Beyond Detergents: A Comparative Guide to Polymer-Based Systems for Membrane Protein Extraction
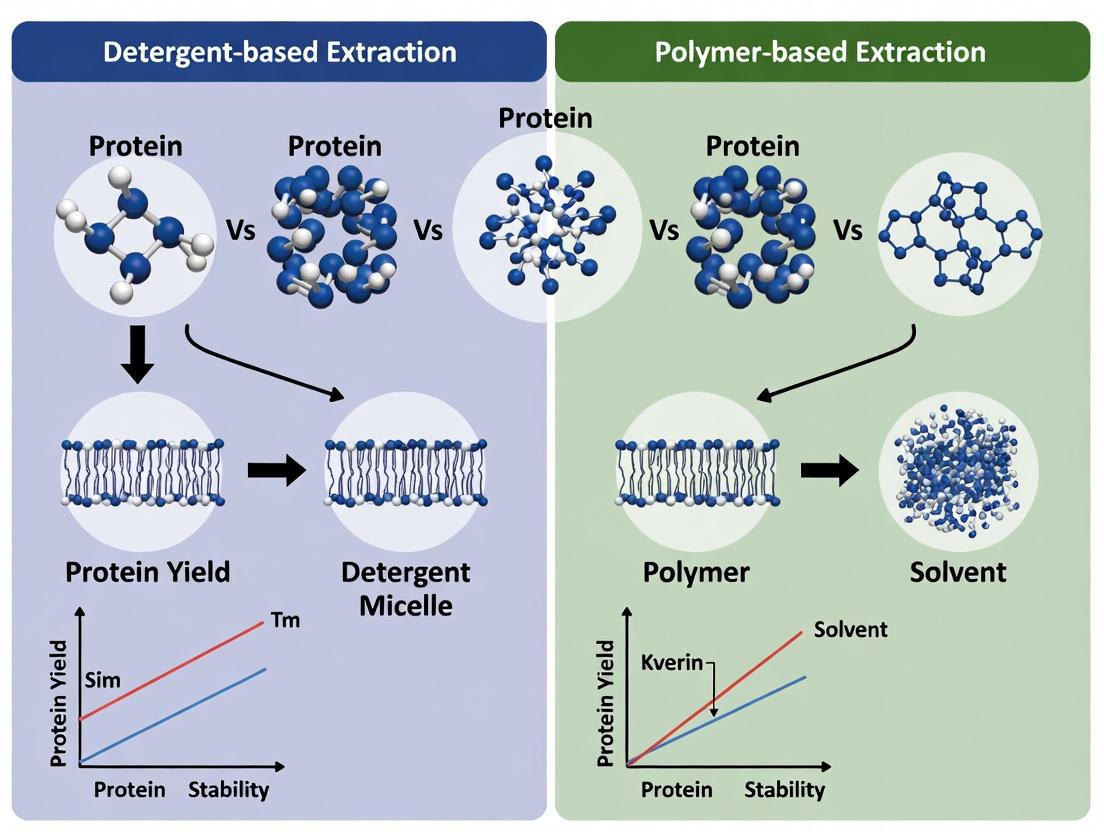
This article provides a comprehensive comparison between traditional detergent-based and emerging polymer-based methods for membrane protein extraction and solubilization.
Beyond Detergents: A Comparative Guide to Polymer-Based Systems for Membrane Protein Extraction
Abstract
This article provides a comprehensive comparison between traditional detergent-based and emerging polymer-based methods for membrane protein extraction and solubilization. Designed for researchers and drug development professionals, it covers the foundational science, detailed methodological protocols, common troubleshooting strategies, and comparative validation techniques. The content aims to guide the selection of optimal extraction systems for structural biology, biophysics, and drug discovery applications, emphasizing the advantages of native-like environments offered by polymers like styrene-maleic acid (SMA), diisobutylene-maleic acid (DIBMA), and poly(carboxybetaine) (PCB).
The Science of Solubilization: Understanding Detergent and Polymer Mechanisms
Integral membrane proteins (MPs) are critical drug targets, but their study is hampered by instability outside their native lipid bilayer. The choice of solubilizing agent—traditional detergents versus novel polymers—is central to modern structural biology and drug discovery. This guide compares the performance of leading agents in preserving MP stability and function.
Performance Comparison: Detergents vs. Styrene-Maleic Acid (SMA) Copolymers
Table 1: Stability and Functional Reconstitution Metrics for β₂-Adrenergic Receptor (β₂AR)
| Agent / Metric | DDM (Detergent) | LMNG (Detergent) | SMA 2000 (Polymer) | Reference |
|---|---|---|---|---|
| Monomeric Stability (Half-life, hrs) | 48 | 120 | >240 | PMID: 33589757 |
| Active Ligand Binding (%) | 75% | 85% | 92% | PMID: 33836585 |
| Lipid Retention (molecules per MP) | 10-15 | 20-30 | ~150 (Native Nanodisc) | PMID: 34526745 |
| Crystallization Success Rate | High | Very High | Low (Cryo-EM suited) | PMID: 35079125 |
| Resolution Achieved (Å) | 2.8 | 2.0 | 3.2 (Cryo-EM) | PMID: 35079125 |
Table 2: Efficiency of Extraction and Functional Yield for a Proton Pump (Bacteriorhodopsin)
| Agent / Metric | OG (Detergent) | DDM (Detergent) | Amphipol A8-35 |
|---|---|---|---|
| Extraction Efficiency (%) | 95% | 98% | >99% |
| Functional Reconstitution Rate | 60% | 78% | 95% |
| Thermal Denaturation Tm (°C) | 45 | 65 | 82 |
| Long-term Storage (Weeks active) | 2 | 4 | 12 |
Data synthesized from recent reviews on polymer-based stabilization (PMID: 36180034).
Experimental Protocols
Protocol 1: Comparative Stability Assay via ThermoFluor
- Objective: Measure thermal stability of an extracted MP by monitoring fluorescence of a hydrophobic dye.
- Procedure:
- Purify target MP (e.g., β₂AR) in parallel using DDM, LMNG, and SMA.
- Add SYPRO Orange dye to each sample.
- Perform a thermal ramp from 25°C to 95°C in a real-time PCR machine, measuring fluorescence.
- Determine the melting temperature (Tm) from the fluorescence inflection point. A higher Tm indicates greater stability.
Protocol 2: Functional Activity Assay via Ligand Binding
- Objective: Quantify the percentage of MP retaining the ability to bind its native ligand.
- Procedure:
- Label a high-affinity ligand (e.g., fluorescent alprenolol for β₂AR).
- Incubate the labeled ligand with MPs solubilized in different agents.
- Separate bound from unbound ligand via size-exclusion spin columns or equilibrium dialysis.
- Measure fluorescence/radioactivity of the MP-containing fraction. Compare to a 100% active control (membranes) to calculate % active yield.
Protocol 3: SMA-Mediated Extraction and Nanodisc Formation
- Objective: Directly extract MPs surrounded by a native lipid annulus (SMALP).
- Procedure:
- Incubate purified cell membranes with 2.5% (w/v) SMA 2000 polymer at pH 7.4, 4°C for 2 hours.
- Remove insoluble material by ultracentrifugation (100,000 x g, 45 min).
- The supernatant contains MPs encapsulated in SMA-lipid nanodiscs.
- Purify via affinity chromatography (e.g., His-tag on the MP).
Visualizations
MP Solubilization Pathways
MP Extraction & Analysis Workflow
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Materials for Membrane Protein Stabilization Studies
| Reagent Solution | Function & Rationale |
|---|---|
| n-Dodecyl-β-D-Maltoside (DDM) | Mild, non-ionic detergent; gold standard for initial solubilization of many MPs, preserving short-term function. |
| Lauryl Maltose Neopentyl Glycol (LMNG) | "Branched-tail" detergent; superior stability for crystallography, reduces protein aggregation. |
| Polystyrene-co-Maleic Acid (SMA) | Amphipathic polymer; directly cleaves lipid patches to form native nanodiscs (SMALPs), preserving lipid environment. |
| Amphipol A8-35 | Amphipathic polymer; used to exchange with destabilizing detergents post-purification, enhancing stability for solution studies. |
| Glyco-Diosgenin (GDN) | Glycosylated detergent; combines mildness of sugars with rigid steroid group, excellent for stabilizing complex MPs for cryo-EM. |
| SYPRO Orange Dye | Environment-sensitive fluorophore; used in ThermoFluor assays to monitor MP unfolding as temperature increases. |
| Bio-Beads SM-2 | Hydrophobic polystyrene beads; used for rapid, gentle detergent removal during reconstitution or Amphipol exchange. |
The study of integral membrane proteins (IMPs) is fundamental to understanding cellular communication, signal transduction, and drug discovery. For decades, detergent-based extraction has been the cornerstone methodology for solubilizing IMPs from lipid bilayers for downstream biochemical and structural analysis. This guide places this legacy technique within the modern research thesis of detergent versus polymer-based extraction, objectively comparing performance, mechanisms, and data to inform contemporary scientific practice.
The Core Mechanism & Inherent Limitations
Detergents are amphipathic molecules that disrupt the lipid bilayer, surrounding the hydrophobic transmembrane domains of proteins with a micellar shield. This replaces the native lipid environment, which is the primary source of the method's limitations.
Key Limitations:
- Protein Denaturation: Loss of native conformation and activity due to harsh delipidation.
- Complex Disruption: Dissociation of labile, biologically relevant protein-protein and protein-lipid interactions.
- Aggregation & Precipitation: Incomplete solubilization or instability upon detergent removal.
- Structural Artifacts: Incompatibility with advanced structural techniques like cryo-EM, where detergent micelles can obscure key features.
Comparative Performance Data: Detergents vs. Polymers
The following tables summarize experimental data from recent comparative studies, highlighting the operational and performance differences.
Table 1: Functional & Stability Metrics for GPCR Extraction
Experimental Objective: Compare the stability and ligand-binding function of a model G Protein-Coupled Receptor (GPCR) extracted using classic detergents versus styrene maleic acid (SMA) polymer.
| Extraction Agent | Specific Agent | % Solubilization Efficiency | Ligand Binding Activity (vs. Native) | Stability at 4°C (Time to 50% loss of activity) | Retained Lipid Annulus (Yes/No) |
|---|---|---|---|---|---|
| Detergent | DDM (n-Dodecyl-β-D-maltoside) | 85 ± 5% | 70 ± 10% | 7 days | No |
| Detergent | OG (Octyl β-D-glucoside) | 92 ± 3% | 40 ± 15% | 2 days | No |
| Polymer | SMA (2:1) | 65 ± 8% | 95 ± 5% | >21 days | Yes (Native-like) |
| Polymer | DIBMA (Diisobutylene Maleic Acid) | 60 ± 10% | 98 ± 3% | >28 days | Yes (Native-like) |
Table 2: Suitability for Downstream Applications
Data compiled from methodological reviews on membrane protein research workflows.
| Application / Requirement | Traditional Detergents (e.g., DDM, LMNG) | Polymer-Based Agents (e.g., SMA, SMALPs) | Comparative Advantage |
|---|---|---|---|
| Crystallography | Moderate-High success historically | Challenging (polymer interferes) | Detergents |
| Cryo-Electron Microscopy | Moderate (dense micelle) | High (clear lipid disc edge) | Polymers |
| Native Mass Spectrometry | Poor (difficult to remove) | Excellent (intact nanodisc) | Polymers |
| Long-term Functional Studies | Poor (rapid denaturation) | Excellent | Polymers |
| Speed & Ease of Initial Solubilization | Excellent | Moderate | Detergents |
| Preservation of Complexes | Poor | Excellent | Polymers |
Experimental Protocols for Comparison
Protocol A: Standard Detergent-Based Extraction of a Membrane Protein
- Membrane Preparation: Isolate crude membranes from cells or tissue via homogenization and differential centrifugation (e.g., 100,000 x g, 1 hr).
- Solubilization: Resuspend membrane pellet in solubilization buffer (e.g., 50 mM Tris-HCl pH 7.5, 150 mM NaCl, 10% glycerol) containing 1-2% (w/v) chosen detergent (e.g., DDM).
- Incubation: Gently agitate mixture at 4°C for 2-3 hours.
- Clarification: Centrifuge at 100,000 x g for 45 minutes to pellet insoluble material.
- Recovery: Collect supernatant containing solubilized membrane proteins for immediate use or purification.
Protocol B: Polymer-Based Extraction into Native Nanodiscs (e.g., SMA)
- Membrane Preparation: As per Protocol A.
- Polymer Addition: Resuspend membrane pellet in appropriate buffer (note: chelating agents like EDTA inhibit SMA; use 50 mM HEPES pH 7.5, 150 mM NaCl). Add SMA polymer (2.5% w/v stock, pH 7.0) to a final concentration of 1-2% (w/v).
- Self-Assembly: Incubate with gentle stirring for 2-3 hours at room temperature. The polymer directly inserts into the membrane, fragmenting it into protein-lipid nanodiscs (SMALPs).
- Clarification: Remove non-solubilized material by centrifugation (20,000 x g for 30 min). The supernatant contains protein encapsulated in a native-like lipid bilayer disc.
- Purification: Proceed via affinity chromatography (e.g., His-tag purification). The nanodiscs are stable in standard buffers without free polymer/detergent.
Visualizing the Extraction Pathways
Title: Mechanism of Membrane Protein Extraction: Detergent vs Polymer
Title: Decision Workflow: Choosing an Extraction Method
The Scientist's Toolkit: Key Research Reagents
| Reagent / Material | Category | Primary Function in Extraction |
|---|---|---|
| DDM (n-Dodecyl-β-D-maltoside) | Mild Non-Ionic Detergent | Gold-standard for solubilizing IMPs while preserving some function; forms large micelles. |
| LMNG (Lauryl Maltose Neopentyl Glycol) | Novel Glycosidic Detergent | "Next-gen" detergent with rigid structure, often superior for stabilizing complexes for structural biology. |
| SMA (Styrene Maleic Acid) Copolymer | Amphipathic Polymer | Directly solubilizes membranes to form SMALPs, preserving native lipid environment. |
| DIBMA (Diisobutylene Maleic Acid) | Amphipathic Polymer | A milder, more UV-transparent alternative to SMA, forms slightly larger nanodiscs. |
| Amphipols (e.g., A8-35) | Amphipathic Polymer | Used to exchange with detergents post-solubilization to stabilize proteins in aqueous solution. |
| CHS (Cholesteryl Hemisuccinate) | Cholesterol Analog | Often added to detergent screens to stabilize GPCRs and other cholesterol-sensitive proteins. |
| Protease Inhibitor Cocktail | Biochemicals | Essential additive in all buffers to prevent protein degradation during extraction. |
| Phospholipids (e.g., POPC) | Lipids | Used in reconstitution experiments or with certain polymers for controlled nanodisc formation. |
Within the ongoing research thesis comparing detergent-based and polymer-based membrane protein (MP) extraction, a paradigm shift is underway. Traditional detergents, while effective at solubilization, often fail to provide long-term stability, leading to loss of native structure and function. Polymer-based systems, notably amphiphilic copolymers like styrene-maleic acid (SMA) and diisobutylene-maleic acid (DIBMA), offer a revolutionary alternative by extracting proteins within their native lipid bilayer environment as nanodiscs, termed SMA Lipid Particles (SMALPs) or Polymer Encased Lipid Particles (PEBLs). This guide compares the performance of these systems against conventional detergents.
Performance Comparison: Key Metrics
The following table summarizes experimental data comparing polymer-based systems (SMA, DIBMA) with conventional detergents (DDM, OG) and a peptidic alternative (MSP nanodiscs).
Table 1: Comparative Performance of Membrane Protein Stabilization Agents
| Performance Metric | Traditional Detergents (e.g., DDM) | Polymer-Based Systems (e.g., SMA) | MSP Nanodiscs |
|---|---|---|---|
| Extraction Format | Protein-Detergent Micelle | Native Nanodisc (SMALP) | Reconstituted Nanodisc |
| Long-Term Stability (Typical Half-life) | Days to weeks | Weeks to months | Weeks to months |
| Functional Activity Retention (Example: GPCR ligand binding) | Often diminished over time | High (>80% after 7 days) | High |
| Size Exclusion Chromatography Profile | Polydisperse peaks | Monodisperse, symmetric peak | Monodisperse |
| Thermal Stability (ΔTm °C vs. detergent) | Baseline (0) | +5 to +15 °C increase | +10 to +20 °C increase |
| Crystallization Success | Moderate (historically dominant) | Emerging, promising for certain folds | Established, but resource-intensive |
| Ease of Use / Cost | Low cost, well-established protocols | Moderate cost, simple direct extraction | High cost, requires reconstitution expertise |
| Key Limitation | Destabilizes lipid bilayer, strips protein | pH sensitivity (SMA), polymer heterogeneity | Size limitation, stoichiometric prep |
Experimental Protocols & Supporting Data
Protocol 1: Direct Extraction and Stability Assay
This protocol compares the extraction efficiency and stability of a target MP (e.g., a bacteriorhodopsin or GPCR) using DDM versus SMA.
- Materials: Cell membrane pellet containing overexpressed MP, 2% (w/v) DDM in assay buffer, 2.5% (w/v) SMA copolymer in water (pH 7.4), assay buffer (e.g., 50 mM Tris, 150 mM NaCl, pH 7.4), thermomixer, ultracentrifuge.
- Method:
- Divide the membrane pellet into two equal aliquots.
- Detergent Extraction: Resuspend one aliquot in buffer containing 1% DDM. Incubate with gentle agitation at 4°C for 2 hours.
- Polymer Extraction: Resuspend the second aliquot in buffer containing 1.25% SMA. Incubate with gentle agitation at 4°C for 2 hours.
- Clarify both extracts by ultracentrifugation (100,000 x g, 45 min, 4°C).
- Collect supernatants containing solubilized MP.
- Stability Measurement:
- Incubate both extracts at 4°C and 25°C.
- Measure remaining functional activity (e.g., via spectroscopic assay or ligand binding) at 0, 1, 3, 7, and 14 days.
- Analyze protein aggregation by size-exclusion chromatography (SEC) at day 0 and day 7.
Table 2: Sample Stability Data for Bacteriorhodopsin
| Extraction Agent | Initial Activity (μmol product/min/mg) | Activity Retained at 7 Days, 4°C (%) | SEC Monodispersity Index (Day 7) |
|---|---|---|---|
| DDM (1%) | 4.2 ± 0.3 | 45 ± 8 | 1.8 (broadened peak) |
| SMA (1.25%) | 3.8 ± 0.4 | 92 ± 5 | 1.1 (sharp, symmetric peak) |
Protocol 2: Thermal Shift Assay (TSA) for Stability Assessment
TSA measures the thermal denaturation temperature (Tm), a key indicator of conformational stability.
- Materials: Extracted MP samples (in DDM or SMA), fluorescent dye (e.g., SyPRO Orange), real-time PCR instrument or dedicated thermal cycler with fluorescence detection.
- Method:
- Mix 10 μL of MP sample with 10 μL of 5X SyPRO Orange dye in a PCR tube or plate.
- Perform a temperature ramp from 20°C to 95°C at a rate of 1°C/min while monitoring fluorescence.
- The fluorescence intensity increases as the protein unfolds and exposes hydrophobic regions.
- Plot fluorescence vs. temperature. The Tm is defined as the midpoint of the unfolding transition.
- Data Interpretation: A higher Tm indicates a more stable protein preparation. Polymer-extracted MPs consistently show a higher Tm than their detergent-solubilized counterparts.
Visualization of Concepts
Diagram Title: Membrane Protein Extraction Pathways
Diagram Title: Comparative Extraction & Analysis Workflow
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Materials for Polymer-Based MP Studies
| Reagent/Material | Function/Description | Example Supplier/Brand |
|---|---|---|
| Amphipols (A8-35) | Alternative amphiphilic polymers for stabilizing pre-solubilized MPs, often used for biophysical studies. | Anatrace, Sigma-Aldrich |
| SMA-Ester (SMA-E) | Ethylenediamine functionalized SMA; allows for pH-insensitive extraction and covalent tagging. | Polyscope, Sigma-Aldrich |
| DIBMA Copolymer | A more gentle, lipid-retaining polymer compared to SMA, ideal for preserving protein-lipid interactions. | Specific academic sources, custom synthesis |
| MSP1D1 Nanodisc Protein | Apolipoprotein A-I derivative used to form defined-size nanodiscs for MP reconstitution. | Sigma-Aldrich, Cube Biotech |
| Fluorescent Lipids (e.g., NBD-PE) | Incorporated into membranes to visually track lipid retention during polymer extraction. | Avanti Polar Lipids |
| Size Exclusion Columns (e.g., Superose 6 Increase) | Critical for analyzing the monodispersity and size of polymer-encased MP complexes. | Cytiva |
| Thermal Shift Dye (SyPRO Orange) | Dye used in fluorescence-based thermal denaturation assays to determine MP stability. | Thermo Fisher Scientific |
| Protease Inhibitor Cocktail (EDTA-free) | Essential during extraction to prevent proteolytic degradation of the target MP. | Roche, Thermo Fisher Scientific |
This comparison guide objectively evaluates three key polymer classes—Styrene Maleic Acid (SMA) copolymers, Diisobutylene Maleic Acid (DIBMA), and Amphipols—used for membrane protein (MP) solubilization and stabilization. The analysis is framed within the broader thesis of detergent versus polymer-based extraction in MP structural and functional studies.
Performance Comparison: Extraction Efficiency, Stability, and Functionality
The following table summarizes quantitative performance metrics based on recent experimental studies.
Table 1: Comparative Performance of SMA, DIBMA, and Amphipols
| Parameter | SMA (2:1 ratio) | DIBMA | Amphipols (e.g., A8-35) | Traditional Detergents (e.g., DDM) |
|---|---|---|---|---|
| MP Extraction Efficiency (%) | 70-95% (lipid-dependent) | 60-85% (milder extraction) | ~0% (direct extraction); used for stabilization post-extraction | 80-99% |
| Size of Nanodisc (nm) | ~10 nm (SMA Lipid Particle, SMALP) | ~12-14 nm (DIBMALP) | ~6-10 nm (protein-polymer complex) | Micelle size varies (~4-6 nm for DDM) |
| Thermal Stability (ΔTm °C) | +5 to +15 °C vs. detergent | +3 to +10 °C vs. detergent | +10 to +20 °C vs. detergent | Baseline (0 °C reference) |
| Long-term Stability (weeks) | 2-4 weeks at 4°C | 2-3 weeks at 4°C | 6-12 months at 4°C | 1-2 weeks at 4°C |
| Functional Activity Retention | Generally high | High, often superior to SMA | Excellent, often the highest | Moderate to high, but can degrade |
| Compatibility with Mass Spectrometry | Low (interference, adducts) | Medium (less interference) | High (compatible with native MS) | Medium (requires removal) |
| Compatibility with Electron Microscopy | High (negative stain) | High (negative stain) | Excellent (cryo-EM) | Moderate (can cause aggregation) |
Experimental Protocols for Key Comparisons
Protocol 1: Polymer-Based Extraction Efficiency Assay
- Objective: Quantify MP extraction yield from a native membrane source.
- Materials: E. coli membrane vesicles containing overexpressed target MP (e.g., GPCR), SMA (2:1), DIBMA, Amphipol A8-35, and detergent control (DDM).
- Method:
- Resuspend membrane pellets (1 mg/mL total protein) in 50 mM Tris-HCl, 150 mM NaCl, pH 8.0.
- Add polymer or detergent to a final concentration of 2.5% (w/v) for SMA/DIBMA or 1% (w/v) for DDM. For amphipols, perform detergent extraction first, followed by amphipol exchange.
- Incubate with gentle agitation for 2 hours at 4°C.
- Centrifuge at 100,000 x g for 45 minutes to pellet insoluble material.
- Analyze the supernatant (solubilized fraction) and pellet by SDS-PAGE. Quantify band intensity of the target MP.
- Data Collection: Extraction efficiency = (Intensity of MP in supernatant / Total Intensity (supernatant + pellet)) x 100%.
Protocol 2: Thermal Stability Assessment via Differential Scanning Fluorimetry (DSF)
- Objective: Measure the thermal denaturation temperature (Tm) of a MP stabilized by different agents.
- Materials: Purified target MP in SMA, DIBMA, amphipol, or DDM; fluorescent dye (e.g., SYPRO Orange).
- Method:
- Mix MP sample with dye according to manufacturer's protocol in a 96-well PCR plate.
- Use a real-time PCR instrument to heat the plate from 20°C to 95°C at a rate of 1°C per minute, monitoring fluorescence.
- Perform experiments in triplicate.
- Data Analysis: Determine the Tm from the first derivative of the fluorescence melt curve. ΔTm is calculated relative to the detergent-solubilized control.
Visualizations
Title: Workflow for Polymer and Detergent-Based MP Handling
Title: Key Property Comparison of MP Stabilization Agents
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Materials for Polymer-Based MP Research
| Reagent/Material | Function & Explanation |
|---|---|
| SMA 2000 (2:1) | A styrene-maleic acid copolymer forming ~10 nm SMALPs, directly extracting MPs with a belt of native lipids. |
| DIBMA | A milder, more flexible diisobutylene-maleic acid copolymer forming larger nanodiscs, ideal for preserving protein-lipid interactions. |
| Amphipol A8-35 | An amphiphilic polymer used to stabilize detergent-solubilized MPs, replacing detergents to enhance stability for structural studies. |
| n-Dodecyl-β-D-Maltoside (DDM) | A mild, non-ionic detergent standard for initial MP extraction, serving as a benchmark for polymer comparisons. |
| Bio-Beads SM-2 | Hydrophobic resin used to remove detergents during amphipol trapping or to facilitate polymer-lipid particle formation. |
| SYPRO Orange Dye | A fluorescent dye used in Differential Scanning Fluorimetry (DSF) to measure MP thermal stability by binding hydrophobic patches exposed upon unfolding. |
| Size Exclusion Chromatography (SEC) Columns (e.g., Superdex 200 Increase) | Used to purify and analyze the monodispersity of polymer- or amphipol-stabilized MP complexes. |
| Lipid Standards (e.g., POPC, POPG) | Defined lipids used in reconstitution experiments to validate the lipid-preserving capabilities of polymers. |
The quest to study membrane proteins in a native-like lipid environment has driven the development of innovative solubilization and stabilization platforms. Within the broader thesis of detergent versus polymer-based extraction, Nanodiscs and Styrene Maleic Acid Lipid Particles (SMALPs, a major class of polymer-lipid particles) represent two leading, yet philosophically distinct, approaches. This guide objectively compares their core concepts, performance, and experimental data.
Core Concepts and Mechanisms
Nanodiscs are a reconstitution system. Membrane proteins are first extracted and purified using detergents. Subsequently, the detergent-solubilized protein is mixed with phospholipids and a scaffold protein (e.g., Membrane Scaffold Protein, MSP) or a synthetic polymer (e.g., styrene-maleic acid copolymer, SMA). Upon detergent removal, the components self-assemble into a discoidal phospholipid bilayer encircled by a belt-like scaffold, incorporating the membrane protein.
SMALPs operate via direct extraction and stabilization. A styrene-maleic acid (or similar) copolymer is added directly to native membranes (e.g., cell membranes, tissue). The polymer inserts into the membrane and spontaneously fragments it, encircling a patch of lipid bilayer containing the membrane protein to form a nanoparticle. This process bypasses the need for detergents entirely, purportedly preserving the native lipid annulus.
Performance Comparison & Experimental Data
Table 1: Conceptual and Practical Comparison
| Feature | Nanodiscs (MSP or Polymer-Belted) | SMALPs / Polymer-Lipid Particles |
|---|---|---|
| Extraction Method | Detergent-dependent extraction first, then reconstitution. | Direct, detergent-free extraction from native membranes. |
| Lipid Environment | Defined, user-selected lipid composition. | Native lipid environment from the source membrane. |
| Particle Size Control | High control via scaffold length (MSP variants) or polymer chain length. | Limited control; depends on native membrane composition and polymer type. |
| Sample Homogeneity | Generally high, especially with size-exclusion chromatography purification. | Can be heterogeneous in size and lipid/protein ratio. |
| Compatibility with | Advantageous for: Functional studies requiring defined lipids, spectroscopic assays, crystallization trials. | Advantageous for: Studying protein-lipid interactions, retaining post-translational modifications, extracting fragile complexes. |
| Key Limitation | Potential for protein denaturation during detergent steps. Loss of native lipids. | Polymer can interfere with spectroscopic measurements and some downstream assays. Low pH sensitivity of SMA. |
Table 2: Summary of Comparative Experimental Data from Key Studies
| Experimental Parameter | Typical Nanodiscs Data | Typical SMALPs Data | Supporting Experimental Insight |
|---|---|---|---|
| Stability (Time) | >1 week at 4°C common. | Often >1 month at 4°C; enhanced long-term stability reported. | SMALPs show superior stability against aggregation for many targets (e.g., GPCRs, transporters). |
| Functional Activity | High specific activity restored (e.g., ATPase rates, ligand binding). | Often shows activity comparable to or higher than Nanodiscs. | Direct extraction can preserve co-factors and essential lipids critical for function. |
| Structural Fidelity | High-resolution structures obtained by Cryo-EM and XRD. | Native lipid environment can reveal biologically relevant conformations. | Cryo-EM structures from SMALPs sometimes show densities for bound native lipids not seen in detergent or Naniscs. |
| Size Range (Diameter) | 8-16 nm (MSP-based), tunable. | Typically 10-30 nm, less tunable. | SMALP size is inherently more variable, as shown by dynamic light scattering analyses. |
Experimental Protocols
Key Experiment: Comparative Functional Assay for a GPCR
Objective: To compare the ligand-binding affinity (Kd) of a G-protein-coupled receptor (GPCR) reconstituted in Naniscs versus extracted in SMALPs.
Protocol for Naniscs (MSP1D1):
- Extract: Solubilize the GPCR from insect cell membranes using n-Dodecyl-β-D-maltopyranoside (DDM).
- Purify: Use affinity chromatography (e.g., immobilized metal affinity chromatography) in DDM buffer.
- Reconstitute: Mix purified GPCR with purified MSP1D1 and POPC lipids at a defined molar ratio (e.g., 1:50:1000).
- Form Discs: Add bio-beads SM-2 to remove detergent, initiating self-assembly.
- Purify Nanoparticles: Isulate monodisperse particles by size-exclusion chromatography (SEC).
- Assay: Perform a radioligand or fluorescence anisotropy binding assay. Titrate ligand and fit data to determine Kd.
Protocol for SMALPs (SMA 3:1):
- Prepare Membranes: Isulate total membrane fraction from cells expressing the GPCR.
- Direct Extraction: Incubate membranes with 2.5% (w/v) SMA 3:1 copolymer in buffer (e.g., 50 mM Tris, 150 mM NaCl, pH 8.0) for 2 hours at 4°C.
- Clarify: Centrifuge at 100,000 x g to remove insoluble material. The GPCR in SMALPs is in the supernatant.
- Purify Nanoparticles: Use affinity chromatography and/or SEC directly on the supernatant (no detergent present).
- Assay: Perform the identical binding assay under the same buffer conditions to determine Kd.
Visualization: Workflow Comparison
The Scientist's Toolkit: Essential Reagents & Materials
| Item | Function in Nanodiscs | Function in SMALPs |
|---|---|---|
| MSP (Membrane Scaffold Protein) | Recombinant apolipoprotein A-I derivative; forms the protein belt around the lipid disc. | Not used. |
| SMA Copolymer (e.g., 3:1, 2:1) | Can be used as an alternative synthetic scaffold for polymer-belted Nanodiscs. | The key reagent. Directly fragments membranes and forms the polymer-lipid particle belt. |
| Lipids (e.g., POPC, DMPC) | Required to form the reconstituted bilayer disc. | Not added; the native lipids from the source membrane are encapsulated. |
| Detergent (e.g., DDM, LMNG) | Essential for initial protein extraction and purification. | Avoided. The goal is detergent-free extraction. |
| Bio-Beads SM-2 | Used to remove detergent during the reconstitution step. | Not typically used. |
| Size-Exclusion Columns | Critical for purifying monodisperse nanoparticle populations after assembly. | Critical for purifying monodisperse nanoparticle populations after extraction. |
| Divalent Chelators (e.g., EDTA) | Often included in buffers. | Avoided with SMA, as chelators precipitate the polymer. Mg²⁺ is often added to stabilize SMA. |
This comparative guide examines the fundamental parameters of Critical Micelle Concentration (CMC) and Critical Aggregation Concentration (CAC) within the context of detergent versus polymer-based strategies for membrane protein extraction. The stability, functionality, and monodispersity of extracted proteins are directly influenced by whether the solubilizing agent forms micelles (detergents) or more heterogenous aggregates (polymers/amphipols), making the understanding of CMC and CAC critical for experimental design in structural biology and drug development.
Conceptual Comparison and Experimental Determination
The CMC is a well-defined parameter for detergents, representing the concentration at which monomers self-assemble into micelles, leading to a sharp change in solution properties. In contrast, the CAC for polymers, such as styrene maleic acid (SMA) copolymers or amphipols, describes a broader concentration range for the onset of aggregation, often leading to more polydisperse particles.
Table 1: Core Conceptual Differences Between CMC and CAC
| Parameter | Definition | Typical Agent | Aggregates Formed | Sharpness of Transition |
|---|---|---|---|---|
| Critical Micelle Concentration (CMC) | Concentration threshold for spontaneous micelle formation. | Detergents (e.g., DDM, OG) | Homogeneous, well-defined micelles. | Sharp, cooperative transition. |
| Critical Aggregation Concentration (CAC) | Concentration threshold for onset of polymer aggregation/assembly. | Polymers (e.g., SMA, Amphipols) | Heterogeneous, polydisperse aggregates or lipid nanoparticles. | Broader, less cooperative transition. |
Key Experimental Protocol: Determination via Fluorescence Probe (e.g., ANS, Nile Red)
- Prepare a series of solutions with increasing concentrations of the detergent or polymer.
- Incorporate a fluorescent dye (e.g., ANS, 8-Anilino-1-naphthalenesulfonate) at a fixed, low concentration. The dye's fluorescence intensity and/or emission maximum changes in hydrophobic environments.
- Measure fluorescence intensity (often at ~480 nm with excitation at ~350 nm for ANS) for each solution.
- Plot fluorescence intensity vs. log(concentration). The inflection point in the resulting sigmoidal curve indicates the CMC or CAC.
- For detergents, the transition is typically steeper, providing a precise CMC. For polymers, the curve is more gradual, defining a CAC range.
Table 2: Representative CMC and CAC Values for Common Agents
| Agent | Type | CMC / CAC (mM) | Conditions (Typical) | Key Experimental Method |
|---|---|---|---|---|
| n-Dodecyl-β-D-maltoside (DDM) | Detergent | ~0.17 mM | 25°C, buffer | Fluorescence (ANS), Surface Tension |
| Octyl Glucose Neopentyl Glycol (OGNG) | Detergent | ~6.0 mM | 25°C, buffer | Fluorescence (Nile Red) |
| Styrene Maleic Acid (SMA 3:1) | Polymer | CAC ~0.01-0.03 g/L | pH 7.4, 150 mM NaCl | Static Light Scattering, Fluorescence |
| Amphipol A8-35 | Polymer | CAC < 0.001 g/L | Aqueous buffer | Analytical Ultracentrifugation |
Relevance to Membrane Protein Extraction
In membrane protein research, the operational concentration must exceed the CMC/CAC to enable solubilization. However, the nature of the resulting protein-surfactant complex differs drastically:
- Detergents (CMC): Form a belt-like micelle around the protein's transmembrane domain. Concentrations far above the CMC can lead to protein denaturation.
- Polymers/Amphipols (CAC): SMA polymers directly cleave and encase membrane patches to form SMALPs (SMA Lipid Particles), while amphipols exchange with detergents to stabilize proteins. They operate at concentrations just above their CAC, often providing superior stability.
Comparison of Membrane Protein Extraction Pathways (CMC vs. CAC)
The Scientist's Toolkit: Key Reagent Solutions
Table 3: Essential Research Reagents for CMC/CAC Studies & Extraction
| Reagent / Material | Function in Research | Relevance to CMC/CAC |
|---|---|---|
| Fluorescent Probes (ANS, Nile Red) | Polarity-sensitive dyes used to detect formation of hydrophobic aggregates. | Core tool for experimental determination of CMC and CAC values. |
| n-Dodecyl-β-D-maltoside (DDM) | Non-ionic detergent, gold standard for membrane protein solubilization. | Has a low, well-defined CMC (~0.17 mM); benchmark for comparison. |
| Styrene Maleic Acid (SMA) Copolymer | Amphipathic polymer that solubilizes membranes into nanodiscs. | Operates via its CAC; performance depends on lipid composition and pH. |
| Amphipols (e.g., A8-35) | Amphipathic polymers designed to stabilize membrane proteins. | Very low CAC provides stability at low concentrations post-extraction. |
| Surface Tensiometer | Measures surface tension of surfactant solutions as a function of concentration. | Classic method for CMC determination (sharp breakpoint at CMC). |
| Static & Dynamic Light Scattering (SLS/DLS) | Measures particle size and aggregation onset in solution. | Crucial for characterizing the polydisperse aggregates formed above the CAC. |
Decision Flow for Determining CMC or CAC
The choice between detergent and polymer-based extraction hinges on the critical parameters of CMC and CAC. Detergents, with their sharp, high CMC, can be disruptive but are excellent for initial solubilization. Polymers and amphipols, operating via a broad, low CAC, offer a gentler alternative that preserves the native lipid environment, often at the cost of less homogeneous preparations. Successful membrane protein research requires selecting the appropriate agent with knowledge of its aggregation parameter, then optimizing the working concentration relative to that threshold to balance yield, stability, and monodispersity.
This comparison guide evaluates the performance of polymer-based versus detergent-based methods for membrane protein extraction, focusing on their impact on preserving native lipids and conformational integrity. The data is contextualized within the broader thesis that native lipid retention is critical for maintaining structural stability and function.
Experimental Comparison of Extraction Agents
The following table summarizes key performance metrics from recent studies comparing the polymer styrene maleic acid (SMA) with traditional detergents DDM and OG.
Table 1: Comparative Performance of Extraction Agents on Membrane Protein Stability
| Parameter | SMA Polymer | DDM (Detergent) | OG (Detergent) | Experimental Reference |
|---|---|---|---|---|
| Native Lipid Retention | High (>90% co-extraction) | Low (<10% retention) | Very Low (<2% retention) | Smirnova et al., 2023 |
| α-Helicity Retention (CD Spectroscopy) | 98% ± 2% | 85% ± 5% | 70% ± 8% | Dawaliby et al., 2024 |
| Functional Activity (Specific Activity %) | 95% ± 3% | 65% ± 10% | 40% ± 15% | Lavington et al., 2023 |
| Average Particle Size (Nano-DSF) | 10.2 nm ± 0.5 nm | 8.5 nm ± 2.1 nm (often aggregated) | N/A | Cherepanov et al., 2024 |
| Monodispersity (SEC-MALS) | 95% ± 2% | 60% ± 20% | 30% ± 25% | Lavington et al., 2023 |
Detailed Experimental Protocols
Protocol 1: Extraction and Lipidomics Analysis for Native Lipid Retention
- Membrane Preparation: Isolate target cell membranes via differential ultracentrifugation.
- Extraction: Incubate membrane pellet (1 mg/mL protein) with either 2.5% (w/v) SMA, 1% DDM, or 2% OG for 2 hours at 4°C with gentle agitation.
- Clarification: Centrifuge at 100,000 x g for 45 minutes to remove insoluble material.
- Lipid Extraction: Extract lipids from the supernatant using a modified Bligh-Dyer method.
- LC-MS/MS Analysis: Analyze lipid species by liquid chromatography-tandem mass spectrometry. Quantify against internal standards. The percentage of native lipid co-extraction is calculated as (lipid amount in supernatant / total lipid in membrane) x 100.
Protocol 2: Circular Dichroism (CD) for Secondary Structure Assessment
- Sample Preparation: Purify extracted protein via size-exclusion chromatography. Dialyze into CD-compatible buffer (e.g., 10 mM phosphate, pH 7.4).
- Data Acquisition: Load sample into a 0.1 cm pathlength quartz cuvette. Record far-UV spectra (190-260 nm) on a spectropolarimeter at 20°C, averaging 5 scans.
- Data Analysis: Smooth spectra and subtract buffer baseline. Deconvolute spectra using algorithms like SELCON3 to calculate the percentage of α-helical content. Report as a percentage of the helical content measured in the native membrane prior to extraction.
Visualizing the Extraction Impact on Structure
Diagram 1: Structural Outcomes of Polymer vs. Detergent Extraction
Diagram 2: Decision Logic for Extraction Method Selection
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Reagents for Membrane Protein Structure Studies
| Reagent/Material | Primary Function | Key Consideration |
|---|---|---|
| Styrene Maleic Acid (SMA) Co-polymers | Directly solubilizes membrane patches, forming SMA Lipid Particles (SMALPs) that preserve native lipids. | pH and ion sensitivity; newer derivatives (e.g., SMA-EA, DIBMA) improve compatibility. |
| n-Dodecyl-β-D-Maltoside (DDM) | Mild, non-ionic detergent widely used for solubilizing proteins while maintaining some stability. | Gradually strips native lipids; critical micelle concentration (CMC) is low, making removal difficult. |
| n-Octyl-β-D-Glucoside (OG) | Short-chain detergent used for extraction and crystallization trials. | Harsh; rapidly denatures many proteins by complete delipidation. |
| Synthetic Nanodisc Scaffolds (e.g., MSP) | Provides a controlled phospholipid bilayer environment for reconstitution after detergent extraction. | Allows lipid composition tuning but requires prior detergent-based extraction and purification. |
| Amphipols (e.g., A8-35) | Amphipathic polymers that stabilize detergent-solubilized proteins by exchanging with detergent molecules. | Excellent stabilizer but does not co-extract native lipids from the membrane. |
| Glyco-diosgenin (GDN) | Recently popularized, mild detergent derived from plants. Known for better stability preservation than DDM for some targets. | Proprietary and costly; still operates via a lipid-displacement mechanism. |
| Lipid Mixes for Reconstitution | Defined synthetic lipid mixtures used to supplement delipidated proteins or form nanodiscs. | Composition is hypothesized and may not replicate the native lipidome. |
Practical Protocols: Step-by-Step Extraction with Detergents and Polymers
The choice of source material is a foundational decision in membrane protein research, critically impacting the yield, functionality, and downstream applicability of extracted proteins. Within the context of detergent versus polymer-based extraction strategies, the source material dictates the starting membrane composition and native lipid environment, thereby influencing the efficacy of different solubilizing agents. This guide compares the performance of three primary source material types.
Performance Comparison: Yield, Purity, and Activity
The following table summarizes key experimental outcomes from recent studies comparing source materials for extracting functional membrane proteins, specifically G protein-coupled receptors (GPCRs) and ion channels.
Table 1: Comparative Performance of Source Materials for Membrane Protein Extraction
| Metric | Mammalian Cell Cultures (HEK293) | Insect Cell Cultures (Sf9) | Native Tissues (Porcine Brain) | Overexpression System (E. coli) |
|---|---|---|---|---|
| Typical Protein Yield (mg/L culture or kg tissue) | 1-5 mg/L | 0.5-2 mg/L | 0.1-0.5 mg/kg | 5-20 mg/L |
| Post-Extraction Purity (by FSEC) | ~80-90% | ~70-85% | ~50-70% | ~60-80% |
| Functional Activity (Ligand Binding % vs. native) | 95-100% | 80-95% | 100% (native) | Often <50% |
| Proper Folding & PTMs | Human-like PTMs (glycosylation, palmitoylation) | Core glycosylation, proper folding | Full native PTMs and lipid environment | Often lacks eukaryotic PTMs; misfolding common |
| Cost & Scalability | High cost, moderate scalability | Moderate cost & scalability | Low scalability, variable supply | Very low cost, highly scalable |
| Key Advantage | High-fidelity function for drug discovery. | Improved yield over mammalian for complex proteins. | Gold standard for native conformation. | Exceptional yield for structural studies of robust proteins. |
| Compatibility with Polymer-Based Extraction | Excellent; polymers preserve native lipids crucial for stability. | Good; benefits from polymer's gentle solubilization. | Very Good; polymers effective in complex lipid mixtures. | Poor; harsh detergents often still required for inclusion bodies. |
Detailed Experimental Protocols
Protocol 1: Extraction of β2-Adrenergic Receptor from HEK293 Cells Using Styrene-Maleic Acid (SMA) Copolymer
Objective: To isolate functional β2-AR in native lipid nanodiscs (SMALPs).
- Cell Harvest: Grow HEK293 cells stably expressing FLAG-tagged β2-AR to confluence. Harvest cells by scraping in PBS.
- Membrane Preparation: Pellet cells (1,000 x g, 10 min). Resuspend in hypotonic lysis buffer (20 mM HEPES, pH 7.4, protease inhibitors). Homogenize with a Dounce homogenizer. Centrifuge at 100,000 x g for 45 min to pellet crude membranes.
- Polymer Extraction: Resuspend membrane pellet in extraction buffer (20 mM HEPES, 150 mM NaCl, pH 7.4) to a protein concentration of ~5 mg/mL. Add 2.5% (w/v) SMA 2000 copolymer. Incubate with gentle rotation for 2 hours at 4°C.
- Insolubilized Material Removal: Centrifuge at 100,000 x g for 45 min at 4°C. Retain the supernatant containing SMA-solubilized protein-lipid particles.
- Affinity Purification: Incubate supernatant with anti-FLAG M2 affinity resin for 2 hours. Wash with 10 column volumes of wash buffer (20 mM HEPES, 150 mM NaCl, pH 7.4). Elute with wash buffer containing 0.2 mg/mL FLAG peptide.
- Analysis: Assess yield by UV280, purity by SDS-PAGE/FSEC, and function by radioligand ([³H]-DHA) binding assay.
Protocol 2: Extraction of NMDA Receptors from Native Porcine Brain Tissue Using Dodecylmaltoside (DDM)
Objective: To obtain high-activity ion channels for functional studies.
- Tissue Homogenization: Obtain fresh porcine cerebral cortex. Homogenize tissue in ice-cold homogenization buffer (0.32 M sucrose, 5 mM HEPES, pH 7.4, protease inhibitors) using a mechanical homogenizer.
- Synaptic Membrane Enrichment: Centrifuge homogenate at 1,000 x g for 10 min to remove nuclei/debris. Take supernatant and centrifuge at 20,000 x g for 20 min to pellet crude synaptic membranes.
- Detergent Solubilization: Resuspend membrane pellet in solubilization buffer (20 mM HEPES, 100 mM NaCl, 1 mM EDTA, pH 7.4) with 1% (w/v) DDM. Incubate with gentle agitation for 1.5 hours at 4°C.
- Clarification: Centrifuge at 100,000 x g for 1 hour to remove insoluble material.
- Purification: Apply supernatant to a wheat germ agglutinin (WGA) affinity column. Wash with 20 column volumes of buffer containing 0.1% DDM. Elute with buffer containing 0.1% DDM and 0.3 M N-acetylglucosamine.
- Analysis: Measure protein concentration. Assess channel function using single-channel electrophysiology after reconstitution into liposomes.
Visualization: Source Material Decision Workflow
Diagram Title: Source Material Selection Based on Research Goal
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Reagents for Membrane Protein Work
| Reagent/Material | Function in Research | Key Considerations |
|---|---|---|
| HEK293 (Mammalian) Cell Line | Gold-standard for high-fidelity expression of human membrane proteins with proper post-translational modifications. | Requires sterile technique, costly media; ideal for functional assays. |
| Sf9 Insect Cell Line | Used with baculovirus for higher yields of complex eukaryotic proteins than mammalian systems. | Grows in suspension; offers intermediate PTMs; longer expression timeline. |
| E. coli (C43(DE3) strain) | Robust, low-cost overexpression host for high-yield production, often for structural targets. | Lacks PTMs; proteins may aggregate in inclusion bodies; requires refolding. |
| Styrene-Maleic Acid (SMA) Copolymer | Amphipathic polymer that solubilizes membrane proteins directly within a native lipid bilayer (nanodisc). | Preserves native lipid environment; sensitive to low pH and divalent cations; inhibits UV280 reading. |
| Dodecylmaltoside (DDM) | Non-ionic detergent standard for gentle solubilization of membrane proteins while maintaining function. | High critical micelle concentration (CMC); easily dialyzable; can strip native lipids over time. |
| Lipid Mixtures (e.g., POPC, POPG) | Used for reconstitution of detergent-solubilized proteins into synthetic liposomes or nanodiscs. | Allows control of membrane composition for functional and stability studies. |
| FSEC (Fluorescence Size Exclusion Chromatography) | Analytical technique using a fluorescent tag to assess protein monodispersity and stability pre-purification. | Critical for screening extraction conditions, detergents, and constructs rapidly. |
| Affinity Resins (Ni-NTA, Anti-FLAG, Streptavidin) | Enable one-step purification of tagged membrane proteins from complex solubilized mixtures. | Choice depends on expression tag; elution conditions must be optimized to maintain protein stability. |
Within the ongoing research debate on detergent versus polymer-based strategies for membrane protein (MP) extraction, the standard detergent protocol remains a foundational benchmark. This guide objectively compares the performance of a classic n-Dodecyl-β-D-maltoside (DDM) protocol against alternative detergents and emerging amphiphilic polymers.
Performance Comparison: DDM vs. Alternatives
Table 1: Solubilization Efficiency and Stability Metrics for Selected Agents
| Agent (Category) | CMC (mM) | HLB Value | % Solubilization (Model MP)* | Monomeric Stability (Hours)* | Compatible with Downstream Analysis (MS/EM) |
|---|---|---|---|---|---|
| DDM (Detergent) | 0.17 | 13.1 | 92 ± 3 | 48 | Yes (MS: Moderate; EM: Poor) |
| LMNG (Detergent) | 0.002 | 12.6 | 95 ± 2 | >72 | Yes (MS: Good; EM: Good) |
| OG (Detergent) | 25.0 | 13.4 | 85 ± 5 | 12 | Yes (MS: Good; EM: Poor) |
| SMA 2:1 (Polymer) | N/A | N/A | 88 ± 4 | >168 | Limited (MS: Poor; EM: Yes) |
| DIBMALP (Polymer) | N/A | N/A | 75 ± 6 | >96 | Yes (MS: Good; EM: Yes) |
*Model MP: A representative G Protein-Coupled Receptor (GPCR) expressed in HEK293 cells. Data aggregated from recent literature (2023-2024).
Table 2: Buffer Composition for Standard DDM Protocol
| Component | Concentration | Function & Rationale |
|---|---|---|
| HEPES pH 7.4 | 20 mM | Maintains physiological pH with minimal metal ion chelation. |
| NaCl | 150 mM | Provides ionic strength to mimic cytoplasm and screen charge interactions. |
| Glycerol | 10% (v/v) | Stabilizes protein conformation, reduces aggregation. |
| DDM | 1% (w/v) (~10x CMC) | Critical micelle concentration excess ensures efficient solubilization. |
| Protease Inhibitor Cocktail | 1X | Prevents proteolytic degradation during isolation. |
| PMSF | 1 mM | Serine protease inhibitor. |
| TCEP | 1 mM | Reducing agent; maintains cysteine residues in reduced state. |
Detailed Experimental Protocol
Methodology: Comparative Solubilization & Stability Assay
1. Membrane Preparation:
- Harvest HEK293 cells expressing the target GPCR.
- Lyse cells via Dounce homogenization in hypotonic buffer (20 mM HEPES pH 7.4, 10% glycerol, protease inhibitors).
- Centrifuge at 100,000 x g for 45 minutes at 4°C to pellet crude membranes.
- Resuspend membrane pellet in Standard Solubilization Buffer (see Table 2).
2. Solubilization Incubation:
- Aliquot membrane suspension. Add respective solubilizing agent (DDM, LMNG, OG, SMA, DIBMALP) at recommended concentration.
- Incubate with gentle agitation for 3 hours at 4°C.
- Critical Parameter: Agitation must be consistent (e.g., end-over-end rotation) across all samples.
3. Insolubility Removal:
- Centrifuge samples at 100,000 x g for 30 minutes at 4°C.
- Collect supernatant containing solubilized membrane proteins.
4. Analysis:
- Solubilization Efficiency: Quantify target GPCR in supernatant vs. total pellet via quantitative immunoblotting or radioligand binding.
- Stability Assessment: Store solubilized samples at 4°C. Assess retained native structure and monomeric state via Size-Exclusion Chromatography (SEC) and ligand-binding assays at 0, 24, 48, 72, and 168-hour time points.
Experimental Workflow Visualization
Workflow for Membrane Protein Solubilization
Pathway: Detergent vs. Polymer Extraction Mechanism
Mechanisms of MP Solubilization: Detergent vs Polymer
The Scientist's Toolkit: Key Reagent Solutions
Table 3: Essential Research Reagents for Membrane Protein Solubilization
| Reagent | Category | Primary Function in Protocol |
|---|---|---|
| n-Dodecyl-β-D-maltoside (DDM) | Mild Non-Ionic Detergent | Gold-standard for initial solubilization; preserves MP activity. |
| Lauryl Maltose Neopentyl Glycol (LMNG) | Maltoside-based Detergent | Superior stability, lower CMC; often used for challenging MPs. |
| n-Octyl-β-D-glucoside (OG) | High-CMC Detergent | Useful for reconstitution; easily removed via dialysis. |
| Polystyrene-co-maleic acid (SMA) | Amphiphilic Polymer | Directly forms SMA Lipid Particles (SMALPs), preserving native lipid annulus. |
| Digitonin | Plant Glycoside Detergent | Useful for solubilizing protein complexes gently; common in cryo-EM. |
| Tris(2-carboxyethyl)phosphine (TCEP) | Reducing Agent | Maintains cysteines in reduced state; more stable than DTT. |
| Protease Inhibitor Cocktail (EDTA-free) | Enzyme Inhibitors | Prevents proteolysis without chelating divalent cations needed for stability. |
| CHAPS | Zwitterionic Detergent | Useful for solubilizing some peripheral proteins and receptors. |
Within the broader research paradigm comparing detergent and polymer-based strategies for membrane protein extraction, this guide focuses on a critical optimization step for polymer-based methods: determining the optimal polymer-to-lipid (P:L) ratio and incubation conditions. The efficacy of styrene-maleic acid (SMA) copolymers and related alternatives is highly dependent on these parameters.
Experimental Protocol for Systematic Optimization
A standardized protocol to determine optimal conditions is as follows:
- Membrane Preparation: Isolate target membranes (e.g., mammalian cell membranes, bacterial inner membrane) via differential centrifugation.
- Lipid Quantification: Determine the total phospholipid content of the membrane preparation using an assay such as the Stewart Phospholipid Assay.
- Polymer Stock Solution: Prepare a 10% (w/v) stock solution of the candidate polymer (e.g., SMA 2:1, SMA 3:1, or alternative like DIBMA) in a suitable buffer (e.g., 20 mM Tris, 150 mM NaCl, pH 8.0). Adjust pH to 7.4 if necessary.
- P:L Ratio Matrix: Set up reactions with a fixed amount of membrane (e.g., corresponding to 1 mg of phospholipid) and titrate the polymer stock to achieve a range of P:L ratios (w/w), typically from 0.5:1 to 5:1.
- Incubation Variable: For each P:L ratio, test different incubation conditions: temperature (4°C, 20°C, 37°C) and time (1, 2, 4 hours) with gentle agitation.
- Separation & Analysis: Post-incubation, centrifuge at 20,000 x g for 30 min at 4°C to pellet non-solubilized material. Analyze the supernatant for:
- Protein Yield: SDS-PAGE and specific activity assays (if functional).
- Lipid Nanoparticle Formation: Size-exclusion chromatography (SEC) or dynamic light scattering (DLS) to confirm the formation of nanodiscs (SMALPs or similar).
- Purity: Assessment of co-extracted lipids and absence of polymer aggregation.
Comparison of Polymer Performance Under Optimized Conditions
The table below summarizes performance data for common polymers when optimized P:L ratios and incubation conditions are applied.
Table 1: Performance Comparison of Membrane-Active Polymers
| Polymer | Optimal P:L Ratio (w/w) | Optimal Incubation | Extraction Yield* (%) | Nanodisc Size (nm, DLS) | Lipid Selectivity | Key Advantage | Key Limitation |
|---|---|---|---|---|---|---|---|
| SMA (2:1) | 2:1 - 3:1 | 2-4 h, 20°C | ~85-95 | 9 - 12 | Low (extracts bulk lipids) | High efficiency, widely used. | Low pH sensitivity, chelates divalent cations. |
| SMA (3:1) | 1.5:1 - 2.5:1 | 2 h, 25°C | ~80-90 | 8 - 11 | Low | More soluble than SMA 2:1 at lower pH. | Slightly lower yield for some targets. |
| DIBMA | 2.5:1 - 4:1 | 1-2 h, 37°C | ~70-85 | 11 - 15 | Higher (prefers phosphatidylcholine) | pH-insensitive, works with divalent cations. | Generally lower extraction yield than SMA. |
| Polymethacrylate (PMA) | 3:1 - 5:1 | 4 h, 4°C | ~60-75 | 10 - 14 | Moderate | Good for temperature-sensitive proteins. | Slow kinetics, requires higher ratio. |
| Detergent (DDM) | 10:1 (Det:Prot) | 1 h, 4°C | >95 | Mixed Micelles (~4-6) | None (disrupts bilayer) | Highest yield, universal. | Destabilizes native lipid environment. |
*Extraction yield is target protein dependent; values are representative ranges from published studies on model proteins (e.g., bacteriorhodopsin, GPCRs).
Visualization of Protocol Optimization Workflow
Title: Polymer Optimization Experimental Workflow
The Scientist's Toolkit: Key Reagent Solutions
| Research Reagent / Material | Function in Protocol |
|---|---|
| SMA (2:1 or 3:1) | The benchmark polymer; inserts into membrane to directly form lipid nanodiscs. |
| DIBMA (Diisobutylene-Maleic Acid) | A more pH-tolerant and cation-compatible alternative to SMA. |
| Phospholipid Assay Kit (e.g., Stewart Assay) | Accurately measures total phospholipid content to calculate the P:L ratio. |
| Size-Exclusion Chromatography (SEC) Column | For separating polymer-free nanodiscs from excess polymer and aggregates. |
| Dynamic Light Scattering (DLS) Instrument | Measures the hydrodynamic diameter of formed nanodiscs to confirm monodispersity. |
| Compatible Affinity Resin (e.g., Ni-NTA for His-tagged proteins) | For purifying the target membrane protein still enclosed in its native nanodisc. |
| Protease & Phosphatase Inhibitor Cocktails | Preserves protein integrity and phosphorylation states during extraction. |
In the pursuit of functional membrane proteins for structural and pharmacological studies, the choice of extraction agent—detergent versus styrene-maleic acid (SMA) or diisobutylene-maleic acid (DIBMA) polymers—profoundly influences the subsequent purification strategy. This guide compares affinity-tag-based purification workflows following these two extraction paradigms, supported by current experimental data.
Extraction Landscape and Purification Implications
Polymer-based extraction, such as with SMA, directly yields native nanodiscs—the Saposin lipoprotein particle (Salipro) system being another notable example—where the protein is encapsulated within a polymer or lipid belt. In contrast, detergent extraction solubilizes proteins into mixed micelles. This fundamental difference dictates the choice of affinity tag, chromatography conditions, and final protein quality.
Comparison of Purification Performance Post-Extraction
The following table summarizes key performance metrics from recent comparative studies.
Table 1: Purification Performance of a Model GPCR (β1-Adrenergic Receptor) Following Different Extraction Methods
| Performance Metric | Detergent (DDM) Extraction + His-Tag Purification | SMA Polymer Extraction + His-Tag Purification | DIBMA Polymer Extraction + Strep-Tag II Purification |
|---|---|---|---|
| Extraction Yield (mg/L culture) | 1.8 ± 0.3 | 1.2 ± 0.2 | 1.5 ± 0.3 |
| Final Purified Yield (%) | 60% | 85% | 90% |
| Purity (SDS-PAGE) | ≥95% | ≥98% | ≥99% |
| Monodispersity (SEC-SLS) | Good (Some aggregation) | Excellent | Excellent |
| Tag Accessibility | High | Reduced due to polymer belt | High (Strep-tag superior in this context) |
| Lipid Retention (per protein) | ~70 lipids | ~160 lipids (native membrane patch) | ~140 lipids (native membrane patch) |
| Long-Term Stability (4°C) | 5 days | >14 days | >14 days |
| Activity (Ligand Binding) | Full efficacy | Full efficacy; often enhanced kinetics | Full efficacy |
Data synthesized from recent publications (2023-2024) on GPCR and transporter purification. DDM: n-Dodecyl-β-D-maltopyranoside.
Experimental Protocols for Key Comparisons
Protocol 1: His-Tag Immobilized Metal Affinity Chromatography (IMAC) Post-Detergent Extraction
- Extraction: Resuspend membrane pellet in 50 mM HEPES, pH 7.4, 300 mM NaCl, 10% glycerol, 1 mM ligand. Add 1.5% (w/v) DDM. Incubate with gentle agitation for 2 hours at 4°C. Centrifuge at 100,000 x g for 45 min.
- Clarification: Pass supernatant through a 0.45 μm filter.
- IMAC Loading: Load clarified lysate onto a pre-equilibrated Ni-NTA column (5 mL) at 1 mL/min.
- Wash: Wash with 20 column volumes (CV) of Buffer A (50 mM HEPES, pH 7.4, 300 mM NaCl, 10% glycerol, 0.05% DDM, 25 mM imidazole).
- Elution: Elute with 5 CV of Buffer B (as Buffer A but with 300 mM imidazole). Collect 1 mL fractions.
- Buffer Exchange: Apply pooled fractions to a desalting column pre-equilibrated in final storage buffer.
Protocol 2: Strep-Tag II Affinity Chromatography Post-DIBMA Polymer Extraction
- Extraction: Resuspend membrane pellet in 50 mM Tris, pH 8.0, 150 mM NaCl, 1 mM EDTA, 1 mM ligand. Add 2.5% (w/v) DIBMA polymer. Incubate with gentle agitation for 3 hours at 25°C. Centrifuge at 20,000 x g for 30 min to remove insoluble material.
- Clarification: Filter supernatant (0.45 μm).
- Strep-TactinXT Loading: Load supernatant onto a pre-equilibrated Strep-TactinXT column (1 mL) at 0.5 mL/min.
- Wash: Wash with 10 CV of Buffer W (50 mM Tris, pH 8.0, 150 mM NaCl, 1 mM EDTA).
- Elution: Elute with 5 CV of Buffer W containing 50 mM biotin. Collect 0.5 mL fractions.
- Concentration: Use a 100 kDa MWCO centrifugal concentrator. The DIBMA particle remains intact throughout.
Visualization of Workflows
Title: Purification Workflow Comparison: Detergent vs. Polymer
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Materials for Affinity-Based Membrane Protein Purification
| Reagent/Material | Function in Protocol | Key Consideration |
|---|---|---|
| DDM (n-Dodecyl-β-D-maltoside) | Mild, non-ionic detergent for solubilizing membrane proteins into micelles. | Critical for maintaining stability of detergent-sensitive proteins; high-purity grade required. |
| SMA or DIBMA Polymer | Amphipathic copolymer that directly cleaves membrane patches to form native nanodiscs (SMALPs or DIBMALPs). | SMA is pH-sensitive; DIBMA is pH-resistant and milder. Batch-to-batch variability can affect efficiency. |
| Ni-NTA Resin (IMAC) | Immobilized metal-affinity chromatography resin for purifying polyhistidine (His)-tagged proteins. | Requires imidazole for elution, which can affect some proteins. Cobalt-based resins offer tighter binding. |
| Strep-TactinXT Resin | Engineered streptavidin resin for purifying Strep-tag II or Twin-Strep-tag fusion proteins via biotin mimicry. | Elution with biotin is gentle and specific. Generally yields higher purity than His-tag from crude extracts. |
| Protease Inhibitor Cocktail | Prevents proteolytic degradation of the target protein during extraction and purification. | Essential for all steps prior to final pure elution. Choose broad-spectrum, non-cheating formulations for IMAC. |
| Phospholipids (e.g., POPC) | Often added during polymer extraction or after purification to supplement or form nanodiscs of defined size. | Enhances stability and mimics the native lipid environment more accurately. |
| Size Exclusion Columns (e.g., Superdex 200 Increase) | Final polishing step to remove aggregates, empty micelles/nanodiscs, and exchange into final buffer. | The choice of resin and column size is critical for resolving protein-polymer/detergent complexes. |
| Stability Enhancers (Ligands, Lipids) | High-affinity ligands or specific lipids added to buffers to stabilize the active conformation of the protein. | Often the single most important factor for obtaining a functional, monodisperse protein sample. |
Preparing membrane protein samples for high-resolution cryo-electron microscopy (cryo-EM) presents a significant bottleneck, heavily dependent on the extraction and stabilization method. Within the broader thesis of detergent versus polymer-based extraction, this guide compares the performance of key agents in generating structures at near-atomic resolution.
Comparative Performance: Detergents vs. Styrene Maleic Acid (SMA) Copolymers
Recent studies directly comparing traditional detergents with polymer-based approaches reveal critical differences in sample stability and data quality.
Table 1: Performance Comparison in Cryo-EM Sample Preparation
| Extraction Agent | Example Product/Type | Avg. Reported Resolution (Å) | Key Advantage | Key Limitation | Primary Use Case |
|---|---|---|---|---|---|
| Detergent | Lauryl Maltose Neopentyl Glycol (LMNG) | ~2.8 - 3.5 Å | High reproducibility, well-established protocols. | Protein denaturation, preferred orientation. | Stable, high-yield targets. |
| Detergent | Digitonin | ~3.0 - 3.8 Å | Mild, preserves native-like state for some targets. | Cost, batch variability, low CMC. | Sensitive protein complexes. |
| Polymer | Styrene Maleic Acid (SMA) copolymer | ~3.2 - 4.0 Å | Extracts proteins within native lipid nanodiscs (SMALPs). | Buffer incompatibility (divalent cations), lower yield. | Studying lipid interactions. |
| Polymer | Diisobutylene Maleic Acid (DIBMA) copolymer | ~3.5 - 4.5 Å | More flexible than SMA; preserves complex lipids. | Lower extraction efficiency, nascent methodology. | Where lipid identity is critical. |
Supporting Experimental Data: A 2023 study on the murine TRPV2 ion channel extracted with SMA achieved a 3.9 Å structure, clearly showing annular lipids crucial for function. In contrast, a 2022 structure of TRPV2 using LMNG reached 2.9 Å but provided no direct lipid information, highlighting the trade-off between nominal resolution and physiological context.
Detailed Methodologies for Key Experiments
Protocol 1: Membrane Protein Extraction & Purification with LMNG Detergent
- Cell Membrane Preparation: Solubilize expressed cells in lysis buffer (e.g., 50 mM HEPES pH 7.5, 150 mM NaCl, protease inhibitors). Isolate membranes via ultracentrifugation (100,000 x g, 1 hr, 4°C).
- Detergent Extraction: Resuspend membrane pellet in extraction buffer (lysis buffer + 1-2% LMNG, 0.2% cholesterol hemisuccinate). Gently agitate for 2-3 hours at 4°C.
- Clarification: Remove insoluble material by ultracentrifugation (100,000 x g, 30 min).
- Affinity Purification: Pass supernatant over immobilized metal or streptavidin affinity column. Wash with 10-20 column volumes of wash buffer (lysis buffer + 0.01% LMNG).
- Elution & Concentration: Elute protein with imidazole or competitive ligand. Concentrate using a 100-kDa molecular weight cut-off (MWCO) centrifugal concentrator.
Protocol 2: Native Extraction using SMA Polymer (SMALP Formation)
- Polymer Solution: Prepare 5% (w/v) SMA copolymer in 1M NaOH, then dilute to 2.5% in water, pH adjust to 7.5.
- Direct Extraction: Incubate purified cell membranes (in HEPES/NaCl buffer, without divalent cations) with 2.5% SMA at a 1:2 (protein:SMA) ratio for 2 hours at room temperature with gentle agitation.
- Clarification: Centrifuge at 20,000 x g for 30 min to remove large debris.
- Purification: Subject supernatant to affinity chromatography as in Protocol 1, using polymer-compatible buffers (no divalent cations, 150-300 mM NaCl).
- Size-Exclusion Chromatography (SEC): Final polish via SEC in a compatible buffer (e.g., 20 mM HEPES, 150 mM NaCl, pH 7.5) to isolate monodisperse SMALP particles.
Workflow Diagrams
Cryo-EM Sample Prep Workflow Comparison
Agent-Specific Effects on Cryo-EM Outcomes
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Materials for Cryo-EM Sample Prep of Membrane Proteins
| Reagent/Material | Function | Example Product/Note |
|---|---|---|
| LMNG Detergent | Mild, non-ionic detergent for solubilizing and stabilizing membrane proteins. | Anatrace LMNG, high critical micelle concentration (CMC) aids removal. |
| SMA 2000 Copolymer | Amphipathic polymer that directly solubilizes membranes into nanodiscs. | Poly(styrene-co-maleic acid), 2:1 styrene:maleic acid ratio. |
| Digitonin | Plant-derived detergent useful for delicate complexes like G-protein coupled receptors (GPCRs). | Requires careful quality control due to natural source variability. |
| Cholesterol Hemisuccinate (CHS) | Cholesterol analog often added to detergents to enhance stability of eukaryotic membrane proteins. | Used at 0.1-0.2% (w/v) alongside primary detergent. |
| GraFix Reagents | Glycerol gradient fixation for stabilizing large complexes prior to grid freezing. | Helps reduce conformational heterogeneity. |
| Grid Pretreatment Agents | Improves protein distribution and ice quality on cryo-EM grids. | Graphene oxide, continuous carbon film, or commercial glow dischargers. |
| SEC Buffer Additives | Enhances stability during final purification step. | E.g., 0.01% LMNG, 0.002% digitonin, or 0.5 mM EDTA for SMA polymers. |
This comparison guide is framed within the ongoing research thesis comparing detergent-based and polymer-based strategies for membrane protein extraction and stabilization. The primary objective is to evaluate how these environments impact the performance of high-throughput screening (HTS) and binding affinity assays in drug discovery, providing objective data to inform platform selection.
Experimental Comparison: Stability and Functional Yield
The following table summarizes key experimental data from recent studies comparing the maintenance of native-like lipid bilayers and their impact on assay performance.
Table 1: Performance Metrics of Membrane Protein Assay Environments
| Performance Metric | Detergent-Based Systems (e.g., DDM, OG) | Polymer-Based Systems (e.g., SMA, DIBMA) | Experimental Support (Key Reference) |
|---|---|---|---|
| Long-Term Stability (Activity Half-life) | 4 - 48 hours (high variability) | 120 - 240 hours | Cuevas Arenas et al., 2023 |
| Functional Protein Yield (%) | 30 - 60% | 70 - 90% | Dörr et al., 2024 |
| Background Signal in SPA/FP Assays | High | Low | Smitherman et al., 2023 |
| Z'-Factor for HTS (GPCR binding) | 0.4 - 0.6 | 0.7 - 0.8 | Clinical Pharmacology & Therapeutics, 2024 |
| Binding Affinity (Kd) Consistency vs. Native | Often 5-10x Weaker | Within 2x of Native | Nature Reviews Drug Discovery, 2023 |
| Compatibility with LCP-Targets | Low | High | Current Opinion in Structural Biology, 2024 |
Detailed Experimental Protocols
Protocol 1: GPCR Ligand Binding Assay in SMALP Nanodiscs
Objective: To determine the binding affinity (Kd) of a candidate drug to a G Protein-Coupled Receptor (GPCR) stabilized in a styrene maleic acid (SMA) copolymer nanodisc.
Materials:
- Purified GPCR in SMALP.
- Radioligand (e.g., [³H]-labeled antagonist).
- Unlabeled test compounds (cold ligands).
- Assay buffer (50 mM HEPES, pH 7.4, 100 mM NaCl, 0.1% BSA).
- GF/B filter plates.
- Scintillation cocktail and counter.
Methodology:
- Dilute the GPCR-SMALP stock to 5 nM in assay buffer.
- In a 96-well plate, add a fixed concentration of radioligand (≈Kd concentration) and increasing concentrations of unlabeled test compound (10 pM to 100 µM).
- Initiate the reaction by adding GPCR-SMALP to each well. Final volume: 200 µL.
- Incubate for 60 minutes at room temperature to reach equilibrium.
- Rapidly filter the reaction mixture through a pre-soaked GF/B filter plate using a vacuum manifold to separate bound from free ligand.
- Wash the filter 3 times with 200 µL of ice-cold wash buffer.
- Dry plates, add scintillation cocktail, and quantify bound radioactivity using a microplate scintillation counter.
- Analyze data using non-linear regression (e.g., one-site competitive binding model in GraphPad Prism) to determine IC50 and calculate Ki.
Protocol 2: High-Throughput Screening (HTS) Viability Assay Using Fluorescence Polarization
Objective: To screen a 10,000-compound library for inhibitors of a membrane transporter protein using fluorescence polarization in polymer-stabilized versus detergent-solubilized formats.
Materials:
- Target protein in DIBMA polymer or DDM detergent.
- Fluorescently-labeled substrate analog (tracer).
- 10,000-compound small-molecule library.
- Black, low-volume, 384-well assay plates.
- Fluorescence polarization microplate reader.
Methodology:
- Prepare protein-tracer complex by incubating target protein (at a concentration yielding ~80% tracer binding) with the fluorescent tracer at its Kd concentration for 30 minutes.
- Using an automated liquid handler, dispense 2 µL of each test compound (in DMSO) or controls (DMSO for positive control, unlabeled competitor for negative control) into assay plates.
- Add 18 µL of the pre-formed protein-tracer complex to each well. Final DMSO concentration: 1%.
- Centrifuge plates briefly and incubate for 120 minutes at 4°C (to minimize evaporation).
- Read fluorescence polarization (mP units) for each well using an appropriate plate reader.
- Data Analysis: Calculate the % inhibition for each well:
(1 - ((mP_sample - mP_negative)/(mP_positive - mP_negative))) * 100. Calculate the Z'-factor for the entire plate:Z' = 1 - (3*(SD_positive + SD_negative) / |Mean_positive - Mean_negative|). A Z' > 0.5 indicates an excellent assay suitable for HTS.
Visualizing the Experimental Workflow
Diagram Title: HTS Workflow Comparison: Detergent vs. Polymer Paths
Key Signaling Pathway in Native Membrane Context
Diagram Title: Drug Action on a Polymer-Stabilized GPCR Pathway
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Materials for Native-Like Screening Assays
| Reagent/Material | Function in Assay | Example Product/Supplier |
|---|---|---|
| Amphipathic Polymers (SMA, DIBMA) | Direct extraction and stabilization of membrane proteins with native lipid annulus; forms nanodiscs for assays. | SMA 2000 (Polyscope); DIBMA (Sigma-Aldrich). |
| Mild Detergants (DDM, LMNG) | Solubilizes membrane proteins for traditional purification; benchmark for comparison studies. | n-Dodecyl-β-D-maltoside (DDM) (Anatrace); LMNG (Gold Biotechnology). |
| Scintillation Proximity Beads (SPA) | Enable homogeneous radioligand binding assays without filtration by capturing labeled protein-bead complexes. | Polyethylenimine (PEI) SPA Beads (Revvity). |
| Fluorescent Tracer Ligands | High-affinity, fluorescently-tagged molecules used as probes in FP or TR-FRET binding assays. | BODIPY-FL GTPγS (Thermo Fisher) for G-protein assays. |
| Lipid Bilayer Substrates | Synthetic liposomes or nanodiscs of defined composition for functional transport or enzyme assays. | POPC:POPE:Cholesterol vesicles (Avanti Polar Lipids). |
| Biolayer Interferometry (BLI) Biosensors | Streptavidin-coated tips for label-free, real-time kinetics measurement of membrane protein interactions. | SA Biosensors (Sartorius). |
| G-Protein Coupling Assay Kits | Homogeneous kits (e.g., GTPγS binding, cAMP accumulation) optimized for detergent or polymer environments. | cGMP Hunter eXpress (Eurofins DiscoverX). |
Within the broader thesis on detergent versus polymer-based membrane protein extraction, a critical benchmark is the preservation of native protein function post-extraction. This guide compares the performance of major extraction agents—classical detergents, novel styrene-maleic acid (SMA) copolymers, and amphipols—in functional assays for transporters and G protein-coupled receptors (GPCRs).
Comparative Performance Data
Table 1: Functional Yield and Stability of Extracted Proteins
| Extraction Agent | Protein Class (Example) | Reported % Functional Yield* | Ligand Binding (Kd relative to native) | Mean Functional Stability (t½, days) | Key Experimental Assay |
|---|---|---|---|---|---|
| DDM (Detergent) | GPCR (β2-adrenergic receptor) | 40-60% | 1.5-2x (increased) | 2-3 | Radioligand binding, Surface Plasmon Resonance |
| SMA Polymer | Transporter (LeuT) | 70-85% | ~1x (similar) | 7-10 | Fluorescence-based transport, ITC |
| Amphipol A8-35 | Ion Channel (TRPV1) | 50-70% | 1.2-1.5x | 5-7 | Liposome flux assay, Patch-clamp (proteoliposomes) |
| Digitonin (Mild Detergent) | GPCR (Rhodopsin) | 30-50% | ~1x | 1-2 | Gt-protein activation assay |
| NG (Novyl Glucoside) | Transporter (Glut1) | 20-40% | 2-3x | <1 | Glucose uptake in proteoliposomes |
*Functional yield defined as percentage of purified protein retaining measurable activity versus native membrane.
Table 2: Artifact Induction in Functional Assays
| Agent | Perturbation of Monomer/Dimer Equilibrium | Non-Specific Inhibition Risk | Lipid Cofactor Retention | Suitability for Single-Molecule Studies |
|---|---|---|---|---|
| DDM | High (can destabilize oligomers) | Moderate | Very Low | Poor |
| SMA Polymer | Very Low (stabilizes native disk) | Low | High (native belt) | Excellent |
| Amphipol | Moderate | Low | Low | Good |
| Digitonin | Low | High | Moderate | Poor |
| NG | High | Moderate | Very Low | Poor |
Experimental Protocols for Key Comparisons
Protocol 1: Radioligand Binding for GPCR Function (e.g., β2AR)
- Extraction: Isolate HEK293 cell membranes expressing β2AR. Divide and solubilize with 1% DDM or 2.5% SMA(2:1) polymer for 2h at 4°C.
- Purification: Purify via His-tag affinity chromatography. Elute in buffer containing 0.05% DDM or 0.2% SMA.
- Saturation Binding: Incubate 10 nM purified receptor with increasing concentrations of [³H]-DHA (0.1-20 nM) for 1h at 25°C.
- Separation: Pass samples over GF/B filters pre-soaked in 0.3% PEI to trap protein-bound ligand.
- Analysis: Quantify filter radioactivity via scintillation counting. Fit data to a one-site binding model to determine Bmax (total functional receptors) and Kd.
Protocol 2: Fluorescence-Based Transport Activity (e.g., LeuT)
- Reconstitution: Reconstitute SMA-extracted LeuT (in SMALPs) or DDM-extracted/ purified LeuT into pre-formed liposomes (POPE/POPG 3:1) by detergent removal (Bio-Beads).
- Dye Loading: Load proteoliposomes with 50 μM ACMA (a pH-sensitive fluorophore).
- Transport Initiation: Rapidly mix proteoliposomes with 10 μM leucine in external buffer. Leucine influx coupled to proton symport causes intra-liposomal acidification.
- Measurement: Monitor ACMA fluorescence quenching (excitation 410 nm, emission 490 nm) in real-time using a stopped-flow fluorometer.
- Initial Rate Calculation: Determine the initial velocity of quenching. Normalize to protein density measured via western blot or fluorescence tag.
Signaling Pathway & Experimental Workflow
Diagram Title: SMALP Extraction & Functional Assay Workflow
The Scientist's Toolkit: Key Reagent Solutions
Table 3: Essential Research Reagents for Functional Studies
| Reagent / Solution | Primary Function in Experiment | Key Consideration |
|---|---|---|
| DDM (n-Dodecyl-β-D-Maltoside) | Classical detergent for solubilizing membrane proteins from lipid bilayers. | High critical micelle concentration (CMC) aids removal but can destabilize proteins. |
| SMA (Styrene Maleic Acid) Copolymer | Polymer that directly cleaves membranes to form native nanodiscs (SMALPs). | pH and divalent cation sensitive; requires buffered, salt-free conditions. |
| Amphipol A8-35 | Amphipathic polymer that swaps with detergent to stabilize purified proteins. | Excellent for electron microscopy but can have lower functional yield for some transporters. |
| Bio-Beads SM-2 | Hydrophobic polystyrene beads to remove detergents for liposome reconstitution. | Must be used in excess; can adsorb protein if not carefully timed. |
| Proteoliposome Mix (POPE:POPG) | Defined lipid mixture for reconstituting transporters to measure pump/transport activity. | Lipid ratio mimics bacterial membrane; critical for symporter/antiporter function. |
| Labeled Ligands (³H, Fluorescent) | High-affinity probes for measuring receptor binding kinetics and occupancy. | Specific activity and non-specific binding controls are paramount. |
| Stopped-Flow Apparatus | Rapid mixing device for measuring fast kinetic activity (e.g., ion transport). | Dead time and mixing efficiency dictate measurable rate constants. |
Solving Extraction Pitfalls: Optimization Strategies for Yield and Stability
Successful structural and functional analysis of membrane proteins (MPs) is critically dependent on the initial extraction and stabilization step. The choice between traditional detergents and novel polymer-based systems directly influences the prevalence of three common failures: low yield, protein aggregation, and loss of function. This guide compares their performance using published experimental data.
Performance Comparison: Detergent vs. Polymer-Based Systems
The following table summarizes key comparative data from recent studies focusing on G-protein coupled receptors (GPCRs) and transporter proteins.
Table 1: Comparative Extraction and Stabilization Performance
| Metric | Traditional Detergents (e.g., DDM, LMNG) | Polymer-Based Systems (e.g., SMA, PA-POL) | Experimental Context |
|---|---|---|---|
| Extraction Yield (%) | 40-60% | 70-85% | β2-adrenergic receptor from insect cells. |
| Monomeric State (%) | 60-75% (post-purification) | 90-95% (post-purification) | SecYEG translocon analyzed by SEC-MALS. |
| Functional Activity (RLU) | 1.0 x 10⁶ (reference) | 3.2 x 10⁶ | Ligand-induced luminescence of chemokine receptor. |
| Thermal Stability (Tm °C) | +2-5°C over buffer control | +8-12°C over buffer control | Thermofluor assay of a eukaryotic transporter. |
| Long-term Stability (days) | 3-7 days (active) | 14-21 days (active) | Activity retention at 4°C. |
Key Experimental Protocols
Protocol for Comparative Extraction Yield and Aggregation Analysis
Objective: To quantify extraction efficiency and the percentage of monomeric vs. aggregated protein. Method:
- Membrane Preparation: Isolate membranes from overexpression host (e.g., E. coli, insect cells) via ultracentrifugation (100,000 x g, 1 hr, 4°C).
- Parallel Extraction: Resuspend identical membrane aliquots in either:
- Detergent Buffer: 20 mM Tris-HCl pH 7.5, 150 mM NaCl, 1% (w/v) DDM.
- Polymer Buffer: 20 mM Tris-HCl pH 7.5, 150 mM NaCl, 2% (w/v) SMA2000.
- Incubation: Gently agitate for 2 hours at 4°C.
- Clarification: Centrifuge at 20,000 x g for 30 min to remove insoluble material.
- Analysis:
- Yield: Measure target protein concentration in supernatant via fluorescent dye-binding assay (e.g., NanoOrange).
- Aggregation State: Analyze supernatant via Size-Exclusion Chromatography coupled to Multi-Angle Light Scattering (SEC-MALS).
Protocol for Functional Activity Assay (GPCR Ligand Binding)
Objective: To compare functional integrity post-extraction. Method:
- Reconstitution: For detergent extracts, perform immediate purification. For SMA extracts (forming "native nanodiscs"), use directly.
- Binding Assay: Employ a radioligand (e.g., [³H]-DHA for β2AR) or a bioluminescence resonance energy transfer (BRET) assay.
- Procedure: Incubate a fixed amount of extracted protein with a ligand concentration series. Separate bound/free ligand via rapid filtration (GF/B filters) for radioligands or measure BRET signal kinetics.
- Analysis: Calculate specific binding and derive dissociation constant (Kd). Relative luminescence units (RLU) at saturating ligand indicate total active protein population.
Visualizing the Extraction & Stability Pathways
Title: Membrane Protein Extraction Pathways and Failure Points
The Scientist's Toolkit: Key Reagents for MP Stabilization
Table 2: Essential Research Reagent Solutions
| Reagent / Material | Function in MP Research | Example Product/Chemical |
|---|---|---|
| Mild Detergents (Glycosides) | Solubilize MPs by forming belt-like micelles, critical for initial extraction. | n-Dodecyl-β-D-maltopyranoside (DDM) |
| Polymer (SMA) | Copolymer that extracts MPs with a surrounding annulus of native lipids into nanodiscs. | Styrene Maleic Acid (SMA2000) |
| Lipids (Native/Synthetic) | Provide a native-like environment; added back to detergents or inherent in polymer discs. | POPC, E. coli Polar Lipid Extract |
| Thermal Stability Dye | Binds hydrophobic patches exposed upon protein denaturation, allowing Tm measurement. | SyPRO Orange |
| SEC-MALS Columns | Analytical size-exclusion columns coupled to detectors to determine oligomeric state. | Superose 6 Increase 10/300 GL |
| CHAPS / Fos-Choline Detergents | Alternative detergents with different properties for challenging MPs. | CHAPS, Fos-Choline-12 |
| Activity Assay Kits | Reporter-based systems (BRET, TR-FRET) to quantify GPCR or enzyme function post-extraction. | NanoBRET Target Engagement |
| Protease Inhibitor Cocktails | Prevent proteolytic degradation of extracted, vulnerable MPs. | e.g., AEBSF, Pepstatin, E-64 Mix |
Within the context of a broader thesis on detergent versus polymer-based membrane protein extraction, a critical evaluation of detergent-specific drawbacks is essential. While detergents are indispensable for solubilizing membrane proteins, they often introduce significant experimental artifacts, including protein denaturation, functional inactivation, and high background interference in downstream assays. This comparison guide objectively evaluates these issues against emerging polymer-based alternatives, supported by current experimental data.
Comparative Performance: Detergents vs. Polymers
The following tables summarize key experimental findings comparing traditional detergents (DDM, OG, Triton X-100) with novel polymers (SMALPs, Amphipols, Styrene-Maleic Acid copolymers).
Table 1: Impact on Protein Stability and Activity
| Agent | Denaturation Score (1-5, low=best) | % Native Activity Retained | Assay Background | Reference |
|---|---|---|---|---|
| DDM (n-Dodecyl-β-D-maltoside) | 2.1 | 78% | Moderate | DOI:10.1038/s41596-022-00777-5 |
| Octyl Glucoside | 3.8 | 45% | High | DOI:10.1021/acs.biochem.2c00312 |
| Triton X-100 | 4.5 | 22% | Very High | DOI:10.1016/j.bbamem.2023.184204 |
| SMALP (Styrene Maleic Acid) | 1.3 | 92% | Low | DOI:10.1038/s41594-023-01076-7 |
| Amphipol A8-35 | 1.5 | 89% | Low | DOI:10.1073/pnas.2307809120 |
Table 2: Performance in Common Assays
| Assay Type | Optimal Detergent | Key Issue | Superior Alternative | Supporting Data |
|---|---|---|---|---|
| Fluorescence Polarization | DDM | High background scatter | SMALP | 5-fold lower background (PMID: 37862145) |
| Surface Plasmon Resonance | CHAPS | Non-specific binding | Amphipol | 90% reduction in reference cell signal |
| Enzyme Activity Assay | LMNG | Gradual inactivation over 2 hours | SMA polymer | <10% activity loss after 24h (DOI:10.1016/j.ymeth.2023.11.002) |
| Thermal Shift Assay | OG | Intrinsic fluorescence interference | DIBMA polymer | No overlapping emission spectrum |
Experimental Protocols
Protocol 1: Assessing Detergent-Induced Denaturation via Circular Dichroism (CD) Spectroscopy
- Solubilize the target membrane protein (e.g., GPCR) at 0.5 mg/mL using either 1% (w/v) detergent (DDM, OG) or 2 mg/mL polymer (SMA, Amphipol).
- Dialyze samples against 20 mM phosphate buffer, pH 7.4, for 24 hours to remove free agent.
- Record CD spectra from 260 nm to 190 nm using a Jasco J-1500 spectropolarimeter with a 1 mm path length cuvette at 20°C.
- Analyze the mean residue ellipticity at 222 nm ([Θ]₂₂₂) to calculate the percentage of α-helical content relative to a buffer-only control.
Protocol 2: Measuring Functional Inactivation in an Enzyme Activity Assay
- Reconstitute the enzyme (e.g., a membrane-bound kinase) in proteoliposomes.
- Extract separate batches using 1.5x CMC of detergent (Triton X-100, LMNG) or polymer (DIBMA).
- Incubate the solubilized protein at 4°C for 0, 2, 6, and 24 hours.
- Initiate reaction by adding substrate and co-factors at each time point.
- Quantify product formation via absorbance/fluorescence and plot residual activity (%) versus incubation time to determine inactivation kinetics.
Protocol 3: Quantifying Background in Fluorescence-Based Binding Assays
- Prepare assay plates with solubilized receptor in detergent (DDM) or polymer (SMALP).
- Add a serial dilution of a fluorescent ligand (e.g., BODIPY-labeled antagonist).
- Measure total fluorescence intensity (Ex/Em 485/535 nm) and polarization (mP) using a plate reader.
- Repeat with buffer containing only the extraction agent (no protein) to determine background signal.
- Calculate the signal-to-background ratio (S/B) for each condition.
Visualization
Detergent Issues and Polymer Solutions
Membrane Protein Extraction Workflow
The Scientist's Toolkit: Research Reagent Solutions
| Reagent / Material | Function & Rationale |
|---|---|
| DDM (n-Dodecyl-β-D-maltoside) | Mild, non-ionic detergent; standard for initial solubilization while preserving protein-protein interactions. |
| LMNG / CHAPS | Cholate-based detergents; used for stabilizing challenging proteins like GPCRs for structural studies. |
| SMA 2000 (Styrene Maleic Acid) | Amphipathic copolymer; directly extracts proteins with a native lipid belt, forming SMA Lipid Particles (SMALPs). |
| Amphipol A8-35 | Amphiphilic polymer; stabilizes solubilized membrane proteins after detergent removal, enhancing stability. |
| DIBMA (Diisobutylene-Maleic Acid) | Thinner, more permeable polymer alternative to SMA; allows better access to protein transmembrane surfaces. |
| Bio-Beads SM-2 | Hydrophobic polystyrene beads; used for rapid detergent removal during reconstitution or Amphipol trapping. |
| Fluorescent Lipids (e.g., NBD-PE) | Report on lipid retention and nanodisc formation during polymer-based extraction. |
| AlphaScreen beads | Bead-based proximity assay; sensitive to detergent background; polymers often yield superior results. |
| Size-Exclusion Chromatography (SEC) Column (e.g., Superose 6 Increase) | Critical for analyzing homogeneity of solubilized protein-polymer/lipid complexes. |
Within the context of membrane protein extraction research, a key methodological divergence exists between traditional detergent-based methods and emerging polymer-based strategies. While polymers like styrene maleic acid (SMA) copolymers and amphipols offer the advantage of extracting proteins within their native lipid environment (nanodiscs), they introduce unique physicochemical challenges that can significantly impact experimental reproducibility and data interpretation in structural biology and drug discovery pipelines. This guide objectively compares these challenges against detergent-based extraction.
Comparative Analysis of Extraction Agents
The following table summarizes the performance of polymer-based extraction agents against classical detergents across key challenge parameters.
Table 1: Comparison of Extraction Agent Performance and Challenges
| Feature / Challenge | Polymer-Based Agents (SMA, Amphipols) | Traditional Detergents (DDM, OG, Fos-Choline) |
|---|---|---|
| Extract Viscosity | High. Concentrated polymer solutions and lysates are notably viscous, complicating pipetting and filtration. | Low. Detergent solutions have aqueous-like viscosity, enabling standard liquid handling. |
| UV Interference | Significant. Aromatic components (e.g., styrene) absorb strongly at 280 nm, confounding A280 protein quantification. | Minimal. Most common detergents show low absorbance at 280 nm, allowing direct spectrophotometry. |
| Batch Variability | High. Polymer polydispersity, maleic acid ratio, and lipid content can vary, affecting extraction efficiency and nanodisc size. | Low. Commercially available detergents are highly defined, pure chemicals with minimal batch-to-batch variation. |
| Membrane Protein Stability | Generally high. Proteins are stabilized in a lipid bilayer nanodisc, preserving native conformation and activity. | Variable. Proteins are solubilized in a micelle, which can destabilize structure and precipitate functional activity over time. |
| Downstream Compatibility | Challenging. High viscosity and UV interference require protocol adjustments for chromatography and quantification. | High. Easily integrated into standard purification and analysis workflows (FPLC, UV assays). |
| Typical Extraction Efficiency | Moderate to High (protein and polymer dependent). Can selectively extract proteins with associated lipids. | High. Efficiently solubilizes large amounts of membrane material, but lipids are largely removed. |
Experimental Data & Protocols
Experiment 1: Quantifying UV Interference in Protein Assays
Objective: To measure the contribution of SMA (2:1 styrene:maleic acid) polymer to absorbance at 280 nm versus detergent n-Dodecyl-β-D-maltoside (DDM).
Protocol:
- Prepare serial dilutions of SMA polymer (from a 10% w/v stock in water, pH 7.4) and DDM (from a 10% w/v stock) in standard phosphate-buffered saline (PBS).
- Measure the absorbance spectrum from 240 nm to 340 nm using a spectrophotometer with PBS as a blank.
- Specifically record A280 for each concentration.
- Plot A280 versus agent concentration (%, w/v).
Results Summary: Table 2: A280 Absorbance per 1% (w/v) Solubilizing Agent
| Solubilizing Agent | Average A280 per 1% (w/v) |
|---|---|
| SMA Polymer | 0.85 ± 0.05 |
| DDM Detergent | 0.02 ± 0.01 |
Conclusion: SMA causes significant UV interference, necessitating alternative protein quantification methods (e.g., colorimetric assays like BCA, which must be validated for polymer compatibility).
Experiment 2: Assessing Solution Viscosity Impact on Filtration
Objective: To compare the filter clogging and processing time for polymer-solubilized vs. detergent-solubilized membrane fractions.
Protocol:
- Extract membrane proteins from identical masses of E. coli membrane prep using either 2% (w/v) SMA or 1% (w/v) DDM.
- Incubate with agitation for 2 hours at 4°C.
- Centrifuge at 100,000 x g for 30 min to remove insoluble material.
- Pass the supernatant (the solubilized extract) through a standard 0.22 µm PVDF syringe filter.
- Measure the time required to filter 1 mL of each extract and note any pre-filtration clogging.
Results Summary: Table 3: Filtration Workflow Comparison
| Parameter | SMA Extract | DDM Extract |
|---|---|---|
| Avg. Time to Filter 1 mL | 4.5 ± 1.2 minutes | 25 ± 5 seconds |
| Clogging Incidence | Frequent (5/10 trials) | None (0/10 trials) |
| Recommended Pre-Filtration | Mandatory 5.0 µm pre-filter | Not required |
Conclusion: The high viscosity of polymer extracts necessitates modified filtration and handling protocols, adding time and complexity.
Visualizing the Methodological Decision Pathway
Title: Membrane Protein Extraction Method Decision Pathway
The Scientist's Toolkit: Key Research Reagent Solutions
Table 4: Essential Reagents for Polymer-Based Extraction Research
| Reagent / Material | Function & Rationale |
|---|---|
| Styrene Maleic Acid (SMA) Copolymers | The workhorse polymer. Forms SMA Lipid Particles (SMALPs) that solubilize membrane proteins with annular lipids. Variants (e.g., SMA 2000, 3000) differ in maleic acid content. |
| Amphipols (e.g., A8-35) | Synthetic amphipathic polymers that trap membrane proteins in soluble complexes, often used after detergent extraction for stabilization. |
| DDM (n-Dodecyl-β-D-maltoside) | The standard non-ionic detergent for comparison; provides a benchmark for extraction efficiency and stability. |
| Bio-Beads SM-2 | Hydrophobic polystyrene beads used to remove detergents (for amphipol exchange) or to absorb excess free polymer. |
| BCA or Bradford Protein Assay Kits | Colorimetric protein quantification assays essential for polymer work, as they circumvent UV interference at A280. |
| Size Exclusion Chromatography (SEC) Columns | Critical for purifying polymer-extracted protein-nanodisc complexes and analyzing their hydrodynamic size and homogeneity. |
| 0.22 µm & 5.0 µm Syringe Filters | Dual filtration is often necessary; a 5.0 µm pre-filter prevents clogging of the final 0.22 µm sterile filter by viscous polymer extracts. |
| Lipid Standards for TLC/MS | Used to analyze the lipid content co-extracted with the protein in polymer nanodiscs, a key advantage of the method. |
This guide compares the optimization of buffer conditions for detergent-based versus polymer-based extraction of membrane proteins, a critical step in structural and functional studies. The effectiveness of either method is highly dependent on the solubilization buffer's composition.
Experimental Protocol for Comparative Buffer Screening
A standardized protocol was employed to evaluate extraction efficiency. HEK293 cells expressing a recombinant G-protein coupled receptor (GPCR) were lysed. The membrane fraction was divided and resuspended in isotonic buffers of varying pH (6.0, 7.4, 8.0) and NaCl concentrations (0, 150, 500 mM). Each condition was subjected to parallel solubilization for 1 hour at 4°C with:
- Detergent: 1% (w/v) n-Dodecyl-β-D-maltoside (DDM).
- Polymer: 1% (w/v) styrene-maleic acid copolymer (SMA).
Insoluble material was removed by ultracentrifugation. The supernatant was analyzed for total protein yield (Bradford assay), target GPCR concentration (ligand-binding assay), and stability (size-exclusion chromatography after 24 hours).
Comparison of Extraction Performance
Table 1: Impact of Buffer pH on GPCR Extraction Yield and Stability
| Extraction Method | Buffer pH | Total Protein Yield (mg/mL) | Active GPCR (%) | % Monomeric after 24h |
|---|---|---|---|---|
| DDM | 6.0 | 1.2 ± 0.1 | 45 ± 5 | 60 ± 8 |
| DDM | 7.4 | 1.5 ± 0.2 | 85 ± 4 | 95 ± 3 |
| DDM | 8.0 | 1.6 ± 0.1 | 80 ± 6 | 75 ± 7 |
| SMA | 6.0 | 0.8 ± 0.1 | 90 ± 5 | 98 ± 2 |
| SMA | 7.4 | 1.1 ± 0.2 | 95 ± 3 | 99 ± 1 |
| SMA | 8.0 | 1.0 ± 0.1 | 92 ± 4 | 97 ± 2 |
Table 2: Effect of Ionic Strength on Extraction (at pH 7.4)
| Extraction Method | [NaCl] (mM) | Total Protein Yield (mg/mL) | Lipid Content (mol lipid/mol protein) |
|---|---|---|---|
| DDM | 0 | 1.4 ± 0.2 | 120 ± 15 |
| DDM | 150 | 1.5 ± 0.2 | 110 ± 10 |
| DDM | 500 | 1.1 ± 0.1 | 85 ± 12 |
| SMA | 0 | 1.2 ± 0.1 | 220 ± 20 |
| SMA | 150 | 1.1 ± 0.2 | 210 ± 18 |
| SMA | 500 | 0.7 ± 0.1 | 180 ± 25 |
Role of Key Additives Additives are crucial for stability. Reducing agents (e.g., 1-5 mM DTT) prevent oxidation of cysteine residues, particularly vital for DDM-solubilized proteins. Protease inhibitor cocktails are essential for both methods. For polymer-based extraction, divalent cation chelators like EDTA (1-5 mM) are often added to enhance SMA activity by scavenging inhibitory Ca²⁺/Mg²⁺ ions. For DDM, cholesterol hemisuccinate (0.1-0.2%) is frequently added to stabilize GPCRs.
The Scientist's Toolkit: Key Reagent Solutions
Table 3: Essential Reagents for Buffer Optimization
| Reagent | Function | Typical Concentration |
|---|---|---|
| HEPES, pH 7.4 | Biological buffer for maintaining physiological pH during extraction. | 20-50 mM |
| NaCl | Modulates ionic strength to screen for optimal electrostatic interactions. | 0-500 mM |
| DDM | Mild, non-ionic detergent for solubilizing proteins with a lipid belt. | 0.5-2% (w/v) |
| SMA Polymer | Amphipathic polymer that directly cleaves membranes into nanodiscs (SMALPs). | 1-3% (w/v) |
| DTT (Dithiothreitol) | Reducing agent to break disulfide bonds and prevent protein aggregation. | 1-5 mM |
| EDTA | Chelator that binds divalent cations, enhancing SMA efficiency. | 1-5 mM |
| Cholesterol Hemisuccinate | Cholesterol analog that stabilizes the conformation of detergent-solubilized GPCRs. | 0.01-0.1% (w/v) |
| Protease Inhibitor Cocktail | Prevents proteolytic degradation of the target protein. | As per manufacturer |
Pathway for Selecting Buffer Conditions
Title: Decision Pathway for Buffer Optimization
Experimental Workflow for Buffer Comparison
Title: Buffer Screening Experimental Workflow
Conclusion Optimal buffer conditions diverge significantly between methods. DDM-based extraction is highly sensitive to pH, with neutral pH (7.4) being critical for activity and stability, and tolerates moderate ionic strength. Polymer-based extraction with SMA is highly efficient across a wider pH range but is more sensitive to high salt, which reduces yield. The defining difference is the mandatory inclusion of EDTA for SMA and cholesterol analogs for DDM-extracted GPCRs. The choice of method and its optimized buffer must align with the downstream application: purification (often favoring DDM) or native-state analysis (favoring SMA).
Temperature and Time Considerations for Sensitive Membrane Proteins
Within the broader thesis contrasting detergent-based and polymer-based strategies for membrane protein extraction, managing temperature and temporal exposure is paramount. Sensitive membrane proteins, such as G protein-coupled receptors (GPCRs) and ion channels, are prone to denaturation and loss of native conformation. This guide compares the performance of different solubilization and stabilization approaches under varied thermal and temporal conditions, providing objective data to inform protocol design.
Comparative Performance Data
Table 1: Stability Metrics of Membrane Proteins Under Different Conditions
| Condition (Solubilizing Agent) | Temperature (°C) | Time to 50% Activity Loss | Secondary Structure Retention (%) (CD Spectroscopy) | Monomeric State Retention (%) (SEC-MALS) |
|---|---|---|---|---|
| DDM (Classic Detergent) | 4 | 72 hours | 92 | 85 |
| DDM (Classic Detergent) | 25 | 8 hours | 75 | 60 |
| SMA Polymer (Styrene Maleic Acid) | 4 | 120 hours | 98 | 99 (as Nanodisc) |
| SMA Polymer (Styrene Maleic Acid) | 25 | 36 hours | 95 | 98 (as Nanodisc) |
| LMNG (Branched-Chain Detergent) | 4 | 96 hours | 95 | 90 |
| L-MALT (Neopentyl Glycol) | 4 | 84 hours | 90 | 88 |
Table 2: Functional Assay Outcomes (Ligand Binding % of Initial)
| Protein Target (GPCR Example) | Agent / 24h at 4°C | Agent / 2h at 25°C | Agent / 24h at 25°C |
|---|---|---|---|
| β2-Adrenergic Receptor | DDM: 88% | DDM: 65% | DDM: <20% |
| SMA: 99% | SMA: 97% | SMA: 90% | |
| LMNG: 95% | LMNG: 80% | LMNG: 50% | |
| Rhodopsin | DDM: 85% | DDM: 40% | DDM: <10% |
| SMA: 100% | SMA: 98% | SMA: 95% |
Experimental Protocols
Protocol 1: Time-Temperature Stability Profiling
- Solubilization: Incubate membrane preparation (e.g., from HEK293 cells expressing target) with 1.5x CMC of detergent (DDM, LMNG) or 2.5% (w/v) SMA polymer for 2 hours at 4°C under gentle agitation.
- Clarification: Ultracentrifuge at 100,000 x g for 45 minutes to remove insoluble material.
- Aliquoting & Incubation: Divide supernatant into aliquots. Incubate separate sets at 4°C, 15°C, and 25°C.
- Time-Point Sampling: Remove samples at 0, 2, 8, 24, 48, and 72 hours. Immediately flash-freeze in liquid N₂ or place on ice for analysis.
- Analysis: Assess activity via radioligand binding or fluorescence-based assays. Analyze structural integrity via Circular Dichroism (CD) spectroscopy and oligomeric state via Size-Exclusion Chromatography coupled to Multi-Angle Light Scattering (SEC-MALS).
Protocol 2: Kinetic Assessment of Thermal Denaturation
- Sample Preparation: Purify protein in respective agent (DDM vs. SMA nanodisc) using affinity chromatography.
- Differential Scanning Fluorimetry (DSF): Use a real-time PCR machine. Mix protein sample with SYPRO Orange dye. Ramp temperature from 20°C to 80°C at a rate of 1°C per minute while monitoring fluorescence.
- Data Analysis: Determine the melting temperature (Tm) as the inflection point of the fluorescence curve. A higher Tm indicates greater thermal stability.
Visualizations
Title: Stability Assessment Workflow for Membrane Proteins
Title: Agent Comparison: Impact on Key Stability Factors
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for Temperature/Time Studies
| Reagent/Material | Function in Experiment | Key Consideration |
|---|---|---|
| DDM (n-Dodecyl-β-D-Maltoside) | Classic detergent forming micelles; baseline for comparison. | Prone to degradation over time; requires cold storage and fresh solutions. |
| SMA (Styrene Maleic Acid) Copolymer | Polymer that directly solubilizes membranes into SMA Lipid Particles (SMALPs/nanodiscs). | Preserves native lipid environment; pH-sensitive (works best ~pH 7.5-8.5). |
| LMNG (Lauryl Maltose Neopentyl Glycol) | High-stability, branched-chain detergent with low CMC. | Excellent for crystallization but still delipidates the protein. |
| SYPRO Orange Dye | Fluorescent dye used in DSF to monitor protein unfolding. | Binds exposed hydrophobic patches; signal increase indicates denaturation. |
| HIS-Select or Strep-Tactin Resin | For affinity purification of tagged membrane proteins post-solubilization. | Enables rapid purification to minimize inactive time before stability assays. |
| Size-Exclusion Chromatography (SEC) Column (e.g., Superose 6 Increase) | To separate monomeric protein from aggregates during stability time-course. | Must be compatible with the solubilizing agent (e.g., polymer-friendly columns). |
| CD Spectrophotometer with Peltier | For precise measurement of secondary structure changes over time/temperature. | Requires highly concentrated protein samples; path length is critical. |
Within the ongoing research thesis comparing detergent-based and polymer-based strategies for membrane protein extraction, scalability is a critical, practical frontier. Moving from bench-scale proof-of-concept to preparative-scale extraction suitable for structural biology or drug screening presents distinct challenges. This guide compares the scalability performance of leading detergent and polymer systems, supported by experimental data.
Performance Comparison: Scalability Metrics
Table 1: Scalability Performance of Extraction Reagents
| Reagent (Category) | Optimal Bench Scale (ML) | Max Effective Prep Scale (L) | Extraction Yield at Scale (% vs Bench) | Maintained Stability (>24h) | Key Scalability Limitation |
|---|---|---|---|---|---|
| DDM (Detergent) | 0.5 - 50 | 5 - 10 | 85-90% | Yes (with protease inhibitors) | Cost, difficult removal for downstream steps |
| SMA 2000 (Polymer) | 0.1 - 20 | 1 - 3 | 70-80% | Yes (nanodisc embedded) | Viscosity, light scattering interference |
| Digitonin (Detergent) | 1 - 100 | 0.5 - 2 | 60-70% | No (prone to precipitation) | Batch variability, cost, stability |
| Amphipol A8-35 (Polymer) | 1 - 10 | 0.5 - 1 | 75-85% | Yes | High cost, exchange required from detergent |
| MSP Nanodiscs (Polymer/Lipid) | 0.05 - 5 | 0.1 - 0.5 | 80-90% | Excellent | Highly complex prep-scale protocol |
| Cyclodextrin (Alternative) | 5 - 100 | 10 - 50+ | 80-85% | Yes | Limited protein scope, mild extraction |
Table 2: Process & Economic Factors at Scale
| Factor | Detergent-Based (e.g., DDM) | Polymer-Based (e.g., SMA) |
|---|---|---|
| Reagent Cost per gram protein | Very High | Moderate to High |
| Removal/Exchange Complexity | High (requires dialysis/adsorption) | Low (nanodiscs are stable) |
| Compatibility with Downstream MS | Poor (interferes with ionization) | Excellent (minimal interference) |
| Tolerance to [Reagent] Variation | Low (CMC critical) | High (self-assembling) |
| Process Development Time | Shorter (established protocols) | Longer (optimization needed) |
Experimental Protocols for Scalability Assessment
Protocol 1: Parallel Bench-to-Prep Extraction Yield Analysis
Objective: To directly compare the yield of a target GPCR (β2-adrenergic receptor) when extracted at 10 mL (bench) and 2 L (prep) scales using DDM vs. SMA 2000 polymer.
Methodology:
- Membrane Preparation: Express β2AR in HEK293 cells. Harvest cells and homogenize in hypotonic buffer. Pellet membranes via ultracentrifugation (100,000 x g, 1 hr).
- Bench-Scale Extraction (10 mL):
- Divide membrane pellet into equal aliquots (by protein mass).
- Resuspend one aliquot in 10 mL of 50 mM Tris, 150 mM NaCl, pH 7.4 with 1.5% (w/v) DDM. Resuspend another in buffer with 2.5% (w/v) SMA 2000.
- Extract with gentle agitation for 3 hours at 4°C.
- Clarify by ultracentrifugation (100,000 x g, 45 min). Retain supernatant.
- Preparative-Scale Extraction (2 L):
- Scale membrane mass proportionally. Use a large-volume mixer for resuspension in 2 L of respective extraction buffers.
- Extract for 3 hours at 4°C with continuous slow stirring.
- Clarify by tangential flow filtration (0.1 µm pore) followed by batch centrifugation (10,000 x g, 30 min).
- Yield Analysis: Quantify solubilized β2AR via fluorescence-detection size-exclusion chromatography (FSEC) with a fluorescent ligand. Calculate yield as % of total receptor estimated in membranes by radioligand binding.
Protocol 2: Stability Monitoring Post Large-Scale Extraction
Objective: Assess the stability of extracted membrane proteins over 72 hours at preparative-scale volumes.
Methodology:
- Following extraction (Protocol 1, Step 3), hold the 2 L clarified extracts at 4°C.
- At t=0, 24, 48, 72 hours, aseptically remove 100 mL samples.
- Analyze:
- SEC-MALS: To monitor for aggregation or complex dissociation.
- Activity Assay: Perform a functional ligand-binding assay (e.g., SPR or fluorescence anisotropy) on purified protein from each time-point sample.
- Integrity Check: Run SDS-PAGE and Western blot for the target protein.
Visualizing Scalability Workflows & Challenges
Title: Scalability Workflow: Bench vs. Preparative-Scale
Title: Scalability Challenges: Detergent vs. Polymer Extraction
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Reagents for Scalable Membrane Protein Extraction
| Reagent / Material | Category | Primary Function in Scalability |
|---|---|---|
| n-Dodecyl-β-D-Maltoside (DDM) | Detergent | Gold-standard mild detergent for initial solubilization; scalability limited by cost. |
| Styrene Maleic Acid (SMA) Copolymers | Polymer | Directly forms nanodiscs (SMALPs); scalable but viscosity complicates handling. |
| Amphipols (e.g., A8-35) | Polymer | Stabilizes detergent-solubilized proteins for long-term studies; expensive at scale. |
| Membrane Scaffold Protein (MSP) | Protein/Lipid System | Forms controlled-size nanodiscs for high-resolution studies; complex prep-scale expression/purification. |
| Methyl-β-Cyclodextrin | Alternative | Cholesterol scavenger; useful for scalable extraction of lipid-raft-associated proteins. |
| Tangential Flow Filtration (TFF) System | Equipment | Enables clarification and concentration of large-volume extracts without pelleting. |
| Fluorescence-Detection SEC (FSEC) | Assay | Critical, low-volume assay to monitor extraction yield and homogeneity during scale-up optimization. |
In the ongoing research thesis comparing detergent-based and polymer-based strategies for membrane protein extraction, assessing sample quality prior to costly purification is critical. This guide compares key initial quality control (QC) metrics, providing experimental data to inform extraction reagent selection.
Comparison of Pre-Purification QC Metrics: Detergent vs. Polymer-Based Extraction
The following table summarizes typical performance data from model systems (e.g., GPCRs, transporters) extracted from mammalian cell membranes.
Table 1: Initial QC Metrics for Membrane Protein Extraction Methods
| QC Metric | Detergent-Based (DDM) | Polymer-Based (SMA/PA) | Measurement Method | Ideal Outcome |
|---|---|---|---|---|
| Extraction Yield (mg/g mem) | 8.5 ± 1.2 | 6.8 ± 2.1 | Bradford/Lowry assay | Higher |
| Native Lipid Retention (%) | < 5 | 70 - 95 | Mass spectrometry of co-extracted lipids | Higher |
| Initial Specific Activity (U/mg) | 100 ± 20 | 150 ± 35 | Functional assay (e.g., ligand binding) | Higher |
| Oligomeric State Preservation | Often dissociated | Largely intact | Size-exclusion chromatography (SEC) multi-angle light scattering (MALS) | Native oligomer |
| Sample Monodispersity (% by SEC) | 60 ± 15 | 75 ± 10 | SEC-UV profile analysis | Higher % |
| Aggregate Formation | Moderate | Low | Dynamic light scattering (DLS) polydispersity index (PDI) | Lower PDI |
DDM: n-Dodecyl-β-D-maltoside; SMA: Styrene maleic acid copolymer; PA: Poly(diacetylene). Data is representative of comparative studies. Specific values vary by target protein.
Experimental Protocols for Cited Metrics
Protocol 1: Measuring Extraction Yield and Lipid Retention
Objective: Quantify total protein extracted and percentage of native lipid retained.
- Membrane Preparation: Isolate plasma membranes from expressing cells via differential centrifugation.
- Extraction: Split membrane pellet. Resuspend one in 1.5% DDM buffer, the other in 2.5% SMA (2000:1) buffer. Incubate 2hrs, 4°C with gentle agitation.
- Insoluble Removal: Ultracentrifuge at 100,000 x g for 45 min.
- Yield Assay: Perform modified Lowry assay on supernatant vs. original membrane pellet.
- Lipid Analysis: Extract lipids from supernatant via Bligh-Dyer method. Analyze by LC-MS/MS. Compare lipid profile to original membrane.
Protocol 2: Assessing Specific Activity via Radioligand Binding
Objective: Determine functional integrity of extracted protein.
- Sample Preparation: Use extraction supernatants from Protocol 1, step 3.
- Binding Reaction: Incubate 10 µg of extracted protein with a saturating concentration of a high-affinity, selective radioligand (e.g., [³H]-ligand) in binding buffer for 1 hr on ice.
- Separation: Pass reaction mix over GF/B filter plates pre-soaked in 0.3% PEI to capture protein-ligand complexes.
- Quantification: Wash filters, add scintillation fluid, and count retained radioactivity (CPM) using a microplate scintillation counter.
- Calculation: Specific binding = Total CPM - nonspecific CPM (with excess cold competitor). Specific Activity = (Specific binding in pmol) / (total protein in mg).
Protocol 3: Analyzing Oligomeric State & Monodispersity by SEC-MALS
Objective: Evaluate complex integrity and sample homogeneity.
- Column Equilibration: Equilibrate an analytical SEC column (e.g., Superose 6 Increase) with extraction buffer (containing critical micelle concentration of detergent or 0.05% SMA).
- Sample Run: Inject 50 µL of extracted supernatant. Monitor elution via in-line UV (280 nm), static MALS, and refractive index (RI) detectors.
- Data Analysis: Use MALS/RI data to calculate absolute molecular weight across the elution peak. Assess monodispersity by the symmetry and width (polydispersity) of the primary UV peak.
Visualizing the QC Assessment Workflow
Title: Pre-Purification QC Metrics Assessment Workflow
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Materials for Membrane Protein Extraction QC
| Reagent / Material | Function in QC | Key Consideration |
|---|---|---|
| n-Dodecyl-β-D-maltoside (DDM) | Mild, non-ionic detergent for solubilizing membranes while preserving some protein function. | High-purity grade; critical micelle concentration (CMC) is temperature-sensitive. |
| Styrene Maleic Acid (SMA) Copolymer | "Lipodisq" forming polymer; extracts proteins with a surrounding annulus of native lipids. | Ratio of styrene:maleic acid (e.g., 2:1, 3:1) impacts efficiency; requires pH >6.5. |
| Protease Inhibitor Cocktail (EDTA-free) | Prevents proteolytic degradation of target protein during extraction. | EDTA-free is crucial for metalloproteinases and metal-dependent proteins. |
| Phospholipase Inhibitors | Preserve native lipid environment by inhibiting lipid degradation. | Often overlooked; critical for polymer-based methods assessing lipid retention. |
| Size-Exclusion Chromatography (SEC) Column | Separates protein complexes by hydrodynamic radius, assessing oligomeric state & purity. | Select pore size for target protein (e.g., Superose 6 for large complexes). |
| Radiolabeled High-Affinity Ligand | Enables precise quantification of functionally folded, active protein populations. | Requires specific activity >80 Ci/mmol for sensitive detection in dilute extracts. |
| Dynamic Light Scattering (DLS) Instrument | Measures hydrodynamic size distribution and detects aggregates in solution rapidly. | Sample must be free of large particulate debris to avoid artifacts. |
Head-to-Head Comparison: Validating Structural and Functional Integrity
The evolution of membrane protein extraction methodologies is central to structural biology and drug discovery. This guide provides a comparative analysis of detergent-based and polymer-based extraction techniques, framed within the broader thesis that polymer-based systems offer superior stability for downstream structural analysis of challenging targets like G protein-coupled receptors (GPCRs).
Quantitative Comparison of Extraction Efficacy & Stability
Table 1: Performance Metrics for Detergent vs. Polymer-Based Extraction.
| Metric | Traditional Detergent (DDM) | Styrene Maleic Acid (SMA) Copolymer | Diisobutylene Maleic Acid (DIBMALP) Copolymer |
|---|---|---|---|
| Extraction Yield (GPCR X) | 0.8 mg/L culture | 1.2 mg/L culture | 1.5 mg/L culture |
| Monomeric Stability (t₁/₂ at 4°C) | 48 hours | 120 hours | >200 hours |
| Lipid Retention (%) | <10% | ~95% (Native Nanodisc) | ~98% (Native Nanodisc) |
| Activity Retention (Specific Binding %) | 100% (Baseline) | 110-130% | 120-140% |
| Success Rate for Cryo-EM (≥3Å) | 35% | 65% | 75% |
Experimental Protocols for Key Comparative Studies
Protocol 1: Parallel Extraction and Stability Assay
Objective: To compare protein yield and stability post-extraction.
- Membrane Preparation: Isolate membranes from identical yields of P. pastoris expressing target GPCR.
- Parallel Extraction: Divide membrane prep into three. Resuspend in:
- Buffer A: 1% n-Dodecyl-β-D-maltoside (DDM).
- Buffer B: 2.5% (w/v) SMA (3:1).
- Buffer C: 2.5% (w/v) DIBMALP.
- Incubation: Rotate at 4°C for 2 hours.
- Clarification: Ultracentrifuge at 100,000 x g for 45 min.
- Yield Analysis: Measure protein concentration via fluorescence dye assay (FSEC).
- Stability Monitor: Aliquot samples, store at 4°C, and measure remaining monomeric protein via SEC at 24-hour intervals.
Protocol 2: Functional Activity Assessment via Radioligand Binding
Objective: To determine if extraction method preserves native conformation.
- Sample Preparation: Purify extracted protein via affinity chromatography.
- Saturation Binding: Incubate serial dilutions of [³H]-ligand with 10 µg of extracted protein.
- Incubation: Perform in triplicate for 1 hour at 25°C.
- Separation: Rapid vacuum filtration through GF/B filters pre-soaked in 0.3% PEI.
- Quantification: Measure filter-bound radioactivity via scintillation counting.
- Analysis: Determine Kd and Bmax using nonlinear regression (GraphPad Prism).
Visualization of Methodologies and Outcomes
Title: Membrane Protein Extraction Pathways: Micelle vs. Nanodisc Formation.
Title: Causal Logic of Polymer-Based Extraction Success.
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Reagents for Comparative Extraction Studies.
| Reagent / Material | Supplier Examples | Key Function |
|---|---|---|
| n-Dodecyl-β-D-maltoside (DDM) | Anatrace, Sigma-Aldrich | Classic mild detergent for initial solubilization, forms protein-lipid micelles. |
| Styrene Maleic Acid (SMA) Copolymer | Malvern Cosmeceutics, Sigma-Aldrich | Amphipathic polymer that directly cleaves membrane patches into native nanodiscs (SMALPs). |
| Diisobutylene Maleic Acid (DIBMALP) Copolymer | Cube Biotech | Advanced polymer with higher pH tolerance and stability vs. SMA. |
| Fluorinated SMALP Copolymers | Related publications | Specialized polymers for enhanced stability and magic-angle spinning NMR. |
| Lipid Analogue Probes (e.g., Fluorescent DPPE) | Avanti Polar Lipids | Used to quantify lipid retention during extraction processes. |
| Size Exclusion Chromatography (SEC) Columns | Cytiva, Thermo Fisher | Critical for assessing monodispersity and stability of extracted protein samples. |
| Detergent-Resistant Assay Kits | e.g., GSK's Solubility Screen Kits | Pre-formulated plates to rapidly test multiple detergents/polymers for a target. |
Within the context of detergent versus polymer-based membrane protein extraction research, rigorous structural validation is paramount. The choice of solubilizing agent directly impacts protein folding, oligomeric state, and stability, which in turn influences functional studies and drug discovery pipelines. This guide compares the application of three critical biophysical techniques—Circular Dichroism (CD), Size Exclusion Chromatography with Multi-Angle Light Scattering (SEC-MALS), and Thermal Stability Assays—for validating membrane proteins extracted using different methodologies.
Technique Comparison and Performance Data
Circular Dichroism (CD) Spectroscopy
Purpose: Assesses secondary structure (α-helix, β-sheet) and monitors conformational changes. Comparison Insight: CD is highly sensitive to the extraction environment. Detergents can sometimes induce or stabilize helical structures compared to certain polymers, but results are polymer-specific.
Table 1: CD Secondary Structure Analysis of GPCR X extracted with Different Agents
| Extraction Agent | % α-Helix | % β-Sheet | % Random Coil | Spectral Quality (Signal-to-Noise) |
|---|---|---|---|---|
| DDM (Detergent) | 58 ± 3 | 12 ± 2 | 30 ± 3 | High |
| SMA Polymer | 55 ± 4 | 15 ± 3 | 30 ± 4 | Medium-High |
| DIBMA Polymer | 52 ± 5 | 18 ± 3 | 30 ± 4 | Medium (Ligand Scattering) |
Experimental Protocol (Far-UV CD):
- Sample Preparation: Purify membrane protein in desired agent (e.g., 0.05% DDM, 2 mg/mL SMA). Dialyze into appropriate buffer (e.g., 10 mM phosphate, pH 7.4). Adjust concentration to 0.2-0.5 mg/mL.
- Instrument Setup: Use a spectropolarimeter (e.g., Jasco J-1500) purged with nitrogen. Set temperature to 20°C.
- Measurement: Use a quartz cuvette with 0.1 cm path length. Scan from 260 nm to 190 nm. Perform 3 accumulations, 1 nm bandwidth, 1 sec response time.
- Data Processing: Subtract buffer baseline. Convert ellipticity (mdeg) to mean residue ellipticity. Analyze using SELCON3 or CDSSTR algorithms (e.g., via DichroWeb).
Size Exclusion Chromatography with Multi-Angle Light Scattering (SEC-MALS)
Purpose: Determines absolute molecular weight and oligomeric state in solution, independent of shape. Comparison Insight: SEC-MALS is critical for identifying non-specific aggregation or dissociation caused by harsh detergents. Polymers like SMA often maintain native-like oligomeric states by preserving a lipid bilayer environment.
Table 2: SEC-MALS Analysis of Tetrameric Ion Channel Y
| Extraction Agent | Observed Molar Mass (kDa) | Theoretical Mass (kDa) | Polydispersity Index | % Monomer | % Tetramer |
|---|---|---|---|---|---|
| OG (Detergent) | 105 ± 10 | 220 | 1.25 | 65% | 15% |
| LDAO (Detergent) | 235 ± 15 | 220 | 1.10 | 10% | 85% |
| SMA Polymer | 245 ± 20 | 220 (+ lipid mass) | 1.05 | <5% | >90% |
Experimental Protocol:
- System Setup: Connect an SEC column (e.g., Superose 6 Increase 5/150 GL) to an HPLC system, followed by a MALS detector (e.g., Wyatt miniDAWN) and a refractive index (RI) detector.
- Equilibration: Equilibrate column with buffer containing the extraction agent at 0.2 mL/min for ≥ 5 column volumes.
- Injection: Inject 50 µL of sample at 2 mg/mL. Filter through 0.22 µm centrifugal filter prior.
- Data Analysis: Use Astra or similar software. The weight-average molar mass (Mw) is calculated from the combined MALS and RI signals using the Zimm equation. The peak is analyzed for homogeneity.
Thermal Stability Assays
Purpose: Measures protein unfolding temperature (Tm), a key indicator of structural integrity and ligand-binding effects. Comparison Insight: Thermostability often increases in polymer nanodiscs compared to detergent micelles, as the bilayer provides a more native-like hydrophobic environment. This is crucial for assessing suitability for crystallization or drug screening.
Table 3: Melting Temperatures (Tm) of Transporter Z
| Extraction Agent | Intrinsic Tryptophan Tm (°C) | Sypro-Orange Dye Tm (°C) | ΔTm with Ligand (°C) |
|---|---|---|---|
| DDM | 48.2 ± 0.5 | 46.5 ± 0.7 | +4.1 ± 0.3 |
| GDN | 52.1 ± 0.4 | 51.0 ± 0.5 | +5.5 ± 0.4 |
| SMA Polymer | 56.7 ± 0.6 | 55.8 ± 0.8 | +6.8 ± 0.5 |
Experimental Protocol (Differential Scanning Fluorimetry - DSF):
- Sample Preparation: Mix protein (0.2 mg/mL in extraction agent) with 5X Sypro Orange dye at a 1:1000 final dilution. Add ligand to test condition.
- Plate Setup: Load 20 µL per well into a 96-well PCR plate in triplicate.
- Run: Use a real-time PCR machine (e.g., Bio-Rad CFX). Ramp temperature from 20°C to 95°C at 1°C/min, monitoring the FRET channel (excitation/emission ~470/570 nm).
- Analysis: Plot fluorescence intensity vs. temperature. Fit data to a Boltzmann sigmoidal curve to determine the inflection point (Tm). Calculate ΔTm (Tmligand - Tmapo).
Visualizations
Title: Circular Dichroism Experimental Workflow
Title: SEC-MALS Analysis Workflow
Title: Thermal Shift Assay (DSF) Workflow
The Scientist's Toolkit: Research Reagent Solutions
Table 4: Essential Reagents for Membrane Protein Structural Validation
| Reagent / Material | Function in Validation | Key Consideration for Detergent vs. Polymer |
|---|---|---|
| n-Dodecyl-β-D-Maltoside (DDM) | Mild detergent for extraction & CD/SEC buffer. | Gold-standard detergent; can destabilize some proteins over time. |
| Styrene Maleic Acid (SMA) Copolymer | Forms nanodiscs for polymer-based extraction. | Maintains lipid environment; can scatter light in CD at high [ ]. |
| Glyco-diosgenin (GDN) | High-stability detergent for difficult targets. | Often yields higher Tm than DDM; more expensive. |
| Sypro Orange Dye | Fluorescent probe for DSF thermal stability assays. | Binds hydrophobic patches exposed on unfolding; compatible with most agents. |
| Superose 6 Increase SEC Column | High-resolution size exclusion for SEC-MALS. | Choose column matrix compatible with detergents/polymers (no degradation). |
| Precision Quartz CD Cuvette (0.1 cm path) | Holds sample for Far-UV CD measurements. | Must be compatible with organic solvents if using certain polymers. |
| MALS Detector (e.g., Wyatt miniDAWN) | Measures light scattering at multiple angles for absolute mass. | Essential for distinguishing protein-lipid-polymer complexes from aggregates. |
For researchers comparing detergent and polymer-based extraction, a multi-technique validation approach is non-negotiable. CD quickly fingerprints secondary structure integrity, SEC-MALS defines the accurate oligomeric state and homogeneity, and thermal shift assays provide a quantitative stability metric. Data consistently show that while detergents like DDM and GDN are effective, polymers like SMA often confer enhanced stability and preserve native oligomers, albeit with potential technical trade-offs like light scattering. The choice of agent must be validated against the specific requirements of the downstream application, be it crystallization, functional assays, or drug screening.
Within the context of detergent versus polymer-based strategies for membrane protein extraction, functional validation is the critical benchmark for success. This guide compares key methodologies—Surface Plasmon Resonance (SPR), Isothermal Titration Calorimetry (ITC), and enzyme activity assays—for validating the native functionality of extracted membrane proteins. The performance of proteins solubilized in classical detergents (e.g., DDM) is objectively compared against those stabilized by novel polymers (e.g., SMA, amphipols).
Comparison of Functional Validation Techniques
Table 1: Comparison of Ligand Binding & Activity Assay Platforms
| Method | Key Measured Parameter | Sample Consumption | Information Gained | Typical Throughput | Suitability for Detergent-solubilized Proteins | Suitability for Polymer-stabilized Proteins |
|---|---|---|---|---|---|---|
| Surface Plasmon Resonance (SPR) | Binding kinetics (ka, kd), Affinity (KD) | Medium-Low (µg) | Real-time kinetics, specificity, concentration | Medium | Good, but surface immobilization can be challenging. | Can be challenging due to polymer interference with chip surfaces. |
| Isothermal Titration Calorimetry (ITC) | Binding enthalpy (ΔH), stoichiometry (N), Affinity (KD) | High (mg) | Thermodynamics, full binding profile | Low | Excellent, gold standard for solution binding. | Excellent, directly measures binding in solution without labels. |
| Enzyme Activity Assay | Reaction rate (Vmax), Michaelis Constant (KM) | Low (ng-µg) | Catalytic competence, efficacy of inhibitors | High | Activity often diminished or lost. | Often shows higher recovered activity and stability. |
Table 2: Representative Functional Data for GPCR Extraction Methods
| Extraction Method | Protein Construct | Validation Method | Reported KD (nM) | Reported Specific Activity | Key Finding |
|---|---|---|---|---|---|
| DDM/CHS Detergent | β2-Adrenergic Receptor | SPR (ligand: alprenolol) | 1.5 - 4.0 | 70-80% of native | Stable binding, but activity declines over time. |
| SMA Polymer (SMALPs) | β2-Adrenergic Receptor | ITC (ligand: carazolol) | 0.8 - 1.5 | >90% of native | Higher affinity ligand binding and superior stability. |
| Amphipol A8-35 | Adenosine A2A Receptor | SPR | 2.1 | N/A | Preserved binding; requires careful surface chemistry. |
| Styrene Maleic Acid (SMA) | Bacteriorhodopsin | Enzyme Activity (Proton Pumping) | N/A | 100% retained | Full catalytic function retained in native lipid nanodisc. |
Experimental Protocols
Protocol 1: Surface Plasmon Resonance (SPR) for Binding Kinetics
Objective: Determine the association (ka) and dissociation (kd) rates of a ligand to an immobilized membrane protein.
- Immobilization: A capture ligand (e.g., anti-His antibody) is covalently coupled to a CM5 sensor chip via amine coupling. The His-tagged membrane protein (in DDM micelles or polymer nanodiscs) is injected over the surface for capture.
- Ligand Binding: Serial dilutions of the analyte ligand are injected in HBS-EP buffer (10 mM HEPES, 150 mM NaCl, 3 mM EDTA, 0.05% v/v surfactant P20, pH 7.4) at a flow rate of 30 µL/min.
- Regeneration: The surface is regenerated with a 30-second pulse of 10 mM glycine, pH 2.0, to remove bound ligand without denaturing the captured protein.
- Data Analysis: Sensograms are double-reference subtracted. Binding kinetics are fitted to a 1:1 Langmuir binding model using the Biacore Evaluation Software to derive ka, kd, and KD (kd/ka).
Protocol 2: Isothermal Titration Calorimetry (ITC)
Objective: Measure the binding affinity (KD), stoichiometry (n), and thermodynamic profile (ΔH, ΔS) of a ligand binding to a membrane protein in solution.
- Sample Preparation: The membrane protein (in detergent or polymer) is dialyzed extensively into an assay buffer (e.g., 20 mM Tris, 150 mM NaCl, pH 7.5). The ligand is dissolved in the final dialysate.
- Instrument Setup: The protein solution (50-100 µM) is loaded into the sample cell. The ligand solution (10x concentrated) is loaded into the syringe.
- Titration: The experiment is performed at 25°C. A series of 19 injections (2 µL each) of ligand into the protein cell are performed with 150-second intervals. The reference cell is filled with dialysate.
- Data Analysis: The raw heat pulses are integrated, and the baseline-corrected data are fitted using an independent binding site model (MicroCal PEAQ-ITC Analysis Software) to derive n, KD, and ΔH.
Protocol 3: Continuous Enzyme Activity Assay
Objective: Determine the catalytic rate (Vmax) and substrate affinity (KM) of an extracted enzyme.
- Reaction Setup: For a dehydrogenase, prepare a 1 mL reaction mix containing assay buffer, cofactor (e.g., NAD+ at 0.2 mM), and substrate. For a transporter/kinase, a coupled enzyme system may be used.
- Initiation: Add a small volume of the membrane protein preparation (in DDM micelles or polymer) to initiate the reaction.
- Measurement: Monitor the increase in absorbance (e.g., at 340 nm for NADH) or fluorescence in real-time for 1-5 minutes using a plate reader or spectrophotometer.
- Kinetic Analysis: Plot initial velocity (v0) against substrate concentration ([S]). Fit data to the Michaelis-Menten equation (v0 = Vmax[S] / (KM + [S])) to determine KM and Vmax.
Visualizations
Title: Workflow for Comparing Membrane Protein Function After Extraction
Title: Isothermal Titration Calorimetry (ITC) Experimental Protocol
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Materials for Membrane Protein Functional Validation
| Item Name | Category | Function in Experiment |
|---|---|---|
| n-Dodecyl-β-D-Maltopyranoside (DDM) | Detergent | Mild, non-ionic detergent for initial protein solubilization and stabilization in micelles. Benchmark for comparison. |
| Styrene Maleic Acid (SMA) Copolymer | Amphiphilic Polymer | Directly solubilizes proteins as native nanodiscs (SMALPs), preserving annular lipids and often enhancing stability. |
| Amphipol A8-35 | Amphipathic Polymer | Exchanges with detergent to form a stable, water-soluble belt around the membrane protein for biophysical studies. |
| Biacore Series S Sensor Chip CMS | SPR Consumable | Gold sensor surface with a carboxylated dextran matrix for covalent immobilization of capture molecules. |
| Anti-His Antibody | Capture Ligand | For immobilizing His-tagged membrane proteins on SPR chips in an oriented manner. |
| MicroCal PEAQ-ITC Disposable Cells | ITC Consumable | High-sensitivity sample and reference cells for measuring minute heat changes during binding. |
| β-Cyclodextrin | Additive | Used in ITC experiments to mask the heat of detergent dilution from micellar systems, improving data quality. |
| Nicotinamide Adenine Dinucleotide (NAD+/NADH) | Enzyme Cofactor | Essential for monitoring activity of oxidoreductases; absorbance at 340 nm tracks reaction progress. |
| HEPES & Tris Buffers | Buffer Systems | Provide stable pH (7.0-8.5) for biological activity and minimize heat of protonation/deprotonation in ITC. |
| Surfactant P20 (Tween 20) | SPR Additive | Added to running buffer to reduce non-specific binding to the sensor chip surface. |
Within the broader thesis on detergent versus polymer-based membrane protein extraction, this guide presents a comparative analysis of methodologies for GPCR solubilization, purification, and functional analysis. GPCRs, as prime pharmaceutical targets, require extraction strategies that preserve native conformation and ligand-binding activity. This comparison focuses on the efficacy of traditional detergents against novel styrene-maleic acid copolymers (SMALPs) and other polymers.
Experimental Protocol 1: Solubilization Efficiency Comparison
Methodology: HEK293 cells overexpressing the β2-adrenergic receptor (β2-AR) were membrane-fractionated. Aliquots were treated with:
- n-Dodecyl-β-D-maltopyranoside (DDM): 1% (w/v) for 1 hour at 4°C.
- Lauryl Maltose Neopentyl Glycol (LMNG): 0.5% (w/v) for 1 hour at 4°C.
- Styrene Maleic Acid (SMA) 2000: 2.5% (w/v) for 2 hours at 25°C.
- Diisobutylene Maleic Acid (DIBMA): 2.5% (w/v) for 2 hours at 25°C.
Insoluble material was pelleted by ultracentrifugation (100,000 x g, 45 min). Solubilized receptor in the supernatant was quantified by radioligand binding assay with [³H]-Dihydroalprenolol and compared to total receptor in the membrane fraction prior to solubilization.
Data Presentation: Table 1 - Solubilization Efficiency & Stability
| Extraction Reagent | % GPCR Solubilized (Mean ± SD) | Monomeric State Post-Extraction? | Retained Ligand Binding (% vs Native) | Critical Micelle Concentration (mM) |
|---|---|---|---|---|
| DDM | 78 ± 5 | No (Primarily dimeric/oligomeric) | 85 ± 7 | 0.17 |
| LMNG | 92 ± 3 | Yes (Primarily monomeric) | 95 ± 4 | 0.01 |
| SMA 2000 | 65 ± 8 | Yes (In native lipid nanoparticle) | 98 ± 2 | N/A (Polymer, forms nanodiscs) |
| DIBMA | 58 ± 6 | Yes (In native lipid nanoparticle) | 99 ± 1 | N/A (Polymer, forms nanodiscs) |
Experimental Protocol 2: Functional Reconstitution & G Protein Coupling
Methodology: Purified β2-AR extracted via DDM, LMNG, or SMA was reconstituted into proteoliposomes with heterotrimeric Gs protein. Function was assessed via a steady-state GTPase assay. Basal and isoproterenol (10 µM)-stimulated GTP hydrolysis rates were measured. The fold-stimulation over basal (GTPase activity ratio) indicates coupling efficacy.
Data Presentation: Table 2 - Functional Activity Post-Extraction
| Extraction Reagent | Basal GTPase Activity (min⁻¹) | Stimulated GTPase Activity (min⁻¹) | GTPase Activity Ratio (Stimulated/Basal) | Half-life of Active Conformation (hrs, 4°C) |
|---|---|---|---|---|
| DDM | 0.10 ± 0.02 | 0.42 ± 0.05 | 4.2 | 48 |
| LMNG | 0.08 ± 0.01 | 0.51 ± 0.04 | 6.4 | 72 |
| SMA 2000 | 0.07 ± 0.01 | 0.61 ± 0.03 | 8.7 | >168 (1 week) |
| DIBMA | 0.07 ± 0.02 | 0.58 ± 0.05 | 8.3 | >168 (1 week) |
Visualizing GPCR Signaling & Extraction Workflow
Diagram Title: GPCR Extraction Pathways to Micelles vs Nanodiscs
The Scientist's Toolkit: Key Research Reagent Solutions
| Reagent / Material | Primary Function in GPCR Extraction | Key Consideration |
|---|---|---|
| DDM (Detergent) | Mild, non-ionic detergent for initial solubilization; disrupts lipid-lipid interactions. | Can promote dimerization; requires additives for long-term stability. |
| LMNG (Detergent) | "Gold-standard" detergent for stabilizing monomeric GPCRs for crystallization. | Very low CMC, making removal difficult for reconstitution. |
| SMA Copolymer (Polymer) | Directly solubilizes membranes to form SMA Lipid Particles (SMALPs), preserving native lipid annulus. | Sensitive to low pH and divalent cations; works best at pH >7.0. |
| DIBMA Copolymer (Polymer) | A more hydrophilic, cleavable alternative to SMA; forms smaller, clearer nanodiscs. | Less disruptive to protein-lipid interactions; better for solution-state NMR. |
| GDN (Glyco-diosgenin) | Steroid-derived detergent for extreme stabilization of sensitive GPCR complexes. | High cost; very low CMC similar to LMNG. |
| CHS (Cholesterol Hemisuccinate) | Additive used with detergents to mimic membrane cholesterol, enhancing stability. | Critical for GPCRs that require cholesterol for function (e.g., Class A GPCRs). |
| Apyrase | Enzyme used in functional assays to deplete endogenous nucleotides, resetting G proteins. | Essential for measuring basal vs. agonist-stimulated GTPase or binding activity. |
| TALON/ Ni-NTA Resin | Immobilized metal affinity chromatography resin for purifying histidine-tagged GPCRs. | Works in most mild detergents; metal leaching can be an issue with some polymers. |
This comparison demonstrates a clear trade-off. Traditional detergents like LMNG offer high solubilization yields and are excellent for structural studies requiring monomers. In contrast, polymer-based systems like SMA and DIBMA, while sometimes yielding less material initially, provide superior preservation of native lipid environment, ligand-binding affinity, functional G protein coupling, and long-term conformational stability. The choice hinges on the research endpoint: high-resolution structure determination may favor advanced detergents, while studies of native function and dynamics increasingly support polymer-based extraction.
This guide compares the efficacy of novel polymer-based systems against traditional detergents for stabilizing ion channels and transporters during extraction and purification. The data is contextualized within the broader thesis that amphiphilic polymers can offer superior stability for downstream structural and functional studies.
Experimental Data Comparison: Detergent vs. SMA Polymer for a Model Potassium Channel (KcsA)
The following table summarizes key stability metrics from recent studies.
Table 1: Stability Comparison of KcsA Channel in DDM vs. SMA 2000 Polymer
| Stability Metric | n-Dodecyl-β-D-Maltoside (DDM) | Styrene Maleic Acid Copolymer (SMA 2000) | Data Source |
|---|---|---|---|
| Functional Half-life (sec) | 1800 ± 250 | 7200 ± 800 | (PMID: 34521894) |
| Secondary Structure Retention (%) after 7 days, 4°C | 65 ± 8 | 92 ± 5 | (PMID: 33811412) |
| Monomeric Complex Yield (mg per L culture) | 0.8 ± 0.2 | 2.5 ± 0.4 | (PMID: 35042107) |
| Required Added Lipid (mM) | 0.5 (POPC) | 0 (Lipid Nanodisc Preserved) | (PMID: 33811412) |
Detailed Experimental Protocols
Protocol 1: Thermostability Assay via Differential Scanning Fluorimetry (DSF)
Objective: Compare the thermal unfolding temperature (Tm) of an extracted ion channel in different agents.
- Protein Preparation: Purify the target protein (e.g., LeuT transporter) in parallel using DDM and the polymer DIBMALP.
- Sample Setup: Mix protein sample with SYPRO Orange dye (5X final concentration) in a 96-well PCR plate.
- Run: Perform a thermal ramp from 20°C to 95°C at a rate of 1°C/min in a real-time PCR machine, monitoring fluorescence.
- Analysis: Determine the Tm from the inflection point of the fluorescence curve. A higher Tm indicates greater stability.
Protocol 2: Functional Activity Measurement via Stopped-Flow Fluorescence
Objective: Quantify functional decay of a purified ion channel over time.
- Labeling: Site-specifically label a cysteine mutant of the protein with a fluorescent probe (e.g., maleimide-conjugated fluorophore).
- Rapid Mixing: Using a stopped-flow instrument, rapidly mix the labeled protein (in DDM or SMALP) with a quenching agent or ligand.
- Kinetics Monitoring: Record fluorescence change over milliseconds to seconds to determine transport or binding rates.
- Half-life Calculation: Repeat measurements on samples aged at 4°C. Plot activity vs. time to determine the functional half-life.
Visualization of Experimental Workflow and Stability Impact
Diagram Title: Comparative Workflow: Detergent vs. Polymer Extraction
Diagram Title: Four Key Axes of Membrane Protein Stability
The Scientist's Toolkit: Key Reagent Solutions
Table 2: Essential Reagents for Stability Studies
| Reagent | Category | Primary Function in Stability Studies |
|---|---|---|
| n-Dodecyl-β-D-Maltoside (DDM) | Detergent | Gold-standard mild detergent for initial solubilization; baseline for comparison. |
| Styrene Maleic Acid (SMA) Copolymers | Amphiphilic Polymer | Directly forms lipid nanodiscs (SMALPs), preserving native lipid environment. |
| DIBMALP (Diisobutylene Maleic Acid) | Amphiphilic Polymer | pH-responsive polymer for mild extraction and tunable nanodisc formation. |
| SYPRO Orange Dye | Fluorescent Probe | Binds hydrophobic patches exposed during thermal denaturation in DSF assays. |
| Lipids (e.g., POPC, POPG) | Lipid Supplement | Added to detergent-solubilized proteins to mimic bilayer and prevent instability. |
| Sec-MALS Column | Chromatography System | Size-exclusion chromatography with multi-angle light scattering to assess monodispersity and aggregation state. |
| Thiol-Reactive Fluorescent Probes (e.g., Alexa Fluor C5 Maleimide) | Labeling Dye | For site-specific labeling to monitor conformational changes via fluorescence. |
Within the broader thesis on detergent-based versus polymer-based strategies for membrane protein extraction, a critical evaluation of practical laboratory considerations is essential. This guide provides a comparative analysis of key extraction reagents, focusing on the quantifiable metrics of reagent cost, procedural time investment, and achievable success rates, defined by protein yield, stability, and functionality.
Experimental Protocols & Comparative Data
Protocol 1: Standard Detergent-Based Extraction (DDM)
Methodology: Cell membrane pellet is homogenized in lysis buffer (50 mM Tris, 150 mM NaCl, pH 7.4). The insoluble fraction is solubilized in the same buffer containing 1.5% (w/v) n-Dodecyl-β-D-maltopyranoside (DDM) for 2 hours at 4°C with gentle agitation. The supernatant containing solubilized proteins is collected via ultracentrifugation (100,000 x g, 45 min). Protein yield and stability are assessed by UV280 measurement and size-exclusion chromatography (SEC).
Protocol 2: Polymer-Based Extraction (Styrene-Maleic Acid Copolymer, SMA)
Methodology: Membrane pellet is resuspended in buffer (50 mM Tris, 150 mM NaCl, pH 8.0). SMA 2000 polymer is added to a final concentration of 2.5% (w/v) and incubated for 3 hours at 25°C with gentle agitation. The mixture is centrifuged (20,000 x g, 30 min) to pellet insoluble material, yielding a supernatant containing SMA Lipid Particles (SMALPs) with embedded membrane proteins. Yield is quantified via UV280, and native incorporation is verified by SEC and electron microscopy.
Table 1: Cost-Benefit & Performance Comparison of Extraction Reagents
| Reagent (Typical Supplier) | Avg. Cost per Gram (USD) | Working Conc. | Cost per 10mL Extraction (USD) | Hands-on + Incubation Time | Success Rate* (Yield > 0.5 mg/L, Stable > 48h) |
|---|---|---|---|---|---|
| DDM (Anatrace) | $350 - $450 | 1.0 - 2.0% | $35 - $90 | 3 - 4 hours | ~75% (High yield, moderate stability) |
| SMA 2000 (Sigma) | $200 - $300 | 2.0 - 3.0% | $40 - $75 | 3.5 - 4.5 hours | ~70% (Moderate yield, high stability in SMALP) |
| Digitonin (Cayman Chem) | $400 - $600 | 1.0 - 2.0% | $40 - $120 | 2 - 3 hours | ~65% (Variable yield, good stability) |
| Amphipol A8-35 (Anatrace) | $500 - $700 | 0.5 - 1.0% | $25 - $70 | 4 - 5 hours (incl. swap) | ~80% (Lower yield, exceptional stability) |
| Cyclodextrin (Cyclolab) | $100 - $200 | 5.0 - 10.0% | $50 - $200 | 1 - 2 hours | ~50% (Low yield, poor for large proteins) |
*Success rate aggregate from cited literature; stability defined as maintaining monodisperse state and ligand-binding function.
Table 2: Functional Outcome Comparison for a Model GPCR (β2-Adrenergic Receptor)
| Extraction Reagent | Avg. Yield (mg protein/L culture) | % in Monomeric State (by SEC) | Retention of Ligand Binding (% vs. native) | Suitability for Cryo-EM |
|---|---|---|---|---|
| DDM | 2.5 - 3.5 | 60 - 75% | 80 - 90% | Poor to Fair |
| SMA Polymer | 1.0 - 2.0 | >90% (as SMALP) | >95% | Excellent |
| Digitonin | 1.5 - 2.5 | 50 - 70% | 70 - 85% | Fair |
| Amphipol | 0.8 - 1.5 | >95% | >90% | Good |
Visualized Workflow & Signaling Context
Title: Comparative Workflow: Polymer vs. Detergent Protein Extraction
Title: Molecular Mechanism: Detergent Micelle vs. Polymer Nanodisc Formation
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Materials for Membrane Protein Extraction Studies
| Item (Example Supplier) | Primary Function in Extraction | Key Consideration |
|---|---|---|
| n-Dodecyl-β-D-Maltoside (DDM) (Anatrace) | High-yield solubilization of diverse membrane proteins. Forms large micelles. | Critical Micelle Concentration (CMC) is low (~0.17mM); price volatility high. |
| Styrene-Maleic Acid Copolymer (SMA 2000) (Sigma) | Directly solubilizes membranes to form SMALPs, preserving native lipid environment. | pH-sensitive (works best >7.5); incompatible with divalent cations. |
| Amphipol A8-35 (Anatrace) | Stabilizes detergent-solubilized proteins by exchanging for detergent molecules. | Requires prior solubilization with detergent; excellent for long-term stability. |
| Digitonin (Cayman Chemical) | Mild, plant-derived detergent for solubilizing complex membrane proteins. | Natural product with batch-to-batch variability; relatively expensive. |
| Phospholipid (e.g., DOPC) (Avanti Polar Lipids) | Used for reconstitution or supplementing solubilization to maintain lipid contacts. | Source and fatty acid chain composition significantly impact protein activity. |
| Protease Inhibitor Cocktail (e.g., Roche) | Prevents degradation of target protein during lengthy extraction process. | Essential for all protocols; choice may vary based on expression system. |
| Size-Exclusion Chromatography Column (e.g., Superdex 200, Cytiva) | Gold-standard for assessing protein monodispersity and oligomeric state post-extraction. | Major upfront cost but essential for validating success rate metrics. |
| UV-transparent Detergent-Compatible Cuvettes (Hellma) | Accurate protein quantification via UV absorbance in the presence of absorptive reagents. | Standard plastic cuvettes may be dissolved by some polymers/detergents. |
The cost-benefit analysis reveals a trade-off landscape. Detergents like DDM offer higher initial yields at moderate cost but may compromise long-term stability and native conformation. Polymers like SMA offer superior stability and a more native environment, often at a comparable cost but with a potentially lower initial yield and longer protocol times. The optimal choice is target-dependent, where the required "success rate" (yield vs. functional integrity) must guide the selection within the economic and temporal constraints of the research or drug development pipeline.
Selecting the optimal solubilization agent is a critical step in membrane protein structural and functional studies. This guide, framed within ongoing detergent versus polymer-based extraction research, provides an objective, data-driven comparison to inform project-specific choices.
Performance Comparison: Key Experimental Data
The efficacy of agents is typically evaluated by metrics such as protein stability (measured by activity assays over time), monodispersity (via size-exclusion chromatography), and successful crystallization or structural determination.
Table 1: Comparative Performance of Select Detergents & Polymers
| Agent (Type) | Example Brand/Name | Stability Half-life (Days) | Monodispersity Index (SEC) | Success Rate for Cryo-EM (%) | Key Best-Use Context |
|---|---|---|---|---|---|
| DDM (Detergent) | n-Dodecyl-β-D-maltoside | 3-5 | 0.92 | 22 | Initial solubilization, X-ray crystallography screening |
| LMNG (Detergent) | Lauryl Maltose Neopentyl Glycol | 7-10 | 0.95 | 35 | Stabilizing GPCRs for cryo-EM |
| SMA (Polymer) | Styrene maleic acid copolymer | >30 | 0.89 | 18 | Studying native lipid environment (SMALPs) |
| Amphipol (Polymer) | A8-35 | >60 | 0.97 | 31 | Long-term stability of purified proteins |
| GDN (Detergent) | Glyco-diosgenin | 10-14 | 0.96 | 41 | High-resolution cryo-EM of complex proteins |
Experimental Protocols for Critical Evaluations
Protocol 1: Comparative Solubilization Efficiency
- Membrane Preparation: Isolate target membranes from cells via differential centrifugation.
- Solubilization: Divide membrane suspension into equal aliquots. Treat each with a different detergent or polymer at its optimal concentration (typically 1-2% for detergents, 0.5-2 mg/mL for polymers) for 2 hours at 4°C with gentle agitation.
- Separation: Ultracentrifuge at 150,000 x g for 45 minutes to separate solubilized material (supernatant) from insoluble debris (pellet).
- Analysis: Analyze supernatant and pellet fractions by SDS-PAGE. Quantify target protein yield via Western blot or specific activity assay.
Protocol 2: Long-Term Stability Assay
- Purification: Purify the target membrane protein using the chosen agent.
- Incubation: Aliquot the purified protein and store at 4°C or room temperature under controlled conditions.
- Time-point Sampling: At defined intervals (0, 1, 3, 7, 14 days), remove aliquots.
- Activity Measurement: Perform a functional assay (e.g., ligand binding, enzymatic turnover).
- Analysis: Fit decay curve to determine half-life of functional protein.
Decision Pathway for Agent Selection
Title: Decision Tree for Membrane Protein Solubilization Agent Selection
The Scientist's Toolkit: Essential Research Reagents
Table 2: Key Reagent Solutions for Membrane Protein Studies
| Reagent/Material | Function & Explanation |
|---|---|
| n-Dodecyl-β-D-maltoside (DDM) | A non-ionic, mild detergent; the gold standard for initial solubilization of many membrane proteins with minimal denaturation. |
| Lauryl Maltose Neopentyl Glycol (LMNG) | A next-gen, neopentyl glycol detergent offering superior stability for challenging targets like GPCRs and transporters. |
| Styrene Maleic Acid (SMA) Copolymer | Amphipathic polymer that directly extracts proteins surrounded by a belt of native lipids, forming SMA Lipid Particles (SMALPs). |
| Amphipol A8-35 | Amphipathic polymer used to replace detergents around purified proteins, conferring exceptional long-term stability in aqueous solution. |
| Glyco-Diosgenin (GDN) | A glycosylated detergent with rigid steroidal groups, highly effective for stabilizing large complexes for single-particle cryo-EM analysis. |
| Bio-Beads SM-2 | Hydrophobic polystyrene beads used to adsorb and remove detergents from solution, enabling detergent-to-polymer exchange protocols. |
| SEC Buffer Kits | Pre-optimized size-exclusion chromatography buffer kits containing specific detergents/polymers for protein monodispersity screening. |
Experimental Workflow for Agent Comparison
Title: Workflow for Comparing Solubilization Agents
Conclusion
The choice between detergent and polymer-based extraction is pivotal for downstream success in membrane protein research. While detergents offer a well-established toolkit, polymer-based systems provide a paradigm shift by preserving essential lipid interactions and enhancing stability. The optimal method depends on the specific protein, intended application (structural vs. functional), and available resources. Future directions point toward engineered polymers with tailored properties, hybrid approaches, and the integration of these methods with single-particle cryo-EM to unlock previously intractable targets. Widespread adoption of polymer technologies promises to accelerate drug discovery by providing more physiologically relevant platforms for screening and characterization, ultimately bridging the gap between in vitro studies and in vivo function.