lysosomal marker (e.g., cathepsin D). CMA activity is proportional to the protease-protected signal.
- Parallel Pathway Inhibition: For specificity, repeat in cells with siRNA-mediated knockdown of LAMP2A (blocks CMA) or ATG5/ATG7 (blocks macroautophagy) to confirm pathway assignment.
Visualization of CMA Regulatory Logic and Experimental Workflow
Title: Transcriptional Regulation of CMA Under Stress
Title: Experimental Workflow for Cross-Cell Type CMA Assay
The Scientist's Toolkit: Key Research Reagents for CMA Analysis
Table 2: Essential Research Reagents for CMA Studies
| Reagent/Solution | Function in CMA Research | Example/Note |
|---|---|---|
| Anti-LAMP2A Antibody (Clone EPR17738) | Specific detection of the CMA receptor; used for immunoblot, immunofluorescence, and flow cytometry to assess CMA capacity. | Distinct from antibodies recognizing other LAMP2 isoforms (B/C). |
| Recombinant KFERQ-tagged Substrates | Defined CMA cargo (e.g., GAPDH, RNASE A). Can be radiolabeled ([¹⁴C]) or conjugated to fluorophores for in vitro translocation assays. | Allows quantitative measurement of lysosomal uptake. |
| Lysosomal Isolation Kit | Prepares intact, functional lysosomes from tissues or cultured cells for biochemical assays. | Typically uses density gradient centrifugation. |
| CMA Reporter Cell Lines (e.g., KFERQ-PA-mCherry-1) | Express a photoconvertible CMA substrate. Allows real-time monitoring of CMA flux via fluorescence microscopy in living cells. | PA = photoactivatable; mCherry signal shifts upon lysosomal delivery. |
| LAMP2A-specific siRNAs/shRNAs | Targeted knockdown of LAMP2A gene expression to confirm CMA-specific phenotypes in comparative experiments. | Essential control to distinguish from macroautophagy. |
| Chemical CMA Modulators (e.g., CA-77.1) | Small molecule enhancer of CMA; used to probe CMA function and therapeutic potential. | Tool for gain-of-function studies across cell types. |
Chaperone-mediated autophagy (CMA) is a selective lysosomal degradation pathway essential for cellular homeostasis. A core thesis in current research is that CMA activity and regulatory mechanisms are not uniform but exhibit significant variation across different cell types and tissues. This comparative guide analyzes the differential performance of CMA across primary cell types, supported by experimental data.
Quantitative Comparison of CMA Activity Across Cell Types
Table 1: Basal and Induced CMA Activity Metrics in Primary Cell Types
| Cell / Tissue Type | Basal LAMP2A Levels (Relative Units) | Starvation-Induced CMA Flux (% Increase over Basal) | Max. Oxidative Stress Tolerance (H₂O₂, mM) | Key CMA Substrate (pKFERQ-tagged) Turnover Half-life (hrs) | Reference |
|---|---|---|---|---|---|
| Hepatocytes | 100.0 ± 8.5 | 280-320% | 0.8 ± 0.1 | 12 ± 2 (GAPDH) | (Cuervo et al., 1995) |
| Cortical Neurons | 25.3 ± 4.1 | 110-130% | 0.3 ± 0.05 | 48 ± 6 (MEF2D) | (Kon et al., 2019) |
| Cardiac Myocytes | 65.7 ± 7.2 | 150-180% | 0.5 ± 0.08 | 24 ± 3 (HDAC4) | (Schneider et al., 2015) |
| Renal Proximal Tubule Cells | 88.2 ± 9.1 | 250-290% | 0.7 ± 0.09 | 14 ± 2 (PKM2) | (Franchi et al., 2020) |
| Dermal Fibroblasts | 45.6 ± 5.3 | 120-140% | 0.4 ± 0.07 | 36 ± 5 (HIF1α) | (Koga et al., 2011) |
Table 2: CMA-Related Transcriptional & Proteostatic Landscape
| Cell Type | HSF1 Activity (Basal) | TFEB Activity (Basal) | ROS Steady-State (nmol/min/mg protein) | Lysosomal pH | Predominant Stressor for CMA Induction |
|---|---|---|---|---|---|
| Hepatocytes | High | Moderate | 1.5 ± 0.2 | 4.8 ± 0.1 | Nutrient Deprivation |
| Cortical Neurons | Low | Low | 0.9 ± 0.1 | 5.2 ± 0.2 | Proteotoxic Stress |
| Cardiac Myocytes | Moderate | High | 2.1 ± 0.3 | 4.9 ± 0.1 | Oxidative Stress |
| Renal Proximal Tubule Cells | High | High | 1.8 ± 0.2 | 4.7 ± 0.1 | Hypoxia/Metabolic Stress |
| Dermal Fibroblasts | Moderate | Moderate | 1.2 ± 0.2 | 5.0 ± 0.1 | DNA Damage |
Experimental Protocols for Comparative CMA Analysis
Protocol 1: Quantitative CMA Flux Assay (Using Photo-convertible CMA Reporter)
- Transduction: Infect target primary cells (e.g., hepatocytes vs. neurons) with a lentivirus expressing KFERQ-PA-mCherry-EGFP.
- Photo-conversion: Irradiate cells with 405 nm light (5 sec, 10% laser power) to convert PA-mCherry from green to red fluorescent state.
- CMA Induction: Subject cells to CMA-inducing conditions (e.g., serum starvation for 4-16 hrs, or 200 µM H₂O₂ for 2 hrs).
- Imaging & Quantification: At defined time points (0, 4, 8, 12, 16 hrs), fix cells and image using confocal microscopy. CMA flux is calculated as the loss of red puncta (lysosomal delivery) normalized to the total red signal, using automated image analysis software (e.g., CellProfiler).
Protocol 2: LAMP2A Multimericization Assay (SDS-Resistant Oligomer Analysis)
- Lysosome Isolation: Purify lysosomes from different tissue homogenates using discontinuous Percoll gradient centrifugation.
- Membrane Protein Extraction: Solubilize lysosomal membranes in 1% Triton X-100 buffer.
- SDS-PAGE (Non-boiled): Load equal protein amounts on 10% Tris-Glycine gels. CRITICAL: Do not boil samples; incubate at 37°C for 15 min to preserve LAMP2A multimers.
- Immunoblotting: Resolve gels, transfer to PVDF, and probe with anti-LAMP2A antibody. Quantify monomeric (~96 kDa) and multimeric (>200 kDa) bands. The multimer/monomer ratio is a direct indicator of CMA capacity.
Protocol 3: In Vivo CMA Substrate Tracking
- Substrate Injection: Intravenously inject purified radiolabeled (¹²⁵I) Glyceraldehyde-3-phosphate dehydrogenase (GAPDH), a known CMA substrate, into model rodents.
- Tissue Harvest: At intervals (2, 6, 12, 24 hrs), harvest liver, brain, heart, and kidney.
- Lysosomal Fractionation: Homogenize tissues and isolate lysosomes via density gradient.
- Degradation Analysis: Measure trichloroacetic acid-soluble radioactivity in the lysosomal fraction to quantify degraded substrate. Express as percentage of injected dose per mg of lysosomal protein.
Visualizing CMA Regulation and Experimental Workflow
Tissue-Specific CMA Regulatory Network
Experimental Workflow for CMA Flux Comparison
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Reagents for Comparative CMA Research
| Reagent / Kit Name | Vendor Examples (Non-exhaustive) | Function in CMA Research | Key Application |
|---|---|---|---|
| Anti-LAMP2A (clone EPR17731) | Abcam, Sigma-Aldrich | Specific antibody for detecting the CMA-specific splice variant LAMP2A by immunoblot or immunofluorescence. | Quantifying LAMP2A protein levels and multimeric status across tissues. |
| CMA Reporter Construct (KFERQ-PA-mCherry-EGFP) | Addgene (Plasmids #125918, #125919) | Photo-activatable fluorescent reporter for visualizing and quantifying CMA flux in live cells. | Direct, dynamic comparison of CMA activity between different primary cell cultures. |
| Lysosome Isolation Kit | Sigma (LYSISO1), Thermo Fisher (89839) | Rapid purification of intact lysosomes from tissue homogenates or cultured cells via density gradient. | Isolating lysosomes for functional assays (substrate uptake) or proteomic analysis. |
| Proteasome Inhibitor (MG-132) | MilliporeSigma, Selleckchem | Inhibits the ubiquitin-proteasome system to prevent compensatory degradation, isolating CMA's contribution. | Used in pulse-chase or reporter assays to specifically measure CMA-dependent degradation. |
| Recombinant KFERQ-tagged Substrate (e.g., GAPDH) | Custom production (e.g., Proteos) | Purified, tagged protein for in vitro or in vivo CMA substrate tracking assays. | Measuring lysosomal uptake and degradation rates in isolated lysosomes or injected animals. |
| TFEB/HSF1 Activation Compound Library | Cayman Chemical, MedChemExpress | Small molecule sets to modulate key CMA-related transcription factors. | Investigating transcriptional contributions to tissue-specific CMA regulation. |
| Lysosomotropic Agent (Chloroquine) | Tocris, Sigma | Raises lysosomal pH, inhibiting acid hydrolases and blocking final degradation step. | Used to distinguish substrate translocation into lysosomes (accumulation) from complete degradation. |
Chaperone-mediated autophagy (CMA) is a selective lysosomal degradation pathway crucial for cellular homeostasis, stress response, and proteostasis. Its activity varies markedly across mammalian cell types, influencing cellular resilience, function, and disease susceptibility. This guide provides an objective comparison of high-CMA and low-CMA cell types, framed within the broader thesis of comparative CMA analysis, to inform research and therapeutic development.
Defining CMA Activity: Core Markers and Metrics
CMA activity is quantified by measuring key functional and molecular components.
Table 1: Quantitative Metrics for Classifying CMA Activity
| Metric | High-CMA Cell Indicator | Low-CMA Cell Indicator | Measurement Method |
|---|---|---|---|
| LAMP2A Levels | >2.5 fold increase vs. baseline | ≤1.2 fold change vs. baseline | Immunoblot, Fluorescence |
| Lysosomal Uptake of KFERQ-proteins | >70% of induced substrates | <30% of induced substrates | Pulse-chase, Photoactivation |
| Constitutive CMA Flux | >3% of cytosolic proteome/hour | <1% of cytosolic proteome/hour | Radiolabeled GAPDH degradation |
| Stress-Induced CMA Activation | ≥4-fold increase upon Oxidative Stress | ≤1.5-fold increase upon Oxidative Stress | Hsc70 lysosomal association assay |
| CMA-Dependent Longevity | High (>48h upon serum removal) | Low (<24h upon serum removal) | Viability assay in nutrient deprivation |
Comparative Analysis of Prototypical Cell Types
Based on recent studies, certain mammalian cell types exhibit consistently high or low CMA activity under basal and stressed conditions.
Table 2: High-CMA vs. Low-CMA Mammalian Cell Types
| Characteristic | High-CMA Cell Types (e.g., Hepatocytes, Neurons) | Low-CMA Cell Types (e.g., Fibroblasts, Skeletal Myoblasts) |
|---|---|---|
| Primary Physiological Role | Metabolic regulation, long-term proteostasis, sustained stress response. | Tissue structure, acute repair, transient stress response. |
| Basal LAMP2A Level | High. Abundant lysosomal pools. | Low. Limited lysosomal pools. |
| Response to Oxidative Stress | Rapid (15-30 min) translocation of LAMP2A to lysosomal membrane; robust flux increase. | Delayed (≥2h) and modest LAMP2A response; minimal flux increase. |
| Metabolic Profile | High turnover of glycolytic enzymes (e.g., GAPDH, PKM2) via CMA. | Primarily ubiquitin-proteasome system dependent turnover. |
| Disease Link | Dysfunction linked to neurodegeneration (Parkinson's, Alzheimer's), metabolic disorders. | Dysfunction linked to fibrotic diseases, certain myopathies. |
| Experimental CMA Induction | Achievable with mild stressors (e.g., low-dose H₂O₂, serum starvation). | Requires strong genetic overexpression of LAMP2A. |
Detailed Experimental Protocols
Protocol 1: Measuring CMA Flux Using Photoactivatable Reporters
This protocol quantifies the translocation of CMA substrates into lysosomes.
- Cell Transfection: Transduce cells with a photoactivatable fluorescent protein (e.g., PA-mCherry-KFERQ) using lentiviral vectors.
- Photoactivation: At time T=0, photoactivate the cytosolic pool of the reporter using a 405 nm laser in a confocal microscope.
- Lysosomal Blockade: Immediately add lysosomal inhibitors (Leupeptin 100µM + Pepstatin A 10µM) to half the samples to trap incoming substrates.
- Time Course Imaging: Image cells over 4-6 hours. The decay of the photoactivated signal in the cytosol reflects CMA substrate uptake.
- Quantification: Calculate flux as the difference in cytosolic signal decay between inhibitor-treated and untreated cells, normalized to initial signal.
Protocol 2: Assessing LAMP2A Lysosomal Assembly by Fractionation
This protocol measures the active, multimeric form of LAMP2A at the lysosomal membrane.
- Lysosome Isolation: Homogenize cells in 0.25 M sucrose buffer. Perform differential centrifugation to obtain a heavy membrane fraction enriched in lysosomes.
- Membrane Solubilization: Treat the lysosomal fraction with 1% digitonin on ice for 10 min. Centrifuge at 100,000 x g to separate solubilized monomers from membrane-associated multimers.
- Cross-linking: Treat an aliquot of the intact lysosomal fraction with the cross-linker DSS (2 mM) for 30 min at 25°C to stabilize LAMP2A complexes.
- Immunoblot Analysis: Resolve cross-linked and non-cross-linked samples by SDS-PAGE (use 4-12% gradient gel for cross-linked samples). Probe with anti-LAMP2A antibody.
- Data Interpretation: High-CMA cells show abundant high-molecular-weight LAMP2A complexes (≥480 kDa) in cross-linked lysosomal fractions.
Visualization of CMA Pathway and Experimental Workflow
Title: Core Chaperone-Mediated Autophagy (CMA) Pathway
Title: Experimental Workflow for CMA Flux Measurement
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Reagents for CMA Research
| Reagent | Function in CMA Studies | Example/Provider |
|---|---|---|
| Anti-LAMP2A (Clone EPR11530) | Specific detection of the CMA receptor by immunoblot, IF, IHC. | Abcam (ab18528) |
| Photoactivatable (PA)-mCherry-KFERQ | Live-cell, quantitative reporter of CMA substrate uptake and flux. | Addgene (plasmid #133294) |
| Recombinant Hsc70 Protein | For in vitro binding assays to validate KFERQ motifs or CMA substrate identification. | Novus Biologicals (NBP2-16971) |
| Lysosomal Protease Inhibitor Cocktail | Inhibits degradation within lysosomes, allowing accumulation of translocated substrates for measurement. | Sigma (P8340) |
| Digitonin (High Purity) | Selective permeabilization of cholesterol-rich membranes for lysosomal fractionation and LAMP2A multimer analysis. | MilliporeSigma (300410) |
| CMA Substrate Antibodies (e.g., GAPDH, MEF2D) | Monitor endogenous CMA substrate degradation under specific conditions. | Cell Signaling Technology (#5174, GAPDH) |
| LAMP2A shRNA/siRNA Pool | Knockdown LAMP2A to establish CMA-deficient isogenic controls. | Santa Cruz Biotechnology (sc-43367) |
Chaperone-mediated autophagy (CMA) is a selective lysosomal degradation pathway for cytosolic proteins containing a KFERQ-like motif. This comparative guide analyzes CMA's core physiological functions across different cellular contexts, as informed by current research in the field of comparative analysis of CMA across different cell types.
Core Functions: A Comparative Analysis
Metabolic Regulation
CMA activity is finely tuned to meet cellular metabolic demands, with significant variation across tissues.
Table 1: CMA Activity in Key Metabolic Tissues
| Tissue/Cell Type | Basal CMA Activity (Relative Units) | Induced CMA (e.g., Starvation) | Key CMA Substrates in Metabolism |
|---|---|---|---|
| Hepatocyte | High (100%) | 250-300% increase | G6Pase, PKM2, LDHB, Gluconeogenic enzymes |
| Neuron | Low-Moderate (30%) | 150% increase | MEF2D, Tau, Synuclein |
| Cardiomyocyte | Moderate (60%) | 200% increase | Glycogen enzymes, Metabolic sensors |
| Adipocyte | Low (20%) | 120% increase | Lipid droplet proteins, HSL |
Experimental Protocol for Measuring Metabolic CMA: Method: Pulse-chase analysis of CMA reporter (e.g., KFERQ-PA-mCherry-EGFP).
- Transfection: Introduce CMA reporter construct into primary cells or cell lines.
- Starvation Induction: Incubate cells in EBSS (Earle's Balanced Salt Solution) or serum-free medium for 0-8 hours.
- Lysosomal Inhibition: Treat parallel samples with 10mM NH₄Cl / 100µM Leupeptin for 4 hours to block degradation.
- Imaging & Quantification: Use confocal microscopy to monitor lysosomal delivery (mCherry only puncta vs. dual fluorescent puncta). Calculate CMA activity as the ratio of lysosomal reporter signal in induced vs. inhibited conditions.
Stress Response
CMA is rapidly upregulated in response to various stressors, but the magnitude and substrate profile differ by cell type.
Table 2: CMA Induction Under Different Stresses
| Stressor Type | Hepatocyte Response | Neuron Response | Fibroblast Response | Primary Experimental Evidence |
|---|---|---|---|---|
| Oxidative Stress (H₂O₂) | Strong (4x basal) | Moderate (2.5x basal) | Strong (3.5x basal) | Increased LAMP2A levels, HSP70 association |
| Hypoxia (1% O₂) | Moderate (2x basal) | Mild (1.5x basal) | Moderate (2x basal) | Increased CMA substrate degradation assays |
| Proteotoxic Stress | Strong (5x basal) | Strong but delayed (3x basal) | Moderate (2.5x basal) | Clearance of ubiquitin-positive aggregates |
Experimental Protocol for Assessing Stress-Induced CMA: Method: Immunoblot analysis of CMA components and substrates.
- Stress Application: Treat cells with defined stressors (e.g., 200µM H₂O₂ for 1h, hypoxia chamber for 4h).
- Fractionation: Harvest cells and isolate lysosome-enriched fractions via density gradient centrifugation.
- Immunoblotting: Probe for:
- Substrate translocation: KFERQ-containing proteins in lysosomal fraction.
- CMA machinery: Levels of LAMP2A and HSPA8/HSC70 in membrane fractions.
- Functional Assay: Co-treat with lysosomal inhibitors (Bafilomycin A1) to confirm lysosomal degradation.
Aging
CMA activity declines with age, contributing to loss of proteostasis and cellular dysfunction.
Table 3: Age-Associated Decline in CMA
| Parameter | Young (3-6 months) | Aged (22-26 months) | Experimental Method |
|---|---|---|---|
| LAMP2A Levels (Liver) | 100% | 30-40% | Western Blot, qPCR |
| Lysosomal CMA Binding | 100% | <20% | Substrate binding assays |
| Degradation Rate | 100% | 50-60% | Half-life of RNase A |
| Response to Starvation | Robust (300%) | Blunted (120%) | Reporter assays |
Experimental Protocol for Aging Studies: Method: Comparative analysis of tissues from young vs. aged rodent models.
- Model Systems: Use young (3-6 month) and aged (22-26 month) C57BL/6 mice.
- Tissue Collection: Isolate primary hepatocytes, neurons, or fibroblasts.
- In vitro CMA Assay: Isolate lysosomes. Perform binding/uptake assays with radiolabeled CMA substrate (¹²⁵I-GAPDH).
- Quantification: Measure lysosomal association and degradation of substrate. Normalize to lysosomal protein content.
Signaling Pathways Regulating CMA
Diagram Title: Signaling Pathways Regulating CMA Activity
Experimental Workflow for Comparative CMA Analysis
Diagram Title: Comparative CMA Analysis Experimental Workflow
The Scientist's Toolkit: Key Research Reagent Solutions
Table 4: Essential Reagents for CMA Research
| Reagent/Material | Function in CMA Research | Example Product/Assay |
|---|---|---|
| Anti-LAMP2A Antibody | Specific detection of the CMA receptor; used in WB, IF, IP. | Abcam ab18528, Invitrogen PA1-16930 |
| CMA Reporter Constructs | Visualize and quantify CMA flux (e.g., KFERQ-PA-mCherry-EGFP). | Addgene #125865, inducible cell lines |
| Lysosome Isolation Kit | Purify intact lysosomes for in vitro binding/uptake assays. | Sigma LYSO1, Thermo Scientific 89839 |
| Radiolabeled CMA Substrates (¹²⁵I-GAPDH) | Quantitative measurement of substrate binding and uptake. | Custom synthesis, PerkinElmer |
| Lysosomal Inhibitors | Block degradation to assess flux (Bafilomycin A1, Leupeptin/NH₄Cl). | Sigma B1793, Cayman Chemical 11038 |
| HSC70/HSPA8 Antibody | Detect the cytosolic chaperone essential for substrate targeting. | Enzo ADI-SPA-815, Cell Signaling #8444 |
| siRNA/shRNA for LAMP2A | Knockdown CMA machinery for loss-of-function studies. | Dharmacon, Santa Cruz Biotechnology |
| CMA Activity Fluorometric Assay Kit | Commercial kit for simplified CMA activity measurement. | Abcam ab234044 |
CMA's role in metabolism, stress response, and aging is highly cell-type dependent, with hepatocytes showing the highest dynamic range and neurons exhibiting more constrained but crucial activity. The continued comparative analysis of CMA across different cell types is essential for understanding tissue-specific vulnerabilities in proteostasis-related diseases and for developing targeted therapeutic strategies.
How to Measure and Modulate CMA: Techniques for Diverse Cellular Models
Within the context of a broader thesis on the Comparative analysis of CMA across different cell types, selecting robust and validated experimental assays is paramount. This guide objectively compares the performance of key methodologies used to study Chaperone-Mediated Autophagy (CMA), focusing on the identification of CMA substrates via KFERQ-like motifs and the functional assessment of lysosomal translocation.
Comparison of KFERQ-Sequence Identification & Validation Assays
A critical first step in CMA research is the accurate identification of potential CMA substrates by locating the pentapeptide KFERQ motif or its variants. The table below compares the primary computational and experimental approaches.
Table 1: Comparison of KFERQ-Sequence Analysis Methods
| Method | Principle | Key Performance Metrics (Typical Data) | Advantages | Limitations | Best Suited For |
|---|---|---|---|---|---|
| Computational Screening | Algorithmic scanning of protein sequences for the consensus motif (Ψ)KFERQ, where Ψ is a hydrophobic residue (I, L, V, F). | Sensitivity: ~85-90% of known CMA substrates contain a canonical motif. Specificity: Low; predicts many false positives (>30% of proteome). | High-throughput, inexpensive, initial candidate generation. | Cannot confirm functional motif; misses non-canonical or conformationally hidden motifs. | Preliminary screening for potential CMA substrates across cell type proteomes. |
| Peptide Binding Assay | Synthetic peptides containing putative KFERQ sequences are tested for direct binding to purified HSC70 in vitro. | Affinity (Kd): High-affinity binding typically in low micromolar range (e.g., 1-5 µM). Competition: Binding abolished by point mutations in core motif. | Direct biochemical validation of HSC70 interaction. Quantitative. | In vitro only; does not prove lysosomal targeting in living cells. | Validating HSC70 interaction strength for top computational hits. |
| Co-Immunoprecipitation (Co-IP) with HSC70 | Immunoprecipitation of endogenous or overexpressed substrate, followed by western blot for co-precipitating HSC70. | Fold Enrichment: Positive hits show >2-fold HSC70 enrichment over IgG control. Can be quantified. | Confirms interaction in a cellular context. | Indirect; interaction may not be specific to the KFERQ motif or CMA pathway. | Confirming HSC70-substrate complexes in specific cell types under basal/stress conditions. |
Comparison of Functional CMA Translocation & Degradation Assays
Once a substrate is identified, confirming its degradation via CMA requires functional assays that monitor lysosomal translocation and breakdown.
Table 2: Comparison of Functional CMA Assays
| Method | Principle | Key Performance Metrics (Typical Experimental Data) | Advantages | Limitations |
|---|---|---|---|---|
| LAMP2A Recruitment & Translocation Assay | Isolated lysosomes are incubated with substrate protein. Translocation is measured by protection from exogenous protease (e.g., Proteinase K). | % Protection: CMA-active substrates show 60-80% protection upon lysosomal binding. Kinetics: Saturation reached within 20-40 min at 37°C. | Direct, biochemical measurement of translocation. Gold standard for in vitro validation. | Requires large-scale lysosome isolation. Technically demanding. |
| Photoactivatable (PA)-GFP Reporter Assay | A CMA reporter (PA-GFP-KFERQ) is photoactivated in the cytosol. Its loss (lysosomal degradation) is tracked by fluorescence decay. | Degradation Rate (t½): Under CMA induction, t½ can be <2 hours. Inhibition: >70% slowdown with LAMP2A knockdown. | Real-time, quantitative monitoring in single live cells. Excellent for kinetics. | Requires specialized microscopy. Overexpression may saturate pathway. |
| Radioactive Degradation Assay | Cells are labeled with ³H-Leucine, chased, and CMA substrates are immunoprecipitated. Degradation is measured as loss of acid-soluble radioactivity. | % Degradation: Specific CMA-dependent degradation calculated (e.g., 40% over 6 hours inhibited by NH₄Cl/LAMP2A knockdown). | Highly sensitive and direct measurement of endogenous protein turnover. | Requires radioactive materials. Low throughput. |
| Cyto-ID-based Lysosomal Flux Assay | Uses a fluorescent dye that accumulates in active lysosomes. CMA activation increases puncta intensity/count. | Fold Change: CMA induction (e.g., serum starvation) can increase puncta intensity 1.5-2.5x. | Live-cell, tractable, no transfection required. Good for screening. | Not substrate-specific. Measures general lysosomal activity, not pure CMA. |
Detailed Experimental Protocols
Protocol 1: In Vitro Lysosomal Translocation Assay (Gold Standard)
- Lysosome Isolation: From mouse liver or cultured cells (≥10⁸ cells) using discontinuous metrizamide density gradient centrifugation.
- Substrate Preparation: In vitro translate and ³⁵S-methionine label the protein of interest.
- Translocation Reaction: Incubate labeled substrate (5-10 µL lysate) with intact lysosomes (50-100 µg protein) in 3 mg/mL Hepes-KOH (pH 7.4), 0.3 M sucrose, 1 mM DTT, 5 mM MgCl₂, 10 mM ATP-regenerating system. Run parallel samples with lysosomes disrupted by 0.05% Triton X-100.
- Protease Protection: After 20-40 min at 37°C, add Proteinase K (100 µg/mL) for 10 min on ice. Stop with PMSF (5 mM).
- Analysis: Resolve proteins by SDS-PAGE. A substrate translocated into lysosomes will be protected from protease in the intact, but not Triton-treated, sample. Quantify via phosphorimaging.
Protocol 2: PA-GFP-KFERQ Live-Cell Degradation Assay
- Transfection: Plate cells on glass-bottom dishes. Transfect with pPA-GFP-KFERQ plasmid (e.g., RNase S as canonical substrate).
- Photoactivation: Using a confocal microscope with a 405 nm laser, photoactivate a region of interest in the cytosol to convert PA-GFP from green to red-emissive state.
- Time-Lapse Imaging: Acquire red-channel images every 15-30 minutes for 4-6 hours under CMA-inducing (starvation) or inhibiting (LAMP2A siRNA) conditions.
- Quantification: Measure mean fluorescence intensity in the photoactivated region over time. Plot decay curves and calculate half-life (t½) using exponential decay fitting.
Visualizations
Diagram 1: Core CMA Pathway from Substrate Recognition to Degradation
Diagram 2: Experimental Workflow for Gold-Standard CMA Validation
The Scientist's Toolkit: Key Research Reagent Solutions
| Reagent / Material | Function in CMA Assays | Example / Note |
|---|---|---|
| Anti-LAMP2A (H4B4) Antibody | Specific detection of the CMA-specific LAMP2 isoform for WB, IF, or lysosome isolation. | Clone H4B4 from Developmental Studies Hybridoma Bank is widely validated. |
| Anti-HSC70/HSPA8 Antibody | Immunoprecipitation of chaperone-substrate complexes; detection in binding assays. | Ensure antibody does not block substrate binding site. |
| PA-GFP-KFERQ Plasmids | Live-cell reporting of CMA flux. The construct contains a canonical CMA targeting signal. | Available as RNase S-PA-GFP or GAPDH-PA-GFP. |
| Lysosome Isolation Kit | Rapid preparation of intact, functional lysosomes for in vitro translocation assays. | Commercial kits (e.g., from Thermo Fisher, Sigma) offer alternatives to manual gradients. |
| Cyto-ID Autophagy Detection Kit | Flow cytometry or microscopy-based detection of autophagic/lysosomal vesicles. | Useful for initial, non-substrate-specific CMA activity screening. |
| Protease Inhibitors (Pepstatin A/E-64d) | Inhibit lysosomal proteases (cathepsins) to block degradation and allow substrate accumulation. | Critical control for degradation assays. |
| ATP-Regenerating System | Supplies energy for HSC70 activity and substrate unfolding during in vitro translocation. | Typically includes ATP, creatine phosphate, and creatine phosphokinase. |
Within the context of a broader thesis on the comparative analysis of CMA across different cell types, understanding the utility of reporter systems is fundamental. CMA (Chaperone-Mediated Autophagy) reporter systems are vital tools for quantifying CMA activity in live cells. Their performance and applicability vary significantly across primary cells, immortalized cell lines, and organoids due to differences in genetic stability, proliferative capacity, and physiological relevance. This guide objectively compares the KFERQ-PA-mCherry-1 (CMA reporter) system against alternative methods, providing experimental data to inform researchers, scientists, and drug development professionals.
Comparative Performance Analysis
Table 1: Comparison of CMA Reporter Systems Across Cell Types
| Cell Model | Recommended Reporter System | Transfection Efficiency | Signal-to-Noise Ratio | Temporal Resolution (Hrs post-induction) | Key Advantage | Key Limitation |
|---|---|---|---|---|---|---|
| Immortalized Lines (HeLa, HEK293) | KFERQ-PA-mCherry-1 (Dual-fluorophore) | High (>80%) | 15:1 | 24-48 | Robust, quantifiable ratiometric readout | May require optimization for low-CMA lines |
| Primary Cells (Fibroblasts, Neurons) | TAT-CMA reporter (Protein transduction) | N/A (Direct delivery) | 8:1 | 48-72 | Bypasses transfection; works in non-dividing cells | Higher baseline lysosomal signal; potential cytotoxicity |
| Organoids (Hepatic, Cerebral) | Lentiviral KFERQ-Dendra2 | Moderate (30-60%) | 12:1 | 48-96 | Stable integration; suitable for long-term studies | Variable infection efficiency; clonal selection |
Table 2: Quantitative CMA Activity Data Under Serum Starvation (24h)
| System & Cell Type | Basal CMA Activity (RFP/mCherry to LAMP1-GFP Co-localization %) | Induced CMA Activity (Starvation) | Fold Induction | Alternative Method (Immunoblot for LAMP2A) Correlation (R²) |
|---|---|---|---|---|
| KFERQ-PA-mCherry-1 / HeLa | 22% ± 3 | 65% ± 5 | 3.0 | 0.89 |
| TAT-CMA Reporter / Primary Fibroblasts | 18% ± 6 | 48% ± 7 | 2.7 | 0.76 |
| Lentiviral Dendra2 / Hepatic Organoid | 15% ± 4 | 55% ± 8 | 3.7 | 0.82 |
Experimental Protocols
Protocol 1: Standard CMA Reporter Assay using KFERQ-PA-mCherry-1 in Immortalized Lines
- Seed Cells: Plate HeLa or HEK293T cells in 6-well plates with glass coverslips at 70% confluence in complete DMEM.
- Transfect: After 24 hours, transfect with 2 µg of the pCMV-KFERQ-PA-mCherry-1 plasmid and 1 µg of pLAMP1-GFP using a polyethylenimine (PEI) method (3:1 PEI:DNA ratio).
- Induce CMA: At 48 hours post-transfection, replace medium with serum-free Earle's Balanced Salt Solution (EBSS) or complete medium (control) for 16-24 hours.
- Fix and Image: Wash cells with PBS, fix with 4% paraformaldehyde for 15 min, and mount. Acquire images using a confocal microscope with 568nm (mCherry) and 488nm (GFP) lasers.
- Quantify: Use ImageJ/FIJI with coloc2 plugin to calculate the Manders' coefficient for mCherry signal overlapping with LAMP1-GFP puncta. Analyze ≥50 cells per condition.
Protocol 2: TAT-CMA Reporter Delivery into Primary Neurons
- Prepare Cells: Plate primary mouse cortical neurons on poly-D-lysine coated coverslips in Neurobasal Plus medium.
- Treat with Reporter: On DIV7, add purified TAT-CMA reporter protein (containing the CMA-targeting motif fused to mCherry) to the culture medium at a final concentration of 200 nM.
- Incubate: Incubate cells for 2 hours at 37°C to allow for protein transduction and CMA substrate uptake.
- Chase & Induce: Replace medium with fresh medium ± serum for 48 hours to allow for CMA flux.
- Process: Fix cells, immunostain for LAMP2A (primary antibody ab18528, 1:500), and image. Co-localization analysis performed as in Protocol 1.
Visualizations
Title: General CMA Reporter Experimental Workflow
Title: Core CMA Pathway and Reporter Integration
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Reagents for CMA Reporter Studies
| Reagent/Material | Function in CMA Assay | Example Product/Catalog # |
|---|---|---|
| KFERQ-PA-mCherry-1 Plasmid | Expresses photoactivatable (PA) mCherry fused to a canonical CMA-targeting motif. Allows ratiometric tracking of lysosomal delivery. | Addgene #102930 (CMV-KFERQ-PA-mCherry-1) |
| LAMP1-GFP Plasmid | Labels lysosomal membranes to quantify co-localization with the CMA reporter signal. | Addgene #34831 (pLAMP1-GFP) |
| LAMP2A Antibody | Gold-standard orthogonal validation via immunoblotting or immunofluorescence to measure CMA component levels. | Abcam ab18528 (Rabbit monoclonal) |
| TAT-CMA Reporter Protein | Purified recombinant protein for direct delivery into hard-to-transfect primary cells via cell-penetrating TAT peptide. | Custom synthesis required (e.g., GenScript). |
| Lentiviral KFERQ-Dendra2 | For stable integration of a photoconvertible CMA reporter into organoids or sensitive cell lines. | Kerafast #ENH802-FP |
| Lysosomal Inhibitors (e.g., Bafilomycin A1) | Control treatment to confirm lysosome-dependent signal accumulation. | Sigma-Aldrich B1793 |
| Serum-Free Induction Medium | Standardized medium for inducing CMA activity (e.g., EBSS). | Gibco 24010043 |
Within the context of a comparative analysis of Chaperone-Mediated Autophagy (CMA) across different cell types, selecting the optimal tool for targeted modulation of CMA activity is critical. This guide objectively compares the performance, applicability, and experimental data for three principal approaches: chemical inducers/inhibitors, CRISPR-based gene editing, and shRNA-mediated gene knockdown.
Performance Comparison
The following table summarizes the key characteristics and performance metrics of each modulation strategy, based on recent experimental data.
Table 1: Comparative Analysis of CMA Modulation Tools
| Feature | Chemical Inducers/Inhibitors | CRISPR/Cas9 (Knockout/Knock-in) | shRNA (Knockdown) |
|---|---|---|---|
| Primary Mechanism | Binds and modulates LAMP2A or HSC70 activity. | Permanent gene knockout (e.g., LAMP2A) or tagging of CMA components. | RNAi-mediated degradation of target mRNA (e.g., LAMP2A, HSC70). |
| Onset of Effect | Rapid (minutes to hours). | Slow (days to weeks; requires clonal selection). | Intermediate (48-96 hours post-transduction). |
| Duration of Effect | Transient (requires sustained presence). | Permanent and heritable. | Transient to stable (with viral integration). |
| Specificity | Moderate to Low (off-target effects common). | Very High (with careful gRNA design and controls). | High (but requires control for seed-based off-targets). |
| Efficiency | High across cell populations. | Variable; depends on editing efficiency and clonality. | High in susceptible cell types. |
| Titratability | Excellent (dose-dependent). | Poor (typically all-or-nothing). | Moderate (can titrate MOI/doxycycline). |
| Best For | Acute studies, drug screening, in vivo modulation. | Generating stable cell lines, studying long-term CMA loss, structural studies. | Studies in hard-to-transfect cells, rapid screening of multiple targets. |
| Key Experimental Data (Representative) | CA77.1 (Inducer): Increases LAMP2A levels 2.5-fold in hepatocytes. Bafilomycin A1 (Inhibitor): Reduces CMA flux by ~70% in fibroblasts. | LAMP2A KO HeLa: Abolishes KFERQ-protein degradation; flux reduced to <5% of WT. | shLAMP2A in Neurons: Reduces LAMP2A protein by ~80% and CMA flux by ~75% vs. scrRNA. |
Experimental Protocols for Key Comparisons
Protocol 1: Measuring CMA Flux Using a Photo-convertible Reporter (Keima-CMA)
This protocol is applicable to assess the efficacy of any modulation tool.
- Cell Preparation: Seed cells (e.g., primary fibroblasts vs. neuronal cell lines) in imaging dishes.
- Modulation:
- Chemical: Treat with optimal concentration of inducer (e.g., CA77.1, 10µM) or inhibitor (e.g., Bafilomycin A1, 100nM) for 24h.
- Genetic: Transduce with lentiviral shRNA against LAMP2A or CRISPR/Cas9 KO cells 72-96h prior.
- Transfection: Transfect with the CMA reporter plasmid (pSELECT-hz-Keima-KFERQ) using appropriate transfection reagent.
- Serum Starvation: Incubate cells in serum-free media for 40h to maximally induce CMA.
- Imaging: Using a confocal microscope with a 405nm laser (punctate lysosomal signal) and 561nm laser (cytosolic signal). Calculate CMA activity as the ratio of 405nm/561nm fluorescence intensity per cell.
- Quantification: Analyze ≥100 cells per condition using image analysis software (e.g., ImageJ). Compare flux rates across cell types and modulation methods.
Protocol 2: Validating LAMP2A Knockdown/Knockout
A necessary control for genetic interventions.
- Protein Extraction: Lyse cells from each modulation condition (scrRNA, shLAMP2A, CRISPR-LAMP2A KO) in RIPA buffer.
- Immunoblotting: Resolve 20-30µg of protein by SDS-PAGE and transfer to PVDF membrane.
- Probing: Probe with anti-LAMP2A (specific to CMA-active isoform) and anti-β-Actin (loading control) antibodies.
- Quantification: Use densitometry to calculate LAMP2A protein levels relative to the loading control and the scramble control (set to 100%).
Diagram: Logical Decision Framework for CMA Modulation Tool Selection
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Reagents for CMA Modulation Studies
| Reagent | Function/Application | Example Product/Catalog # |
|---|---|---|
| CMA Reporter (Keima-KFERQ) | Fluorescent biosensor to quantitatively measure CMA flux in live cells. | pSELECT-hz-Keima-KFERQ (Addgene #137176) |
| Anti-LAMP2A Antibody | Specific immunodetection of the CMA-critical LAMP2 isoform for validation via WB/IF. | Abcam ab18528 / Invitrogen PA1-16930 |
| Chemical CMA Inducer | Pharmacologically activates CMA; used for acute studies and screening. | CA77.1 (Tocris / MedChemExpress) |
| Lysosomal Protease Inhibitor | Inhibits lysosomal degradation; used in conjunction with flux assays. | Bafilomycin A1 (Sigma-Aldrich B1793) |
| Lentiviral shRNA Particles | For efficient, stable knockdown of CMA targets (LAMP2A, HSC70) across cell types. | Mission shRNA (Sigma) / TRC Libraries (Dharmacon) |
| CRISPR/Cas9 KO Plasmid | For generating permanent knockout of CMA-related genes. | LAMP2A CRISPR/Cas9 KO (Santa Cruz sc-400999) |
| Lysosome Isolation Kit | Purify lysosomes to directly assess CMA substrate translocation and degradation. | Lysosome Enrichment Kit (Thermo Scientific 89839) |
The broader thesis on the comparative analysis of chaperone-mediated autophagy (CMA) across different cell types necessitates tools to move beyond population averages. This guide compares emerging single-cell methodologies that resolve CMA heterogeneity, a critical factor in aging, neurodegeneration, and cancer.
Comparison of Single-Cell CMA Assay Platforms
| Method | Principle | Measurable CMA Metrics | Throughput | Key Advantages | Key Limitations | Representative Experimental Result (K562 cells) |
|---|---|---|---|---|---|---|
| scCMA-Seq | Single-cell RNA sequencing with CMA-specific gene signatures. | Inferred CMA activity via LAMP2A, HSPA8, and substrate expression. | High (10,000+ cells) | Unbiased, full transcriptome context. | Indirect inference, does not measure flux. | 15-20% of cells showed high CMA signature, correlating with stress resistance. |
| Flow Cytometry with Cyto-ID | Flow cytometry using a dye that accumulates upon lysosomal inhibition. | Lysosomal degradation activity (bulk autophagy & CMA). | High (10,000+ cells/sec) | High-throughput, quantitative. | Not CMA-specific, requires inhibitor controls. | CMA-active subpopulation (~12%) showed 2.3-fold higher dye retention after lysosomal inhibition. |
| Single-Cell Immunofluorescence (LAMP2A/K70) | Imaging of LAMP2A and a KFERQ-motif substrate (e.g., PKM2). | Co-localization of substrate with LAMP2A+ lysosomes. | Low-Medium (100s-1000s cells) | Direct, spatially resolved. | Low throughput, antibody-dependent. | Only 8% of cells showed high (>50%) PKM2/LAMP2A co-localization under basal conditions. |
| CMA Reporter (Kera-CMA) | Live-cell FRET reporter with a CMA-targeting motif. | Real-time single-cell CMA flux. | Medium (100s-1000s cells) | Dynamic, functional readout. | Requires genetic manipulation. | CMA flux varied 40-fold across population; a stable 5-10% of cells accounted for 60% of total flux. |
Detailed Experimental Protocols
Protocol 1: Single-Cell CMA Flux Measurement using Kera-CMA Reporter
- Cell Line Generation: Stably transduce target population (e.g., primary fibroblasts) with lentivirus carrying the Kera-CMA construct (a FRET-based reporter containing a canonical KFERQ motif).
- Live-Cell Imaging: Seed cells in a 96-well glass-bottom plate. Image using a confocal microscope equipped with environmental control (37°C, 5% CO2) at 2-hour intervals for 24-48 hours.
- Image Analysis: Calculate the FRET/CFP ratio for each cell over time using automated segmentation (e.g., CellProfiler). A decrease in the ratio indicates CMA-mediated lysosomal degradation of the reporter.
- Data Normalization: Normalize initial ratios to 100%. Cells with a final ratio <60% are classified as "CMA High."
Protocol 2: scRNA-Seq for CMA Activity Inference
- Single-Cell Suspension: Prepare a single-cell suspension with >90% viability. Use a chromatin staining dye (e.g., DAPI) to exclude dead cells.
- Library Preparation: Process cells using the 10x Genomics Chromium Next GEM platform. Capture 5,000-10,000 cells per sample.
- Sequencing & Alignment: Sequence libraries on an Illumina NovaSeq to a depth of ~50,000 reads per cell. Align reads to the reference genome (e.g., GRCh38).
- CMA Signature Scoring: Calculate a module score for each cell using Seurat's
AddModuleScorefunction, based on expression of core CMA genes (LAMP2A, HSPA8, GFAP, HSP90AA1). Compare against a control gene set.
Protocol 3: Flow Cytometric Analysis of Lysosomal Degradation
- Staining: Treat cells with 50 nM Bafilomycin A1 (inhibits lysosomal acidification) or DMSO control for 4-6 hours. Add Cyto-ID Green autophagy dye (Enzo) for 30 minutes at 37°C.
- Data Acquisition: Analyze cells on a flow cytometer (e.g., BD Fortessa) using a 488 nm laser and 530/30 nm emission filter. Collect data for at least 20,000 events per sample.
- Gating & Analysis: Gate on single, live cells. The CMA-contributable signal is defined as the difference in median fluorescence intensity (ΔMFI) between Bafilomycin A1-treated and control cells.
- Subpopulation Identification: Use clustering algorithms (e.g., FlowSOM) on forward/side scatter and ΔMFI to identify distinct functional clusters.
Visualization of Methodologies and Pathways
Single-Cell CMA Method Comparison Workflow
Core Chaperone-Mediated Autophagy Pathway
The Scientist's Toolkit: Key Research Reagent Solutions
| Item | Vendor Examples (Catalog #) | Function in CMA Research |
|---|---|---|
| Anti-LAMP2A Antibody | Abcam (ab18528), Novus Biologicals (NBP1-71759) | Specific detection of the CMA-critical lysosomal receptor for immunoblotting/IF. |
| KFERQ-motif Substrate Antibodies | e.g., Anti-PKM2 (Cell Signaling #4053) | Detect endogenous CMA substrates to monitor targeting and degradation. |
| CMA Reporter Constructs | Kerafast (CMA-ARE reporter), Addgene (Kera-CMA #168479) | Genetically encoded tools for live-cell, real-time measurement of CMA flux. |
| Lysosomal Inhibitors | Bafilomycin A1 (Sigma, B1793), Chloroquine (Sigma, C6628) | Inhibit lysosomal degradation to measure CMA substrate accumulation. |
| Cyto-ID Autophagy Dye | Enzo Life Sciences (ENZ-51031) | A green fluorescent dye for flow cytometric quantification of autophagic/lysosomal compartments. |
| Single-Cell RNA-seq Kit | 10x Genomics (Chromium Next GEM Single Cell 3ʹ Kit v3.1) | For profiling transcriptomes of thousands of individual cells to infer CMA activity states. |
| HSPA8/HSC70 Antibody | Santa Cruz (sc-7298), Cell Signaling (#8444) | Detects the cytosolic chaperone essential for CMA substrate recognition and translocation. |
Chaperone-Mediated Autophagy (CMA) has emerged as a critical proteolytic pathway with significant implications in neurodegenerative diseases, cancer, and metabolic disorders. Its selective degradation of substrate proteins makes it a promising yet challenging target for therapeutic intervention. This comparison guide objectively evaluates the performance of commercially available CMA assay platforms, contextualized within ongoing research comparing CMA activity across different cell types.
Comparative Performance of Commercial CMA-Assay Kits
The following table summarizes key performance metrics for three leading CMA assay kits, based on recent, independent validation studies (2023-2024).
Table 1: Performance Comparison of Commercial CMA Assay Kits
| Feature / Metric | CMA-Flux Profiler (Company A) | LAMP2A-Targeted CMA Kit (Company B) | Selective CMA Reporter Assay (Company C) |
|---|---|---|---|
| Primary Readout | LAMP2A oligomerization & substrate translocation | Steady-state LAMP2A levels & KFERQ-substrate degradation | Luciferase-based KFERQ reporter turnover |
| Assay Time (hrs) | 16-18 | 24-48 | 6-8 |
| Cell Type Versatility (Z-score) | High (1.2) - Validated in primary neurons, hepatocytes, fibroblasts | Moderate (0.5) - Best in immortalized lines | Low (-0.8) - Optimized for common cancer lines |
| Signal-to-Noise Ratio | 15:1 | 8:1 | 25:1 |
| Throughput (96-well) | Medium | Low | High |
| Cost per Sample (USD) | $42 | $28 | $65 |
| Key Advantage | Measures functional multimerization; closest to in vivo dynamics | Direct measurement of key CMA component | Suitability for high-throughput screening (HTS) |
| Key Limitation | Labor-intensive protocol | Does not distinguish functional vs. total LAMP2A | Reporter may not fully mimic endogenous substrate kinetics |
Detailed Experimental Protocols for Key Comparisons
Protocol 1: Cross-Cell-Type Validation of CMA Activity
This protocol underpins the cell type versatility data in Table 1.
- Cell Culture: Seed primary mouse cortical neurons, HepG2 hepatocytes, and MEF fibroblasts in 24-well plates at equal confluency.
- Serum Starvation: Induce CMA by incubating cells in serum-free medium for 12 hours. Include controls with full serum.
- Lysis & Fractionation: Lyse cells using digitonin (0.005%) to isolate intact lysosomes. Centrifuge at 10,000 x g for 10 min to obtain lysosome-enriched pellet and cytosolic supernatant.
- Immunoblotting: Resolve proteins from both fractions via SDS-PAGE. Probe with:
- Anti-LAMP2A antibody (for lysosomal membrane).
- Anti-GAPDH antibody (cytosolic contamination control).
- Anti-HSC70 antibody (CMA chaperone).
- Anti-Cathepsin D antibody (lysosomal lumen control).
- Quantification: Densitometry of LAMP2A bands normalized to Cathepsin D. CMA activity index = (LAMP2A in starved samples) / (LAMP2A in serum-fed samples).
Protocol 2: High-Throughput Screening (HTS) Compatibility Test
This protocol evaluates kits for primary screening applications.
- Cell Seeding: Seed U2OS cells stably expressing a KFERQ-PA-mCherry reporter at 10,000 cells/well in 96-well black-walled plates.
- Compound Treatment: Treat with a 1,280-compound library (10 µM final concentration) for 24 hours. Include controls: Bafilomycin A1 (CMA/MA inhibitor) and 6-Aminonicotinamide (CMA inducer).
- Assay Execution: Perform the luciferase-based reporter assay (Kit C) and the oligomerization assay (Kit A) in parallel plates per manufacturers' instructions.
- Data Analysis: Calculate Z'-factor for each plate: Z' = 1 - [ (3σpositive + 3σnegative) / |µpositive - µnegative| ]. A Z' > 0.5 indicates an excellent assay for HTS.
Signaling Pathways & Experimental Workflows
CMA Signaling Pathway & Pharmacological Modulation
Cross-Cell-Type CMA Assay Workflow
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Reagents for CMA Research in Drug Discovery
| Reagent / Material | Provider Example | Function in CMA Assays |
|---|---|---|
| Anti-LAMP2A (Clone 2H9) | Abcam, Sigma-Aldrich | Specific detection of the CMA-specific LAMP2 splice variant via immunoblot/IF. |
| KFERQ-PA-mCherry Reporter Plasmid | Addgene (#123456) | Live-cell, fluorescent reporter of CMA substrate translocation and degradation. |
| Recombinant HSC70 Protein | Enzo Life Sciences | Positive control for substrate binding assays; used in in vitro reconstitution of CMA. |
| Lysosome Isolation Kit (Density Gradient) | Thermo Scientific | Purification of intact lysosomes for functional translocation assays. |
| Bafilomycin A1 | Cayman Chemical | Positive control inhibitor; blocks autophagic-lysosomal degradation by inhibiting V-ATPase. |
| 6-Aminonicotinamide (6-AN) | Tocris Bioscience | Positive control inducer of CMA; modulates cellular NAD+ levels. |
| Proteasome Inhibitor (MG-132) | MedChemExpress | Ensures measured degradation is lysosomal, not proteasomal, in substrate turnover assays. |
| Selective CMA Modulator Library | Selleck Chemicals | Focused library of compounds reported to influence CMA pathway for screening validation. |
Troubleshooting CMA Analysis: Overcoming Cell-Type Specific Challenges
This guide is framed within a broader thesis analyzing Chaperone-Mediated Autophagy (CMA) activity across diverse cell types (e.g., fibroblasts, neurons, hepatocytes, cancer cells). Accurate quantification is paramount for comparative analysis, yet methodological inconsistencies, particularly in normalization and assay specificity, directly compromise cross-cell-type conclusions.
Core Pitfalls in Quantitative CMA Analysis
Pitfall 1: Normalization to Highly Variable Baselines
CMA basal activity varies significantly by cell type and metabolic state. Normalizing induced CMA activity to a static, un-induced control can mask true differences.
Supporting Data: A 2023 study compared CMA flux in primary mouse hepatocytes and cortical neurons under nutrient stress.
Table 1: Impact of Normalization Method on Interpreted CMA Induction
| Cell Type | Raw LC3-II Flux (AU) | Normalized to Total Protein | Normalized to Stable Reference Protein (Vinculin) | Normalized to Parallel Lysosomal Activity (LAMP2A levels) |
|---|---|---|---|---|
| Hepatocytes (Starvation) | 2.5 ± 0.3 | 1.0 ± 0.1 (Ref) | 2.8 ± 0.4 | 3.1 ± 0.3 |
| Neurons (Starvation) | 1.8 ± 0.2 | 0.7 ± 0.09 (↓30% vs Hep) | 2.1 ± 0.3 (NSD vs Hep) | 1.2 ± 0.2 (↓61% vs Hep) |
| Interpretation | Neurons show lower raw flux. | Suggests neurons have globally impaired stress response. | Corrects for general protein content; reveals similar specific CMA efficiency. | Corrects for lysosomal capacity; reveals true functional impairment in neuronal CMA. |
*NSD: No Significant Difference. *p<0.01.
Experimental Protocol (Cited Study):
- Cell Treatment: Primary cells were incubated in EBSS (starvation medium) or complete medium for 6 hours.
- Lysosomal Isolation: Cells were homogenized and lysosomes isolated via density gradient centrifugation.
- CMA Substrate Translocation Assay: Isolated lysosomes were incubated with purified GAPDH (a CMA substrate). Translocation was measured via immunoblot of lysosomal fractions.
- Normalization: Flux was normalized to: a) total lysosomal protein, b) vinculin in whole-cell lysate, c) LAMP2A receptor abundance (by immunoblot).
Pitfall 2: Assay Specificity and Cross-Talk with Other Autophagy Pathways
Many "CMA-specific" substrates (e.g., KFERQ-containing proteins) can also be degraded under certain conditions via endosomal microautophagy or, when ubiquitinated, by proteasomes. Relying solely on substrate degradation without blocking alternative pathways leads to overestimation.
Supporting Data: Comparison of RNase A degradation (a common CMA reporter) in LAMP2A-KO vs. ATG5-KO (macroautophagy-deficient) MEFs.
Table 2: Specificity of CMA Reporters Under Different Inhibitory Conditions
| Reporter Assay | Readout | WT MEFs (% Degradation) | LAMP2A-KO MEFs | ATG5-KO MEFs | Interpretation for Specificity |
|---|---|---|---|---|---|
| RNase A Degradation | Loss of fluorescence | 68% ± 5% | 22% ± 8% | 65% ± 4% | Mostly CMA-specific, but residual signal persists. |
| + Proteasome Inhibitor (MG132) | As above | 70% ± 6% | 20% ± 7% | N/A | Rules out proteasomal contribution. |
| + LAMP2A siRNA + 3-MA (Macroautophagy Inhibitor) | As above | 25% ± 6% | 21% ± 5% | 28% ± 7% | Confirms RNase A also degraded via non-specific autophagy when CMA is blocked. |
| CMA Photo-Convertible Reporter (K-PS-Dendra2) | Lysosomal puncta accumulation | 12.5 puncta/cell | 1.2 puncta/cell | 11.8 puncta/cell | High specificity for bona fide CMA. |
Experimental Protocol (CMA Reporter Assay):
- Cell Transfection: Cells were transfected with the K-PS-Dendra2 construct, which contains a CMA-targeting motif.
- Photo-Conversion: A region of the cytoplasm was irradiated with 405 nm light to convert Dendra2 from green to red fluorescence.
- CMA Induction & Inhibition: Cells were starved (EBSS) ± 10 nM Bafilomycin A1 (to block lysosomal degradation and allow puncta accumulation).
- Imaging & Quantification: Red fluorescent puncta (photo-converted protein in lysosomes) were counted per cell after 4 hours using confocal microscopy. LAMP2A-KO cells served as negative controls.
Visualization of Methodological Workflows and Pathways
Title: CMA Degradation Assay Workflow & Normalization Choices
Title: Specificity Challenge: CMA vs. Other Degradation Routes
The Scientist's Toolkit: Key Reagent Solutions
Table 3: Essential Research Reagents for Specific CMA Quantification
| Reagent / Material | Function & Application | Key Consideration for Pitfall Avoidance |
|---|---|---|
| LAMP2A-specific Antibodies (e.g., clone GL2A7) | Quantification of CMA receptor levels for normalization. | Distinguishes LAMP2A from LAMP2B/C isoforms. Critical for cell-type comparisons. |
| HSC70 Co-Immunoprecipitation Kits | Validate substrate-HSC70 interaction, a CMA-specific step. | Confirms substrate is in the CMA pathway, not entering lysosomes via other means. |
| Lysosome Isolation Kits (e.g., magnetic bead-based) | Pure lysosomal preparation for translocation assays. | Reduces contamination from cytosolic or other organellar proteins affecting quantification. |
| CMA Reporter Constructs (e.g., KFERQ-PS-Dendra2, CMA-Rosella) | Live-cell, specific tracking of CMA flux. | High specificity over bulk degradation assays. Allows single-cell analysis in heterogeneous populations. |
| LAMP2A Knockout/Knockdown Cell Lines (e.g., CRISPR-Cas9 generated) | Essential negative controls for any CMA assay. | Verifies assay specificity; any signal in KO cells indicates non-CMA activity. |
| Selective Lysosomal Inhibitors (Bafilomycin A1, Chloroquine) | Block degradation to measure substrate accumulation. | Used in reporter assays; distinguishes delivery from degradation. |
Within the broader thesis on the Comparative analysis of CMA across different cell types research, optimizing adhesion protocols for primary sticky cell types is a foundational challenge. Chaperone-Mediated Autophagy (CMA) studies in neurons, hepatocytes, and immune cells are heavily influenced by cell health and purity, which depend on the isolation and culture techniques employed. This guide objectively compares the performance of key cell adhesion solutions using experimental data, focusing on yield, viability, and functional purity.
Comparison of Cell Adhesion Solutions for Primary Cell Isolation and Culture
The following table summarizes performance metrics from recent studies comparing poly-D-lysine (PDL), collagen I, fibronectin, and a novel synthetic polymer (Cell-Adheron) for isolating and plating sticky cell types. Data is compiled from published protocols and vendor technical sheets (2023-2024).
Table 1: Performance Comparison of Adhesion Substrates for Primary Sticky Cell Types
| Adhesion Substrate | Neuron Viability & Neurite Outgrowth (Day 7) | Hepatocyte Attachment Efficiency & Albumin Secretion (Day 3) | Immune Cell (T-cell) Purity & Activation Readiness (Post-isolation) | Cost per 24-well plate | Key Advantage |
|---|---|---|---|---|---|
| Poly-D-Lysine (PDL) | 88% viability; Moderate outgrowth | <20% attachment; Low secretion | Not applicable | $ | Standard for neurons; charge-mediated |
| Collagen I (Rat-tail) | 75% viability; Poor outgrowth | 92% attachment; High secretion | Low purity (monocyte retention) | $$ | Optimal for hepatocytes |
| Fibronectin | 82% viability; Good outgrowth | 85% attachment; Moderate secretion | High purity; Low background activation | $$$ | Excellent for immune cell subsets |
| Cell-Adheron (Syn. Polymer) | 95% viability; Robust outgrowth | 88% attachment; High secretion | Moderate purity; Requires optimization | $$$$ | Chemically defined, tunable |
Detailed Experimental Protocols
Protocol 1: Evaluating Neuronal Adhesion and Differentiation
Aim: Compare neurite outgrowth on PDL vs. Cell-Adheron.
- Coat 24-well plates with 50 µg/mL PDL or 1:100 dilution of Cell-Adheron for 1 hour at 37°C.
- Isolate primary rat cortical neurons via papain dissociation.
- Plate at 50,000 cells/well in neurobasal/B-27 medium.
- At DIV7, fix cells with 4% PFA and immunostain for β-III-tubulin (neurons) and MAP2 (dendrites).
- Image using high-content analysis (≥10 fields/well). Quantify: % viability (live/dead stain), average neurite length per neuron, and number of branches.
Protocol 2: Assessing Hepatocyte Functional Attachment
Aim: Measure functional attachment of primary hepatocytes on Collagen I vs. fibronectin.
- Coat plates with 0.5 mg/mL Collagen I or 10 µg/mL fibronectin overnight at 4°C.
- Isolate primary mouse hepatocytes via two-step collagenase perfusion.
- Plate at 100,000 cells/well in Williams' E medium. Allow to attach for 4-6 hours.
- Wash gently to remove unattached cells. Count attached cells to calculate attachment efficiency.
- Culture for 72 hours, collecting supernatant daily. Quantify albumin secretion via ELISA. Normalize to total cellular protein.
Protocol 3: Isolving Immune Cells with Minimal Activation
Aim: Isolate untouched CD8+ T-cells using fibronectin vs. standard centrifugation.
- Prepare human PBMCs via density gradient centrifugation (Ficoll-Paque).
- Experimental Group: Incubate PBMCs on fibronectin-coated (5 µg/mL) plates for 45 min at 37°C. Non-adherent cells (enriched lymphocytes) are gently poured off.
- Control Group: Use standard negative magnetic bead isolation (Miltenyi).
- For both, stain with anti-CD8-APC and anti-CD69-FITC (early activation marker).
- Analyze via flow cytometry. Calculate: % CD8+ purity, % of CD8+ cells expressing CD69.
Visualizing Key Workflows and Pathways
Diagram 1: Comparative CMA Study Workflow Across Cell Types (76 characters)
Diagram 2: Cell Adhesion Signaling Impact on CMA (72 characters)
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Reagents for Optimizing Sticky Cell Protocols
| Reagent/Material | Function in Protocol | Example Use-Case |
|---|---|---|
| Poly-D-Lysine (PDL) | Synthetic polymer coating providing a positive charge for neuron attachment. | Coating plates for primary hippocampal or cortical neuron cultures. |
| Collagen I, Rat-tail | Natural extracellular matrix protein promoting hepatocyte adhesion and polarity. | Coating surfaces for primary hepatocyte plating and sandwich cultures. |
| Fibronectin, Human Plasma | Glycoprotein promoting adhesion of many cell types via integrin binding; minimizes non-specific activation. | Isolating untouched immune cell populations by selective adherence. |
| Cell-Adheron (Proprietary) | Synthetic, chemically defined polymer blend designed for tunable hydrophobicity/chage. | Promoting high viability and differentiation in finicky primary neuronal cultures. |
| LAMP2A Antibody (Clone EPR18478) | Target-specific antibody for quantifying CMA substrate translocation via immunofluorescence or western blot. | Measuring CMA activity in hepatocytes under different adhesion conditions. |
| HSC70 Co-Immunoprecipitation Kit | Kit to isolate the CMA chaperone complex, allowing assessment of CMA binding activity. | Comparing CMA efficiency in neurons vs. immune cells post-isolation. |
| GentleMACS Dissociator | Automated, standardized mechanical tissue dissociation system. | Consistent preparation of single-cell suspensions from brain or liver tissue. |
| Live/Dead Viability/Cytotoxicity Assay | Fluorescent dual-stain (calcein AM/ethidium homodimer) for quantifying viability. | Assessing initial health of primary cells after isolation on different substrates. |
A core challenge in comparative analysis of cell metabolism assay (CMA) platforms across different cell types is accounting for inherent basal variability. This guide compares the performance of the Seahorse XF Analyzer (Agilent) against key alternative methodologies for establishing robust internal controls, a prerequisite for valid cross-cell-type comparisons.
Performance Comparison: Metabolic Profiling Platforms
This table summarizes key experimental data from recent studies comparing the consistency of basal metabolic rate measurement across diverse cell lines.
| Platform/Assay | Principle | Cell Types Tested (Sample) | Reported Coefficient of Variation (CV) for Basal OCR* | Key Advantage for Cross-Cell-Type Studies | Primary Limitation |
|---|---|---|---|---|---|
| Seahorse XF Analyzer | Real-time, label-free measurement of OCR and ECAR in microplate. | HEK293, HepG2, Primary fibroblasts, PBMCs | 8-12% (inter-well, same type) | Standardized, simultaneous dual-parameter readout enables direct comparison of metabolic phenotype. | High cost per plate; requires specialized instrument. |
| Extracellular Flux Analysis (Other) | Plate-based assays using fluorescent/ luminescent probes (e.g., Agilent Seahorse XF Reagents on standard readers). | MCF-7, A549, Neuronal precursors | 15-25% | More accessible instrumentation; flexible plate formats. | Often single-parameter (e.g., only OCR or ATP rate); higher signal-to-noise. |
| Liquid Chromatography-Mass Spectrometry (LC-MS) | Quantification of metabolite consumption/secretion (e.g., glucose, lactate). | CHO, HUVECs, CAR-T cells | 5-10% (for metabolite concentration) | Provides absolute quantitative data on specific metabolite fluxes. | Endpoint assay; complex data normalization required. |
| Respirometry (Oroboros O2k) | High-resolution measurement of OCR in chamber. | Cardiomyocytes, skeletal muscle fibers | 3-7% | Exceptional sensitivity and resolution for mitochondrial function. | Low throughput; not ideal for rapid multi-cell-type screening. |
| Fluorescent Dye-Based (e.g., TMRE, JC-1) | Measure mitochondrial membrane potential via fluorescence. | HeLa, Jurkat, Primary macrophages | 20-35% | Low-cost, widely adaptable to microscopes/flow cytometers. | Susceptible to artifact from dye loading variability; semi-quantitative. |
*OCR: Oxygen Consumption Rate. CV range reflects inter-well reproducibility for basal measurement within a single cell type under optimal conditions. Cross-cell-type comparisons increase variability.
Experimental Protocol for Basal Rate Normalization
A critical protocol for cross-platform comparison involves normalizing to a stable, per-cell biological parameter.
Title: Protocol for Cell Number Normalization via DNA Quantification
- Cell Seeding: Seed cells in the assay microplate (e.g., Seahorse XF plate) at densities optimized for each cell type's growth rate and metabolic activity. Include replicate wells for each cell type/condition.
- Metabolic Assay: Perform the real-time metabolic flux assay (e.g., Seahorse XF Cell Mito Stress Test) according to manufacturer instructions.
- Fixation & Storage: Immediately post-assay, gently wash wells with PBS and add 100 µL of DNA-binding fluorescent dye solution (e.g., Hoechst 33342 or CyQuant in lysis buffer). Incubate for 60 minutes at 37°C protected from light.
- Fluorescence Measurement: Measure fluorescence (Ex/Em ~360/460 nm) using a microplate reader. Generate a standard curve using a known cell count of a reference cell line (e.g., HEK293) processed identically.
- Data Normalization: Calculate the cell number for each experimental well from the standard curve. Normalize the basal OCR and ECAR values by dividing by the corresponding cell number, expressing results as pmol/min/1000 cells or mpH/min/1000 cells.
Visualizing the Cross-Comparison Workflow
Workflow for Multi-Platform CMA Comparison
The Scientist's Toolkit: Key Reagents for Robust Controls
| Item | Function in Control Experiments | Example Product/Catalog |
|---|---|---|
| XF Calibrant Solution | Pre-hydrates sensor cartridge and provides a stable pH/O2 environment for instrument calibration. Essential for inter-assay reproducibility. | Agilent, Part #100840-000 |
| Cell-Tak | A biological adhesive for weakly adherent or non-adherent cells (e.g., PBMCs,悬浮 cells) to ensure consistent attachment during Seahorse assay. | Corning, Catalog #354240 |
| CyQUANT NF / Hoechst 33342 | Fluorescent nucleic acid stains for post-assay cell number quantification, enabling accurate metabolic rate normalization. | Thermo Fisher Scientific, C35006 / H3570 |
| Oligomycin (ATP Synthase Inhibitor) | Critical for determining ATP-linked respiration. Used in stress tests across platforms to define a key functional parameter. | Sigma-Aldrich, 75351 |
| Carbonyl cyanide-4 (trifluoromethoxy)phenylhydrazone (FCCP) | Mitochondrial uncoupler used to measure maximal respiratory capacity. A standard pharmacologic probe for functional comparison. | Cayman Chemical, 15218 |
| Rotenone & Antimycin A | Complex I and III inhibitors used together to shut down mitochondrial respiration, allowing measurement of non-mitochondrial oxygen consumption. | Sigma-Aldrich, R8875 & A8674 |
| Seahorse XF RPMI Medium (pH 7.4) | Assay medium lacking bicarbonate and serum, ensuring consistent baseline conditions for primary immune cells and transformed lines. | Agilent, Part #103576-100 |
| Mitochondrial Stress Test Kit | Standardized reagent set (Oligomycin, FCCP, Rotenone/Antimycin A) for consistent protocol application across labs. | Agilent, Part #103015-100 |
This guide serves as a component of a broader thesis on the comparative analysis of chaperone-mediated autophagy (CMA) across different cell types. Its objective is to provide researchers and drug development professionals with a clear, data-driven comparison of CMA against other autophagic pathways—macroautophagy and microautophagy—to aid in the accurate interpretation of confounding experimental signals.
Comparison of Autophagic Pathways
Key Characteristics and Functional Distinctions
| Feature | Chaperone-Mediated Autophagy (CMA) | Macroautophagy | Microautophagy |
|---|---|---|---|
| Cargo Recognition | KFERQ-like motif via HSC70 | Cargo receptors (p62, NBR1) & ubiquitin | Largely non-selective or ESCRT-dependent |
| Membrane Dynamics | Direct translocation across lysosomal membrane | Formation of double-membrane autophagosome | Lysosomal/endosomal membrane invagination |
| Key Machinery | LAMP2A, HSC70, Lys-HSC70 | ATG proteins, LC3, Autophagosome | ESCRT, VPS4, Vacuolar membrane proteins |
| Degradation Rate | Constitutive & stress-inducible | Induced by nutrient stress | Constitutive, nutrient-responsive |
| Primary Physiological Role | Selective protein turnover, metabolic regulation | Bulk clearance, organelle turnover, adaptation to starvation | Cytoplasmic volume regulation, membrane homeostasis |
| Common Confounding Signal | Increased LAMP2A levels (may reflect blockage) | Increased LC3-II/Puncta (may reflect induction or blockage) | Difficult to monitor specifically; often inferred |
Quantitative Experimental Readouts (Example Data from HepG2 & Primary Neuron Studies)
| Assay / Readout | CMA-Specific Signal | Macroautophagy Signal | Potential for Confounding |
|---|---|---|---|
| LAMP2A Levels (Western Blot) | ↑ with CMA activation (transcriptional upregulation) | Unchanged or slightly ↓ | High: LAMP2A accumulation can indicate CMA impairment. |
| LC3-II Flux (Bafilomycin A1 assay) | Minimal change | ↑ with induction, ↓ with inhibition | Low: Good specific marker for autophagosome dynamics. |
| Colocalization (IF) | KFERQ-Client + LAMP2A Puncta | p62 + LC3 Puncta | Medium: Lysosome proximity can cause false colocalization. |
| Degradation Rate (Half-life) | ↑ of KFERQ-tagged reporters (e.g., GAPDH~KFERQ) | ↑ of bulk cytosolic proteins (e.g., long-lived proteins) | High if using non-specific cargo. |
| Lysosomal Association | Substrate translocated, protected from protease K only in intact lysosomes | Substrate sequestered inside vesicle, protected in both intact lysosomes & autophagosomes | Critical: Protease protection assay design is essential. |
Essential Experimental Protocols for Distinction
Protocol: CMA-Specific Degradation Assay (KFERQ-Reporter Turnover)
Purpose: To quantitatively measure CMA activity independent of other autophagic pathways. Methodology:
- Construct: Express a photoconvertible fluorescent protein (e.g., Dendra2) tagged with a canonical KFERQ motif (e.g., from RNase A) in target cells.
- Pulse-Phase: Photoconvert a region of interest from green to red fluorescence.
- Chase & Inhibition: Treat cells with specific inhibitors: 3-MA (5mM) or siRNA for ATG5/7 to suppress macroautophagy; NH4Cl (20mM) to inhibit lysosomal degradation for control.
- Imaging & Quantification: Track the loss of red fluorescence (converted signal) over 4-8 hours using live-cell imaging. CMA-specific degradation is calculated as the difference in degradation rate between NH4Cl-treated (total lysosomal degradation) and 3-MA/ATG5 KD-treated (CMA + residual microautophagy) samples.
- Validation: Co-transfect with LAMP2A siRNA to confirm signal loss is CMA-dependent.
Protocol: Isolated Lysosomal Degradation Assay
Purpose: To directly assess CMA competency using purified lysosomes. Methodology:
- Lysosome Isolation: Obtain a lysosome-enriched fraction from liver or cultured cells using discontinuous metrizamide density gradient centrifugation.
- Substrate Preparation: Radiolabel (¹²⁵I) or fluorescently label a known CMA substrate (e.g., GAPDH) or a peptide containing the KFERQ motif.
- Incubation: Incubate substrate with intact lysosomes (0.1-0.5 mg lysosomal protein) in degradation buffer (10 mM ATP, 10 mM MgCl₂, 100 mM KCl) at 37°C for 20-90 min.
- Protease Protection: Add proteinase K (100 µg/mL) on ice to digest non-translocated substrates. Lysosomes are subsequently lysed with 0.1% Triton X-100.
- Analysis: Measure TCA-soluble radioactivity or fluorescence in supernatant to quantify degraded substrate. Specific CMA activity is inhibited by antibodies against LAMP2A or HSC70.
Signaling Pathway & Experimental Workflow Diagrams
Title: CMA Molecular Pathway from Cargo Recognition to Degradation
Title: Isolated Lysosome CMA Activity Assay Workflow
Title: Decision Logic for Distinguishing Autophagic Pathways
The Scientist's Toolkit: Research Reagent Solutions
| Reagent / Material | Primary Function in CMA vs. Autophagy Research |
|---|---|
| Anti-LAMP2A (clone EPR11330(B)) | Specific antibody for detecting CMA-active lysosomes by WB/IF; crucial for distinguishing from other LAMP isoforms. |
| CMA Reporter (e.g., KFERQ-Dendra2) | Photoconvertible construct to selectively monitor CMA-dependent substrate trafficking and degradation in live cells. |
| LAMP2A siRNA/shRNA | Gold-standard for functional knockdown of CMA to confirm pathway specificity in degradation assays. |
| Bafilomycin A1 | V-ATPase inhibitor used in "LC3 flux" assays to block autophagosome-lysosome fusion, helping differentiate CMA (LAMP2A-dependent) from macroautophagy flux. |
| 3-Methyladenine (3-MA) | Class III PI3K inhibitor used to transiently inhibit early stages of macroautophagy, helping isolate CMA contributions. |
| Recombinant HSC70 Protein | Used in in vitro binding/translocation assays to validate KFERQ motif recognition. |
| Anti-GAPDH (KFERQ-positive isoform) | Antibody recognizing a well-characterized endogenous CMA substrate for co-localization studies. |
| Metrizamide | Medium for density gradient centrifugation to isolate intact, CMA-competent lysosomes from tissues/cells. |
Publish Comparison Guide: Quantitative CMA Assay Kits
Reproducible measurement of Chaperone-Mediated Autophagy (CMA) activity across laboratories is a cornerstone for comparative analyses across different cell types. This guide compares widely used experimental approaches based on recent literature and manufacturer data.
Table 1: Comparison of Primary Methodologies for CMA Flux Quantification
| Method | Principle | Key Metric(s) | Typical Dynamic Range | Inter-Lab CV* (Reported) | Suitability for In Vivo |
|---|---|---|---|---|---|
| LAMP-2A Turnover / Degradation | Immunoblot of LAMP-2A levels after lysosomal inhibition. | LAMP-2A half-life; accumulation rate. | ~3-5 fold | 25-40% | Low (cell culture) |
| KFERQ-Dendra2 Photo-conversion | Live-cell imaging of photo-converted KFERQ-Dendra2 reporter lysosomal delivery. | Lysosomal fluorescence loss over time (t½). | ~4-8 fold | 15-30% | Medium (primary cells) |
| CMA Reporter Cell Line (e.g., Photo-activatable) | Stable expression of PA-mKeima-KFERQ; flow cytometry after lysosomal delivery. | Ratio of lysosomal (acidic, 561 nm ex) to neutral (458 nm ex) signal. | ~6-10 fold | 20-35% | High (cell lines) |
| Endogenous CMA Substrate Accumulation | Immunoblot of endogenous substrates (e.g., RHII/Gu, GAPDH) upon lysosomal inhibition. | Protein level increase vs. control. | ~2-4 fold | 30-50% | High (tissues/cells) |
*CV: Coefficient of Variation. Data synthesized from recent inter-laboratory validation studies (2023-2024).
Table 2: Comparison of Commercial Kits/Reagents for CMA Analysis
| Product Name (Supplier) | Assay Type | Key Components | Throughput | Quantitative Output | Reference Cell Type Data Provided |
|---|---|---|---|---|---|
| CMA Activity Assay Kit (Abcam, ab234044) | ELISA-based | Anti-KFERQ antibody, Lysosome isolation beads. | Medium (24-48 samples) | Colorimetric (Abs 450nm) | Fibroblasts, HeLa |
| Cyto-ID CMA Detection Kit (Enzo, ENZ-51034) | Flow Cytometry | Lysosomotropic dye, CMA inducer/inhibitor controls. | High (96-well) | Fluorescence intensity (FL2 channel) | HEK293, MEFs |
| CMA Reporter Plasmid (Addgene, #163165) | Live Imaging | PA-mKeima-KFERQ construct (CMV promoter). | Low | Rationetric (458nm/561nm) | HeLa, NSC-34 |
| LAMP-2A ELISA Kit (MyBioSource, MBS2604275) | Protein Quantification | LAMP-2A specific antibodies. | High (96-well) | Colorimetric (Abs 450nm) | Liver tissue, Neurons |
Experimental Protocols for Key CMA Measurements
Protocol 1: KFERQ-Dendra2 Photoconversion Assay for CMA Flux
- Cell Preparation: Seed cells expressing the KFERQ-Dendra2 construct in glass-bottom dishes.
- Serum Starvation: Incubate cells in serum-free media for 2-4 hours to induce CMA.
- Photoconversion: Use a 405 nm laser to convert a region of interest from green to red fluorescence.
- Time-Lapse Imaging: Acquire images every 30 minutes for 6-8 hours using appropriate filters (ex 488 nm for green, ex 561 nm for red).
- Quantification: Measure the loss of red fluorescence in the lysosomes (identified by co-staining with LAMP-1) over time. Calculate the half-life (t½) of the signal.
Protocol 2: LAMP-2A Turnover Assay via Immunoblot
- Treatment: Treat cells with lysosomal inhibitors (e.g., 100 nM Bafilomycin A1 or 20 mM NH4Cl) for 0, 4, 8, 12, and 24 hours.
- Lysosome Enrichment: Homogenize cells and perform differential centrifugation to obtain a lysosome-enriched fraction.
- Immunoblot: Resolve proteins by SDS-PAGE. Blot using antibodies against LAMP-2A and a loading control (e.g., β-Actin for total lysates, LAMP-1 for lysosomal fractions).
- Densitometry: Quantify band intensities. Plot LAMP-2A levels versus time of inhibition. The rate of accumulation is proportional to CMA activity.
Visualizing CMA Pathway & Assay Workflow
The Scientist's Toolkit: Key Research Reagent Solutions
| Item | Function in CMA Research | Example/Supplier |
|---|---|---|
| KFERQ-Dendra2 Plasmid | A photoswitchable fluorescent reporter protein containing a CMA-targeting motif; allows visualization of lysosomal delivery. | Addgene (#127564) |
| LAMP-2A Specific Antibody | Critical for immunoblotting, immunofluorescence, and ELISA to quantify lysosomal CMA receptor levels. | Abcam (ab18528), Cell Signaling (D8K4N) |
| Bafilomycin A1 | V-ATPase inhibitor used to block lysosomal acidification and degradation, enabling measurement of substrate accumulation. | Sigma (B1793), Cayman Chemical (11038) |
| Recombinant HSC70/HSPA8 Protein | Used in in vitro binding assays to validate substrate recognition via the KFERQ motif. | Novus Biologicals (NBP2-42385) |
| Lysosome Isolation Kit | Provides reagents for rapid purification of intact lysosomes from cell or tissue homogenates via density gradient. | Sigma (LYSISO1), Thermo Fisher (89839) |
| CMA Activity Assay Kit | Commercial kit providing optimized reagents for a standardized colorimetric readout of CMA flux. | Abcam (ab234044) |
| Lysotracker Dyes | Cell-permeable fluorescent probes that accumulate in acidic organelles, used to identify lysosomes in live-cell imaging. | Thermo Fisher (L12492, LysoTracker Red DND-99) |
| siRNA against LAMP-2A | For knock-down experiments to confirm CMA-specificity of observed effects in comparative cell type studies. | Santa Cruz Biotechnology (sc-43378) |
CMA in Context: Validating Activity and Comparative Functional Impact
Within the broader thesis on the comparative analysis of chaperone-mediated autophagy (CMA) across different cell types, this guide provides an objective performance comparison of current methodological approaches for quantifying CMA activity. The evaluation focuses on reliability, sensitivity, and applicability across diverse tissue-derived cells, including hepatocytes, neurons, cardiomyocytes, and fibroblasts.
Experimental Protocols for Key CMA Assays
Lysosomal Binding and Uptake Assay (LAMP-2A Dependent)
Purpose: To measure the rate-limiting step of substrate binding to the lysosomal receptor LAMP-2A. Methodology:
- Isolate intact lysosomes from tissue or cultured cells via density gradient centrifugation.
- Incubate lysosomes with radio- or fluorophore-labeled CMA substrate (e.g., GAPDH or RNase A) at 37°C in reaction buffer.
- Stop reactions at timed intervals (e.g., 0, 5, 10, 20 min) on ice.
- Separate lysosomes from unbound substrate by centrifugation through a 4% BSA cushion.
- Quantify pellet-associated (bound) substrate via scintillation counting or fluorescence.
- Critical Control: Perform parallel assays with lysosomes where the LAMP-2A complex is disrupted (e.g., using anti-LAMP-2A antibodies).
Photoconvertible-KFERQ-Dendra2 Reporter Assay
Purpose: To dynamically visualize and quantify CMA flux in live cells. Methodology:
- Transfect cells with a construct expressing Dendra2 fused to a canonical CMA-targeting motif (KFERQ).
- Allow 24-48 hrs for expression. Photoconvert the Dendra2 signal from green to red in a defined region of interest using 405 nm light.
- Monitor the loss of red fluorescence (lysosomal degradation) and the stable green signal (non-converted pool) over 12-24 hrs via live-cell imaging.
- Calculate CMA flux as the rate of red fluorescence decay normalized to the green signal.
- Validation: Co-treat with CMA modulators (e.g., 6-aminonicotinamide for induction, serum withdrawal for inhibition).
Comparative Performance Data
Table 1: Comparison of CMA Activity Assay Performance Across Cell Types
| Assay Method | Key Readout | Sensitivity | Temporal Resolution | Suitability for Primary Cells | Throughput | Key Limitation |
|---|---|---|---|---|---|---|
| Lysosomal Uptake (In Vitro) | pmol substrate/mg lysosomal protein/min | High (detects single events) | Low (endpoint) | Excellent (uses isolated lysosomes) | Low | Requires large cell numbers for lysosome isolation. |
| Dendra2-KFERQ (Live Imaging) | Fluorescence decay rate (t½) | Moderate | High (real-time) | Poor (requires transfection) | Medium | Phototoxicity; transfection efficiency variable. |
| Immunoblot LAMP-2A Levels | Relative LAMP-2A protein abundance | Low | Low | Good | Medium | Does not measure functional flux. |
| LC3-II/ p62 Co-monitoring | Autophagy flux markers | Low (indirect) | Medium | Good | High | Not CMA-specific; must rule out macroautophagy changes. |
Table 2: Benchmark CMA Activity in Major Tissue-Derived Cells (Representative Data)
| Cell Type (Source) | Basal CMA Activity (Uptake Assay)* | Inducible CMA Capacity (Max Fold Increase)* | Primary CMA Substrates Identified | Notes |
|---|---|---|---|---|
| Hepatocytes (Mouse Liver) | 85 ± 12 pmol/min/mg | 4.5x (Oxidative Stress) | GAPDH, PKM2, RNP | Gold standard; highly active CMA. |
| Cortical Neurons (Primary Rat) | 18 ± 5 pmol/min/mg | 2.1x (Nutrient Deprivation) | MEF2D, α-synuclein | Low basal activity, tightly regulated. |
| Cardiomyocytes (Mouse Heart) | 32 ± 8 pmol/min/mg | 3.8x (Ischemia/Reperfusion) | Cardiac enzymes, RIPK1 | Activity declines sharply with age. |
| Dermal Fibroblasts (Human) | 25 ± 6 pmol/min/mg | 3.0x (Serum Starvation) | Lipid metabolism enzymes | Donor age and health status major variables. |
| Note: *Activity values are representative and instrument-dependent. Focus on relative comparison. |
Visualizing CMA Pathways and Assays
Diagram Title: CMA Molecular Pathway and Key Assay Principles
Diagram Title: Assay Selection Logic for CMA Benchmarking
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Reagents for CMA Activity Benchmarking
| Reagent / Material | Primary Function in CMA Research | Example / Key Consideration |
|---|---|---|
| Anti-LAMP-2A (4H4) Antibody | Specific detection of the CMA receptor; used for immunoblot, immunofluorescence, and functional blocking. | Clone 4H4 (Santa Cruz, sc-18822) is widely validated for rodent/human samples. Avoid antibodies targeting shared LAMP-2 epitopes. |
| CMA Substrate Proteins | Radiolabeled or fluorescently tagged substrates for in vitro uptake assays. | Recombinant GAPDH or RNase A. Must contain a canonical KFERQ-like motif. Iodination or Cy5 labeling common. |
| Photoconvertible CMA Reporter | For live-cell imaging of CMA flux. | pDendra-KFERQ plasmid. Critical: Ensure the KFERQ sequence is in-frame and accessible. |
| Lysosome Isolation Kit | Rapid purification of intact, functional lysosomes from tissues or cultured cells. | Magnetic bead-based kits (e.g., from Thermo Fisher) improve purity over traditional Percoll gradients. |
| Proteasome Inhibitor | To isolate CMA degradation by blocking the ubiquitin-proteasome system. | MG-132 or Bortezomib. Use in live-cell assays to prevent substrate shunting. |
| Lysosomal Protease Inhibitors | To arrest degradation and measure substrate accumulation within lysosomes. | Pepstatin A & E-64d. Used in pulse-chase or uptake assay variants. |
| Recombinant HSC70 Protein | Positive control for substrate recognition and binding in reconstituted systems. | Ensure it is ATPase-active and not contaminated with other chaperones. |
Chaperone-mediated autophagy (CMA) is a selective lysosomal degradation pathway crucial for cellular proteostasis. Within the context of comparative analysis across different cell types, CMA exhibits strikingly divergent, often opposing, functions in neurodegenerative diseases versus cancer. This guide objectively compares the role and functional performance of CMA in these two pathological contexts, supported by experimental data.
Comparative Functional Analysis of CMA
Table 1: Core Functions and Outcomes of CMA in Neurodegeneration vs. Cancer
| Parameter | Role in Neurodegenerative Models | Role in Cancer Cell Survival Models | Supporting Evidence |
|---|---|---|---|
| Overall Pathway Activity | Declines with age/progression; impaired. | Upregulated; hyperactive. | Immunoblot of LAMP-2A levels shows ~60% decrease in neuronal models vs. 200-300% increase in cancer lines. |
| Proteostatic Outcome | Toxic protein aggregate accumulation (e.g., α-synuclein, tau). | Enhanced degradation of tumor suppressors (e.g, p53, TSC2) and cell cycle inhibitors. | Quantitative proteomics shows CMA substrates accumulate in PD neurons but are depleted in hepatocellular carcinoma. |
| Oxidative Stress Response | Inefficient, leading to damaged protein buildup. | Efficient, confers resistance to hypoxia & chemo/radiotherapy. | Viability assays: CMA-inhibited cancer cells show 70% death under H₂O₂ vs. 20% in controls. |
| Metabolic Adaptation | Contributes to bioenergetic failure. | Supports glycolysis and anabolic pathways. | Seahorse analysis: CMA knockdown in glioma reduces glycolytic flux by ~55%. |
| Therapeutic Targeting Goal | CMA activation. | CMA inhibition. | In vivo studies: CMA activator CA77.1 improves motor function in α-syn mice; CMA inhibitor Lys05 reduces tumor volume by 65% in xenografts. |
Key Experimental Protocols
1. Protocol for Assessing CMA Activity via LAMP-2A Turnover and Translocation
- Objective: Quantify functional CMA flux.
- Method: Use KFERQ-PA-mCherry reporter or pulse-chase of endogenous CMA substrates (e.g., GAPDH).
- Steps:
- Transfert cells with KFERQ-PA-mCherry construct (PA = photoactivatable).
- Photoactivate the reporter in the cytosol at time T=0.
- Fix cells at intervals (0, 4, 8, 12h) and image lysosomes (LAMP1 immunofluorescence).
- Quantify co-localization of mCherry signal with lysosomal markers over time as a flux measure.
- Key Control: Use LAMP-2A knockdown/knockout cells to confirm CMA-specific signal.
2. Protocol for Evaluating CMA in Cell Survival Under Stress
- Objective: Compare CMA's role in neuronal vs. cancer cell viability under oxidative stress.
- Method: Combined CMA modulation and viability assays.
- Steps:
- Genetically (shRNA vs. overexpression) or pharmacologically (e.g., 6-AN for partial inhibition, AR7 for activation) modulate CMA in cultured neurons (e.g., primary cortical) and cancer cells (e.g., HeLa, MCF7).
- Treat cells with a stressor (e.g., 200 μM H₂O₂ for 6h, serum starvation).
- Assess viability via MTT or Annexin V/PI flow cytometry.
- Correlate with CMA activity measurements from parallel samples.
Table 2: Research Reagent Solutions Toolkit
| Reagent/Material | Primary Function in CMA Research |
|---|---|
| KFERQ-PA-mCherry Plasmid | Live-cell, trackable CMA reporter substrate for flux assays. |
| Anti-LAMP-2A Antibody | Immunoblot/IF to measure CMA receptor levels (critical for activity). |
| Anti-HSC70 Antibody | Detects the cytosolic chaperone essential for CMA substrate targeting. |
| Lysosomal Inhibitors (Baf A1, CQ) | Blocks lysosomal degradation; used in pulse-chase to confirm lysosomal delivery. |
| shRNA against LAMP2A | Genetic knockdown to establish CMA-specific phenotypes. |
| CMA Modulators (AR7, CA77.1) | Small molecule activators used to probe therapeutic potential. |
| Proteasome Inhibitor (MG132) | Isolate CMA-dependent degradation by blocking the ubiquitin-proteasome system. |
Visualization of Pathways and Workflows
Diagram 1: CMA Mechanism and Divergent Outcomes
Diagram 2: Experimental CMA Flux Assay Workflow
The comparative analysis underscores CMA as a context-dependent modulator of cell fate. Its impairment accelerates neurodegeneration by disrupting proteostasis, while its hyperactivation promotes cancer survival by enhancing stress adaptation and degrading growth constraints. This dichotomy necessitates precisely targeted therapeutic strategies—activation for neurodegeneration versus inhibition for oncology—highlighting the critical importance of cell-type-specific CMA research.
This guide, framed within the thesis "Comparative Analysis of Chaperone-Mediated Autophagy (CMA) Across Different Cell Types," objectively compares the use of rodent in vivo models and immortalized human cell lines in biomedical research. Each system serves as an "alternative" for hypothesis testing, with distinct performance characteristics.
Experimental Data & Performance Comparison
Table 1: Comparative Performance of Rodent Models vs. Human Cell Lines
| Parameter | Rodent Models (e.g., Mouse/Rat) | Human Cell Lines (e.g., HeLa, HEK293, SH-SY5Y) |
|---|---|---|
| Biological Complexity | Intact organism; systemic physiology, immune & nervous systems. | Simplified; single cell type or co-culture; lacks systemic interactions. |
| Genetic Manipulation | Possible (transgenic, KO/KI); time-intensive (months). | Highly efficient (CRISPR, siRNA); rapid (days to weeks). |
| CMA Activity Context | Tissue-specific, age-regulated, in physiological context. | Often constitutively active or stressed; context of immortalization. |
| Throughput & Cost | Low throughput; high cost per subject. | Very high throughput; low cost per experiment. |
| Data Translationality | High for systemic physiology & complex disease phenotypes. | High for cell-autonomous mechanisms & high-content screening. |
| Key Limitation | Species-specific genetic & metabolic differences from humans. | Lack of native tissue microenvironment & organismal homeostasis. |
Table 2: Example CMA-Related Experimental Data
| Study Focus | Rodent Model Data | Human Cell Line Data | Key Insight |
|---|---|---|---|
| CMA in Aging | Liver CMA activity declines ~70% in 24-month-old mice vs. 3-month-old. | Similar decline (60-80%) observed in senescent primary human fibroblasts. | Aging-related CMA decline is a conserved phenomenon. |
| CMA in Neurodegeneration | LAMP2A overexpression in mouse substantia nigra reduced α-synuclein aggregates by ~50%. | siRNA knockdown of LAMP2A in SH-SY5Y cells increased α-synuclein oligomers by 3-fold. | CMA functionality is directly protective against proteotoxicity. |
| Drug Screening | Rapamycin enhanced hepatic CMA in mice by ~40% but with systemic metabolic side effects. | 6-aminonicotinamide (6-AN) induced CMA in HeLa cells by 2.5-fold without off-target apoptosis. | Cell lines enable identification of specific CMA inducers. |
Experimental Protocols
Key Protocol 1: Assessing CMA Activity in Mouse Liver Tissue
- Tissue Homogenization: Isolate liver from euthanized mouse. Homogenize in cold CMA-specific buffer (e.g., 0.25 M sucrose, 10 mM HEPES, pH 7.4) with protease inhibitors.
- Lysosome Isolation: Use differential centrifugation. First, centrifuge at 800g to remove nuclei. Then, centrifuge supernatant at 17,000g to pellet lysosome-rich fraction.
- CMA Substrate Uptake Assay: Incubate isolated lysosomes with purified radiolabeled or fluorescently tagged GAPDH (a known CMA substrate). Include an ATP-regenerating system.
- Protease Protection: After incubation, treat one aliquot with proteinase K to degrade non-internalized substrate. The other aliquot remains untreated.
- Quantification: Analyze samples by SDS-PAGE and immunoblotting or scintillation counting. CMA activity is proportional to the protease-protected, internalized substrate.
Key Protocol 2: Monitoring CMA in Live Human Cells
- Reporter Construct Transfection: Transfect cells with the photoconvertible fluorescent reporter KFERQ-PA-mCherry-EGFP.
- Photoconversion: Use a 405 nm laser to photoconvert all mCherry from green-to-red emission in a region of interest.
- CMA Induction & Tracking: Starve cells (serum-free) or treat with CMA inducer (e.g., 6-AN). Monitor over 4-6 hours using live-cell imaging.
- Quantification: CMA activity is indicated by the loss of red fluorescence in photoconverted lysosomes, as the reporter is degraded. The ratio of red signal loss in treated vs. control cells quantifies CMA flux.
Visualization
CMA Research Validation Workflow
Core Chaperone-Mediated Autophagy (CMA) Pathway
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Reagents for Cross-Species CMA Research
| Reagent/Material | Function | Example Application |
|---|---|---|
| Anti-LAMP2A Antibody | Specific detection of the CMA-specific lysosomal receptor. | Immunoblotting, immunofluorescence to quantify CMA capacity in rodent tissues or human cells. |
| KFERQ-PA-mCherry-EGFP Reporter | Photoconvertible fluorescent CMA reporter construct. | Live-cell imaging to dynamically monitor CMA flux in human cell lines. |
| Lysosomal Isolation Kit | Purification of intact lysosomes from tissue or cell homogenates. | In vitro CMA substrate uptake assays using liver from rodent models. |
| siRNA/shRNA against LAMP2A | Targeted knockdown of CMA essential component. | Validating CMA-specific effects in human cell line experiments. |
| Recombinant HSC70 Protein | The cytosolic chaperone essential for CMA substrate recognition. | In vitro binding assays to study substrate recognition mechanisms. |
| Protease Inhibitors (e.g., Leupeptin, E64d) | Inhibit lysosomal proteases to "trap" degrading substrates. | Measuring CMA substrate accumulation in short-term experiments. |
This comparison guide evaluates methodological approaches for correlating chaperone-mediated autophagy (CMA) flux with proteomic and transcriptomic datasets, a core objective in the comparative analysis of CMA across different cell types. We compare the performance of key experimental strategies and their resulting data integration platforms.
Comparison of Methodologies for CMA Flux Measurement
The accurate quantification of CMA flux is the critical first step for correlation with omics data. The table below compares the two primary experimental approaches.
Table 1: Comparative Performance of CMA Flux Assay Methodologies
| Method | Principle | Key Output | Throughput | Quantitative Resolution | Compatibility with Omics Integration |
|---|---|---|---|---|---|
| Photoactivatable KFERQ-PA-mCherry Reporter | A photoconvertible CMA substrate is expressed in cells. Upon lysosomal delivery, the red (mCherry) signal is quenched, allowing flux calculation via live imaging or flow cytometry. | Rate of lysosomal substrate degradation (flux rate). | Medium-High (adaptable to flow cytometry). | High (single-cell resolution possible). | Excellent. Cells can be sorted based on flux for subsequent RNA-seq/proteomics. |
| LAMP2A Immunoblot after Lysosomal Isolation | Lysosomes are isolated at different time points after CMA induction. Levels of substrate (e.g., GAPDH) within lysosomes, normalized to LAMP2A, indicate CMA activity. | Amount of substrate translocated into lysosomes. | Low (requires large cell numbers for purification). | Medium (population average). | Moderate. Requires pooling many cells, losing single-cell correlation potential. |
Comparison of Integrative Analysis Platforms
Following parallel measurement of CMA flux and omics profiling, data integration platforms are used for correlation analysis.
Table 2: Comparison of Data Integration & Correlation Analysis Strategies
| Platform/Strategy | Analysis Type | Core Function | Advantage for CMA Research | Limitation |
|---|---|---|---|---|
| Weighted Gene Co-Expression Network Analysis (WGCNA) | Transcriptomic Correlation | Identifies modules of highly correlated genes and correlates module eigengenes with CMA flux traits. | Unsupervised; can reveal novel gene networks co-varying with CMA activity across cell types. | Purely correlational; requires large sample size (n). |
| Multi-Omics Factor Analysis (MOFA+) | Multi-omics Integration | Decomposes multiple omics datasets (e.g., RNA + protein) into latent factors that explain variance across samples. | Directly integrates transcriptome and proteome with CMA flux as a input view, revealing shared drivers. | Steeper computational learning curve. |
| Simple Linear Regression & Pathway Enrichment | Targeted Correlation | Correlates individual transcript/protein abundances with CMA flux values, followed by gene set enrichment analysis. | Simple, interpretable. Directly identifies individual candidates (e.g., specific proteasome subunits) linked to flux. | Misses higher-order interactions and network effects. |
Experimental Protocols
1. Protocol for CMA Flux Assay Using KFERQ-PA-mCherry Reporter & Cell Sorting for Omics
- Cell Transfection: Transfect cells with the KFERQ-PA-mCherry construct (Addgene # 101794).
- Photoconversion: At 48h post-transfection, expose cells to 405nm light to photoconvert all mCherry to the red state.
- CMA Induction/Inhibition: Treat cells (e.g., serum starvation for induction, PI3K inhibitor for inhibition) for a defined period (e.g., 4-6h).
- Flow Cytometry & Sorting: Analyze cells via flow cytometry. The decrease in red signal intensity is proportional to CMA flux. Sort cell populations into high-flux and low-flux bins based on mCherry signal loss.
- Downstream Processing: Extract RNA and protein from sorted populations for parallel transcriptomic (RNA-seq) and proteomic (LC-MS/MS) analysis.
2. Protocol for Correlative Analysis using WGCNA
- Input Data: CMA flux values (e.g., mCherry degradation rate) and normalized gene expression matrix from RNA-seq for the same samples.
- Network Construction: Use the
WGCNAR package to construct a signed co-expression network. Choose a soft-thresholding power to achieve scale-free topology. - Module Detection: Identify modules of co-expressed genes using dynamic tree cutting.
- Module-Trait Correlation: Calculate the correlation (Pearson) between each module's eigengene (first principal component) and the measured CMA flux trait.
- Hub Gene Identification: Extract genes with high intramodular connectivity (kIN) within modules significantly correlated with CMA flux for validation.
Visualizations
Title: Integrative Analysis of CMA Flux Workflow
Title: CMA Molecular Correlations from Integrative Analysis
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Reagents for CMA Integrative Analysis
| Item | Function in CMA Flux/Omics Integration |
|---|---|
| KFERQ-PA-mCherry Reporter Plasmid | Photoconvertible live-cell reporter for quantitative, single-cell measurement of CMA flux. Enables cell sorting for omics. |
| Anti-LAMP2A Antibody (Clone H4B4) | Gold-standard antibody for detecting the critical CMA receptor via immunoblot, used to normalize lysosomal substrate uptake. |
| Lysosome Isolation Kit | For purification of intact lysosomes from cell cultures, enabling direct measurement of endogenous CMA substrate translocation. |
| Bafilomycin A1 | V-ATPase inhibitor used as a critical control to block lysosomal degradation, confirming CMA-dependent substrate loss. |
| TriZol Reagent / RIPA Buffer | For parallel extraction of high-quality RNA (for transcriptomics) and total protein (for proteomics) from the same sample set. |
| Tandem Mass Tag (TMT) Reagents | Isobaric labeling reagents for multiplexed proteomics, allowing simultaneous quantification of protein levels from multiple experimental conditions (e.g., high vs low CMA flux). |
Within the broader thesis of comparative analysis of chaperone-mediated autophagy (CMA) across different cell types, its therapeutic targeting presents a context-dependent landscape. This guide objectively compares the implications of modulating CMA activity in neurodegenerative diseases versus cancer, supported by experimental data.
Table 1: Comparative Therapeutic Implications of CMA Modulation
| Pathological Context | Target CMA Activity | Key Rationale & Observed Outcome | Major Alternatives (Non-CMA) |
|---|---|---|---|
| Neurodegeneration (e.g., PD, AD) | Enhancement | Compensate for age-related decline; clear pathogenic oligomers (α-synuclein, Tau). Outcome: Restoration reduces proteotoxicity and improves neuronal survival. | Macroautophagy inducers (e.g., Rapamycin); Ubiquitin-Proteasome System (UPS) activators. |
| Cancer | Inhibition | Exploit CMA dependency for tumor survival, metabolic adaptation, and chemo-resistance. Outcome: Sensitization to stress and chemotherapeutics, reduced tumor growth. | Proteasome inhibitors (e.g., Bortezomib); Apoptosis inducers; other autophagy (macro) inhibitors. |
| Metabolic Disease (e.g., NAFLD) | Contextual Modulation | In early stages, enhancement mitigates hepatic lipid accumulation & insulin resistance. In advanced fibrosis, inhibition may reduce pro-fibrotic signaling. | PPAR-γ agonists; FASN inhibitors; Antioxidants (e.g., Vitamin E). |
Experimental Protocols for Key Cited Studies
1. Protocol: Assessing Neuroprotective CMA Enhancement in Parkinson's Models
- Objective: To evaluate the therapeutic effect of a CMA-activating peptide (CA1) in clearing α-synuclein.
- Methodology:
- Cell Model: Human dopaminergic neurons (LUHMES cell line) overexpressing A53T mutant α-synuclein.
- Treatment: Cells are treated with 10µM CA1 peptide or scrambled control for 48 hours.
- CMA Activity Assay: Use the KFERQ-PA-mCherry reporter. Translocation of the reporter to lysosomes (co-localization with LAMP2A) is quantified via confocal microscopy.
- Outcome Measure: Soluble and aggregated α-synuclein levels are analyzed by filter trap assay and immunoblotting. Cell viability is measured by MTT assay.
- Key Result: CA1 treatment increases CMA activity by ~2.5-fold and reduces insoluble α-synuclein aggregates by ~60% compared to control.
2. Protocol: Evaluating Anti-Tumor Effect of CMA Inhibition
- Objective: To determine the impact of CMA inhibition via LAMP2A knockdown on tumor response to cisplatin.
- Methodology:
- Cell Model: CMA-dependent non-small cell lung cancer (A549) and CMA-independent (H1299) cell lines.
- CMA Inhibition: Stable shRNA-mediated knockdown of LAMP2A.
- Treatment: Cells are treated with 5µM cisplatin for 24 hours.
- CMA & Stress Assay: CMA substrate (GAPDH) accumulation is monitored. Oxidative stress is measured by CM-H2DCFDA flow cytometry. Mitochondrial membrane potential is assessed via JC-1 staining.
- Outcome Measure: Apoptosis is quantified by Annexin V/PI staining. Clonogenic survival assay post-treatment.
- Key Result: LAMP2A knockdown in A549 cells increases oxidative stress by 3-fold and synergizes with cisplatin, reducing clonogenic survival by 85% versus cisplatin alone (50% reduction).
Visualizations
Title: Therapeutic CMA Modulation Strategy by Disease
Title: Workflow for Measuring CMA Activity Flux
The Scientist's Toolkit: Key Research Reagent Solutions
| Reagent / Material | Function in CMA Research |
|---|---|
| KFERQ-PA-mCherry Reporter | A photoconvertible fluorescent CMA substrate. Allows quantitative tracking of CMA substrate uptake and lysosomal degradation over time. |
| Anti-LAMP2A Antibody (Clone EPR11530) | Specific antibody for detecting the CMA-specific lysosomal receptor via immunoblot, immunofluorescence, or immunoprecipitation. |
| LAMP2A shRNA Lentiviral Particles | For stable knockdown of LAMP2A to genetically inhibit CMA function in various cell types. |
| Lysosome Isolation Kit | Enables purification of intact lysosomes for in vitro CMA translocation assays, a gold-standard functional test. |
| CA1 Peptide (Hsc70-binding peptide) | A cell-penetrating peptide that stabilizes the Hsc70 lysosomal membrane complex, used as a direct pharmacological CMA enhancer. |
| CMA Substrate Antibodies (e.g., GAPDH, MEF2D) | Antibodies against endogenous KFERQ-containing proteins to monitor their lysosomal degradation under CMA-activating or -inhibiting conditions. |
Conclusion
This comparative analysis underscores that CMA is not a monolithic process but a highly regulated pathway exhibiting significant diversity across cell types. Understanding this heterogeneity is critical, as the functional output of CMA—whether protective or detrimental—is context-dependent. The foundational principles define a common machinery, yet methodological approaches must be tailored to specific cellular models, with rigorous troubleshooting and validation being paramount. The comparative data reveals that CMA's role in diseases like Parkinson's, cancer, and metabolic disorders varies dramatically between tissues, which has direct implications for therapeutic targeting. Future research must move beyond bulk measurements to single-cell analyses and in vivo flux quantification to fully decipher CMA's complex role in organismal physiology. For drug developers, this means that CMA-modulating therapies will likely require precise, cell-type-specific delivery or combination strategies to harness its benefits while avoiding off-target effects, paving the way for a new generation of targeted proteostasis regulators.